Journal:Bridging the collaboration gap: Real-time identification of clinical specimens for biomedical research
| Full article title | Bridging the collaboration gap: Real-time identification of clinical specimens for biomedical research |
|---|---|
| Journal | Journal of Pathology Informatics |
| Author(s) | Durant, Thomas J.S.; Gong, Guannan; Price, Nathan; Schilz, Wade L. |
| Author affiliation(s) | Yale New Haven Hospital, Yale New Haven Health |
| Primary contact | Email: Log in required |
| Year published | 2020 |
| Volume and issue | 11 |
| Article # | 14 |
| DOI | 10.4103/jpi.jpi_15_20 |
| ISSN | 2153-3539 |
| Distribution license | Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License |
| Website | http://www.jpathinformatics.org/article.asp |
| Download | http://www.jpathinformatics.org/temp/JPatholInform11114-7412119_203521.pdf |
Abstract
Introduction: Biomedical and translational research often relies on the evaluation of patients or specimens that meet specific clinical or laboratory criteria. The typical approach used to identify biospecimens is a manual, retrospective process that exists outside the clinical workflow. This often makes biospecimen collection cost prohibitive and prevents the collection of analytes with short stability times. Emerging data architectures offer novel approaches to enhance specimen-identification practices. To this end, we present a new tool that can be deployed in a real-time environment to automate the identification and notification of available biospecimens for biomedical research.
Methods: Real-time clinical and laboratory data from Cloverleaf (Infor, NY, NY) were acquired within our computational health platform, which is built on open-source applications. Study-specific filters were developed in NiFi (Apache Software Foundation, Wakefield, MA, USA) to identify the study-appropriate specimens in real time. Specimen metadata were stored in Elasticsearch (Elastic N. V., Mountain View, CA, USA) for visualization and automated alerting.
Results: Between June 2018 and December 2018, we identified 2,992 unique specimens belonging to 2,815 unique patients, split between two different use cases. Based on laboratory policy for specimen retention and study-specific stability requirements, secure e-mail notifications were sent to investigators to automatically notify them of availability. The assessment of throughput on commodity hardware demonstrates the ability to scale to approximately 2,000 results per second.
Conclusion: This work demonstrates that real-world clinical data can be analyzed in real-time to increase the efficiency of biospecimen identification with minimal overhead for the clinical laboratory. Future work will integrate additional data types, including the analysis of unstructured data, to enable more complex cases and biospecimen identification.
Keywords: biobanking, biomedical research, biospecimen science, clinical specimens, real-time identification, translational research
Introduction
In the era of precision medicine, human biospecimens are an important resource for basic, translational, and clinical research and are increasingly needed to advance our understanding of human physiology, disease, treatment response, and outcomes. The field of biobanking has undergone significant optimization efforts by national and international communities to improve and harmonize biospecimen curation to support this need.[1][2] However, the operationalization and maintenance of biobanks is resource-intensive and often cost prohibitive for many institutions. In addition, long-term biobanking may be suboptimal for some types of testing, such as for studies that rely on labile analytes.[3][4] As a result, comprehensive access to human biospecimens remains limited, and there is a persistent need for efficient solutions that can provide access to high-quality and recently acquired human biospecimens.[5]
Human biospecimens can always be found in clinical laboratories, but access for the research is complicated by a series of technical, logistic, regulatory, and ethical challenges. Beyond the demands of delivering clinical results, laboratories lack efficient processes for biospecimen identification, human resources for specimen acquisition, and procedural infrastructure for biospecimen collection under the provisions of human interventional ethics committees. Despite these challenges, the clinical laboratory is a promising resource for the acquisition of biospecimens, and researchers are beginning to investigate curation methods that can integrate with existing clinical workflows and leverage electronic health record (EHR) metadata for biospecimen identification and annotation.[6][7]
One of the first automated biospecimen identification systems was Crimson, an application used to identify the discarded blood samples accessioned into the clinical laboratory by querying the laboratory information system (LIS). Specimens, which met predetermined inclusion criteria, were electronically reaccessioned into a deidentified research database that could be accessed by researchers with institutional review board (IRB) approval.[8] While biobanks routinely link health information between specimen and participant postenrollment, such solutions demonstrate how EHR integration and associated metadata can be used for targeted and automated biospecimen selection. However, examples of this framework remain limited, both in the literature and in practice, which typically focus on retrospective specimen identification for long-term biobanking. With the increased digitization of healthcare and modern data architectures that allow for real-time analysis of clinical data, biospecimens can be identified as samples that are processed through the clinical laboratory. This approach offers the benefit of increasing access to specimens of interest, including those with labile analytes, while not disrupting routine clinical workflows.[4]
In this report, we present Prism, a new tool built on open-source technology that can efficiently identify and notify the investigators of biospecimen availability in near real time. We describe the pipeline architecture and our experience with two IRB-approved pilot projects within our department (IRB Protocol IDs: Babesia – 2000023123; Diabetic biomarkers – 2000022266).
Methods
Prism platform architecture
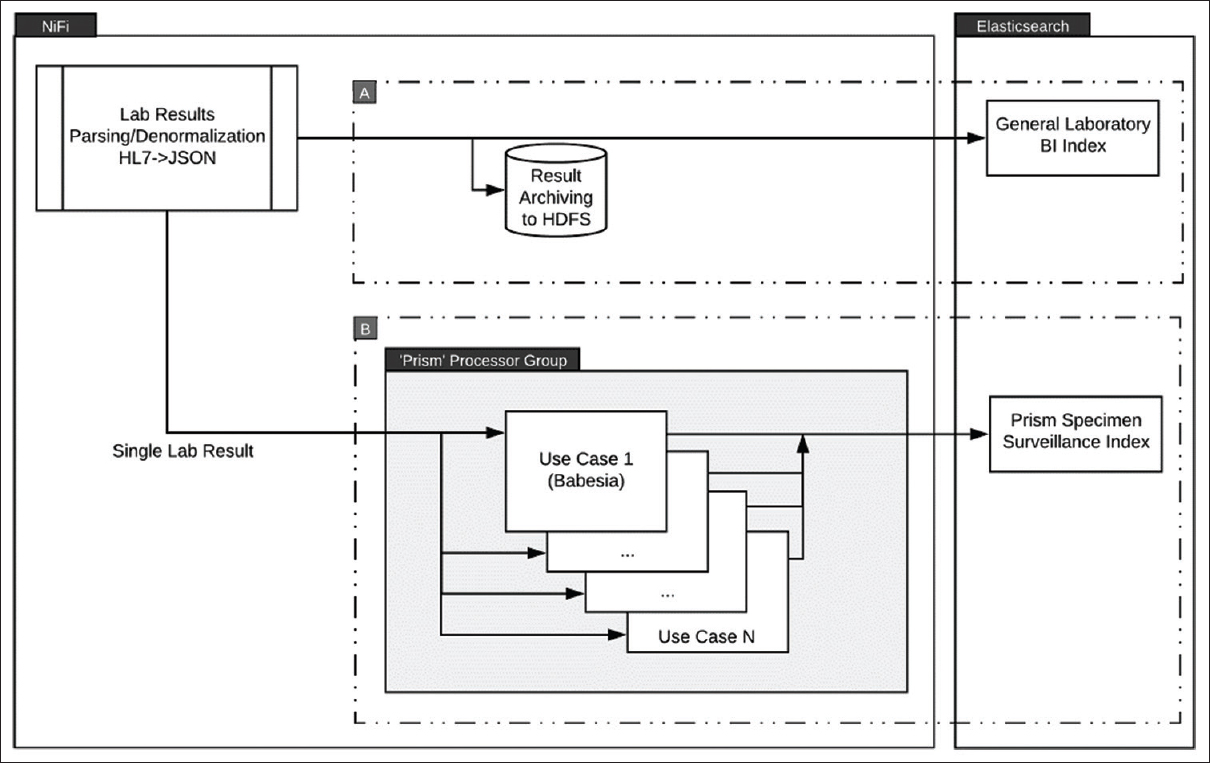
We implemented a real-time pipeline, called Prism, that consists of three key components: real-time data acquisition, stream processing, and end-user alerting, to support case and specimen identification (Fig. 1). Parameterized processors for Apache's open-source dataflow (stream processing) system NiFi were used to filter and identify the clinical specimens based on study-specific inclusion criteria extracted from corresponding laboratory result metadata. Specimens that met inclusion criteria were indexed within Elastic search (Elastic NV; Mountain View, CA, United States). Alerting was done through Watcher (Elastic NV; Mountain View, CA, United States) and secure e-mail, with a reporting dashboard built in Kibana (Elastic NV; Mountain View, CA, United States).
|
This framework was deployed within our organization's computational health care platform, Baikal, which has been previously described (see Supplemental Figure 1 in Additional file 1).[9] The Baikal platform is built on open-source technology and provides a mechanism to manage and analyze high-volume and high-frequency clinical data in real time, including laboratory results.
Throughput assessment
Scalability and computing resource needs for Prism were estimated through the deployment in a standalone workstation environment with a single CPU with six cores (Intel Core i7-6850K CPU @ 3.60GHz) and 256 GiB of memory. Apache NiFi was deployed within a Docker (version 19.03.2, build 6a30dfc; Docker, Inc., San Francisco, CA, United States) container under the Ubuntu (Version: 16.04.6 LTS (Xenial); Canonical Ltd; London, UK) operating system. We ran a modified version of the Prism dataflow using file-based record I/O instead of streaming data from a network interface. Data for the assessment were obtained by randomly selecting data from our production Health Level 7 (HL7) feed, and the data were assembled into three datasets of increasing size. These datasets contained 1 × 105, 1 × 106, and 1 × 107 JavaScript Object Notation-transformed HL7 ORU messages, resulting in 0.75 GB, 7.27 GB, and 72.8 GB of data, respectively. Two series of five trials were performed with the 1 × 106 record data set. In the first series, all five trials were run consecutively. In the second series, Docker was restarted between each trial to assess for any possible performance impacts in long-running containers. Throughput was measured using built-in NiFi monitoring tools to assess record count and throughput.
Results
Babesia specimen identification
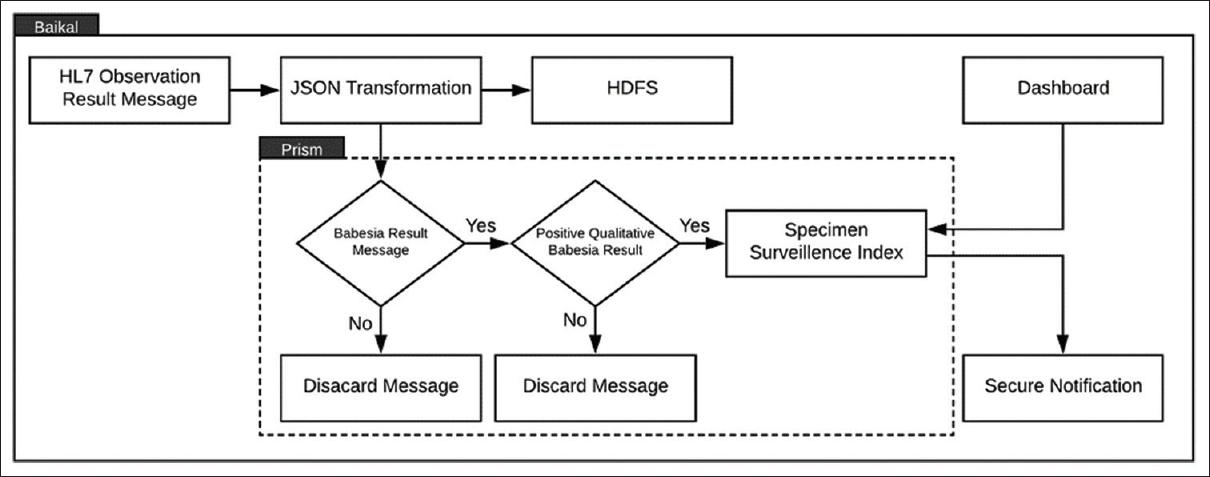
Babesia is a tick-borne hemoprotozoan, which infects human erythrocytes and can be life-threatening for patients who are asplenic, immunocompromised, or elderly. The gold standard for the laboratory diagnosis is microscopic analysis of a peripheral blood smear. For research into the automation of digital microscopic analysis using the computer vision, the researcher needed peripheral blood smears, which were identified as containing Babesia. Incoming HL7 messages corresponding to a Babesia result record with a “Positive” result value were flagged and sent to the Prism index in Elasticsearch (Fig. 2). Researchers were securely notified of all “Positive” Babesia specimens identified every four hours.
|
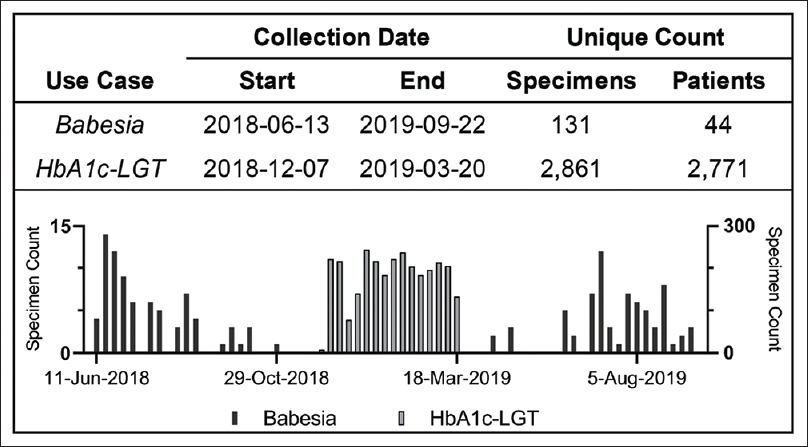
Specimen identification for positive Babesia specimens went live in May 2018. In a collection period of 16 months (June 2018–September 2019), Prism identified 131 unique lavender-top tubes, belonging to 44 unique patients, which were identified as positive for Babesia by manual light microscopy. The collection period for this project was extended beyond the anticipated time requirement as Babesia exhibits a strong seasonal prevalence, and positive specimen rates dropped over the colder months (Fig. 3).
|
Diabetic biomarker specimen identification
The development of type 2 diabetes can be prevented or delayed in prediabetic individuals with lifestyle modifications such as dietary changes or increased physical activity. Accordingly, there is a need to identify the biomarkers to guide preventative interventions.[10][11] To identify possible biomarkers, a researcher at our institution was interested in obtaining blood specimens from patients with and without diabetes, with borderline cases excluded, as a prelude to a larger prospective biomarker study. The deidentified samples would undergo metabolomic analysis by liquid chromatography–mass spectrometry to identify the metabolites that were significantly changed between the two groups as candidate biomarkers.
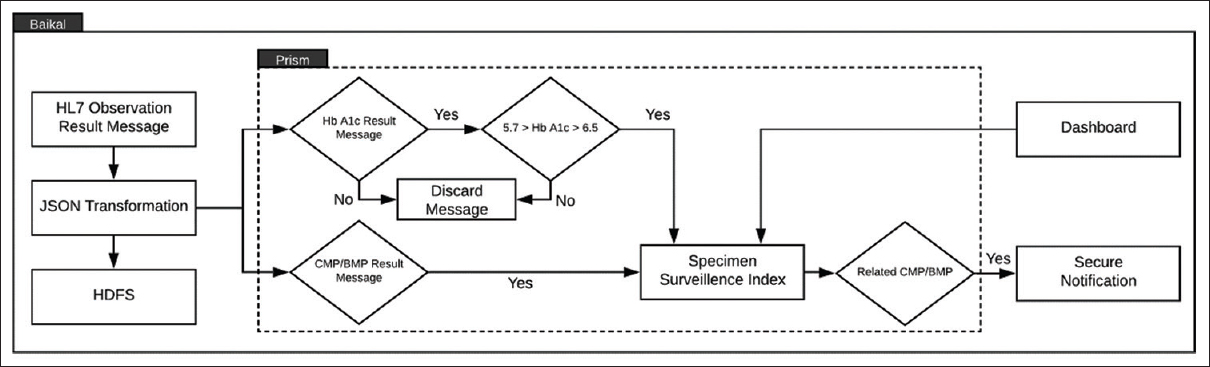
Hemoglobin A1C values < 5.7 and > 6.5 were used to delineate between diabetic and nondiabetic patients, with additional inclusion criteria of outpatient specimen collection and patient age range 18–70 years. Of note, the preferred collection container for Hgb A1c at our institution is a lavender-top tube, which does not contain gel-separation barriers. In an effort to optimize biomarker recovery, plasma from light-green-top (LGT) tubes was requested for this study. Accordingly, LGT tubes were flagged when a paired sample with an Hgb A1c within the appropriate range was found within seven days (Fig. 4). Researchers were securely notified every morning by e-mail of all matching LGT specimens present in the Prism index reported within 24 hours prior.
|
Specimen identification for diabetic biomarker discovery went live in December 2018. In a collection period of four months (December 2018–March 2019), Prism identified 2,861 unique LGT specimens from 2,771 unique patients (Fig. 3).
Throughput assessment
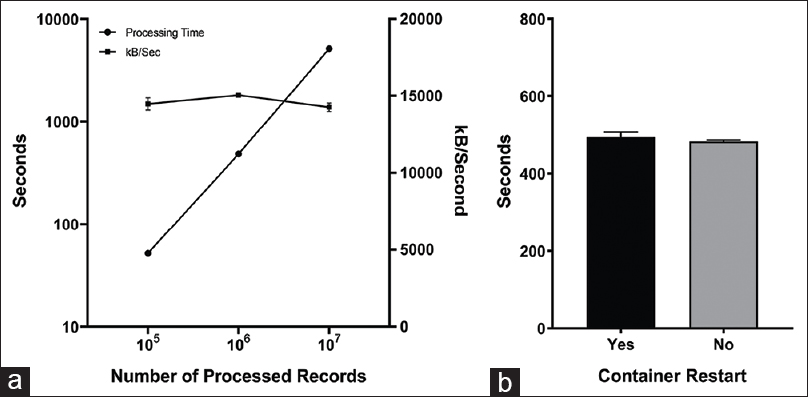
We assessed the processing throughput to ensure the pipeline could scale to large environments and consistently manage high-volume data. Our institution's computational health platform processes approximately 350,000 discrete HL7 ORU messages per day. Accordingly, we evaluated processing time across five trials and observed an average execution time of approximately eight minutes for one million records, which represents slightly less than three days of laboratory result volume. Processing time was observed to be linear over two orders of magnitude in dataset size (Fig. 5a), and the average total execution time to process one million messages differed by 2% between runs with (494 seconds) and without (483 seconds) Docker container restart (Fig. 5b).
|
Discussion
In this report, we describe a novel data analysis pipeline called Prism that can be used to improve the efficiency of biospecimen collection. This workflow has been deployed to identify the biospecimens in near real-time for two biomedical research use cases. We demonstrated that this solution is highly scalable to meet the needs of even large academic centers and reference laboratories. We also found, consistent with our prior work, that virtualization of this workflow within a microservices environment does not introduce a performance penalty.[12]
In 2000, it was estimated that 300 million human biospecimens were preserved in the United States, with a projected 7% annual growth rate.[13] However, researchers continue to report difficulty in obtaining specimens for biomedical research and express underlying concerns in the validity of their results when using specimens subjected to long-term storage conditions.[5] In addition, while many biospecimens are being stored, a large proportion is expected to remain unused, and there is increasing concern that untargeted collection of biospecimens consumes resources that could be better allocated.[14][15][16] Accordingly, as institutions seek to expand biomedical research efforts, particularly in the era of personalized medicine, novel approaches for improving access to high-quality human biospecimens should be evaluated.
The quality of biomedical research is dependent on the integrity of biospecimens and as with clinical testing, analyte recovery is subject to a significant number of preanalytical considerations.[4] While biobanking procedures have seen significant optimization in recent years, poor reproducibility of studies that use biospecimens has been thought to be caused, in part, by the variable quality and inadequate documentation of biospecimen metadata.[5] To this end, biobanks are beginning to emulate testing procedures found in the clinical laboratory to optimize analyte recovery and test reproducibility.[17][18] Tools that can identify the samples accessioned to the clinical laboratory, such as Prism, would align with these efforts by identifying the specimens that have been collected and processed under clinical conditions.
Despite ongoing adoption of clinical procedures in biospecimen science, the collection and processing of labile analytes remain challenging, and some components may require unique processing protocols.[17][19] Proteomic and molecular analytes are particularly sensitive to specimen transport delays, matrix effects, and optimal-storage environments.[20] Accordingly, some components of interest may require sample processing techniques that exist outside routine clinical workflows. In this setting, real-time streaming analytics could also be envisioned to identify patients which match study-specific inclusion criteria to guide targeted subject enrollment and subsequent collection.
In addition to specimen identification and collection, annotation with patient metadata remains an important and challenging facet of contemporary biobanking. Large scale biobanks such as the U.K. Biobank rely on a combination of data sources for curating specimen metadata, including participant enrollment surveys, physical measures (e.g., blood pressure and spirometry), and linkage to digital health information.[21][22] Indeed, while the majority of national biobanking resources capture data from both inpatient and outpatient medical records, there is also interest in capturing data that is not stored in the EHR.[21][23][24] As digital health information continues to expand, health care systems are increasingly working to develop clinically integrated data management tools for the centralization of disparate data resources.[9] Deployment of automated specimen identification tools in these frameworks may facilitate correlation with these data and would align with national efforts to do so.
It should be noted that the use cases described in this report were selected based on the immediate needs among researchers in our department. However, similar open-source tools could be similarly envisioned to integrate with anatomic pathology data and the EHR, to automatically phenotype tissue specimens as they are processed in the laboratory. While the majority of data elements in the clinical laboratory are discrete, identifying tissue specimens in the anatomic pathology laboratory may require technologies such as natural language processing (NLP) to process semistructured and unstructured data, such as those commonly found in pathology reports.[25] While not used for this implementation, custom NiFi processors would allow the users to develop more complex filters and integrate NLP or machine learning-based technology for free text or nested data structures commonly found in anatomic pathology. Similarly, the platform can also be used to identify the patients who may be eligible to consent and enroll in studies, rather than simply for biospecimen collection.
In the era of digital and personalized medicine, novel approaches to increase the efficiency of biospecimen identification will be crucial to accelerate discovery. Modern data architectures as described here can be used to address the fundamental challenges in the procurement of biospecimens in support of biomedical research. Future work will seek to integrate additional data types, including the analysis of unstructured data, to enable more complex case and biospecimen identification.
Acknowledgements
Disclosure
Wade Schulz was an investigator for a research agreement, through Yale University, from the Shenzhen Center for Health Information for work to advance intelligent disease prevention and health promotion. Schulz also collaborates with the National Center for Cardiovascular Diseases in Beijing; is a technical consultant to HugoHealth, a personal health information platform, and co-founder of Refactor Health, an AI-augmented data management platform for healthcare; and is a consultant for Interpace Diagnostics Group, a molecular diagnostics company.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
References
- ↑ van Ommen, G.-J.B.; Törnwall, O.; Bréchot, C. et al. (2015). "BBMRI-ERIC as a Resource for Pharmaceutical and Life Science Industries: The Development of Biobank-Based Expert Centres". European Journal of Human Genetics 23 (7): 893-900. doi:10.1038/ejhg.2014.235. PMC PMC4463510. PMID 25407005. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4463510.
- ↑ Langhof, H.; Kahrass, H.; Illig, T. et al. (2018). "Current Practices for Access, Compensation, and Prioritization in Biobanks. Results From an Interview Study". European Journal of Human Genetics 26 (11): 1572–1581. doi:10.1038/s41431-018-0228-x. PMC PMC6189200. PMID 30089824. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6189200.
- ↑ Shabihkhani, M.; Lucey, G.M.; Wei, B. et al. (2014). "The Procurement, Storage, and Quality Assurance of Frozen Blood and Tissue Biospecimens in Pathology, Biorepository, and Biobank Settings". Clinical Biochemistry 47 (4–5): 258-66. doi:10.1016/j.clinbiochem.2014.01.002. PMC PMC3982909. PMID 24424103. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3982909.
- ↑ 4.0 4.1 4.2 Ellervik, C.; Vaught, J. (2015). "Preanalytical Variables Affecting the Integrity of Human Biospecimens in Biobanking". Clinical Chemistry 61 (7): 914-34. doi:10.1373/clinchem.2014.228783. PMID 25979952.
- ↑ 5.0 5.1 5.2 Massett, H.A.; Atkinson, N.L.; Weber, D. et al. (2011). "Assessing the Need for a Standardized Cancer HUman Biobank (caHUB): Findings From a National Survey With Cancer Researchers". Journal of the National Cancer Institute Monographs 2011 (42): 8–15. doi:10.1093/jncimonographs/lgr007. PMID 21672890.
- ↑ Moore, H,M.; Jelly, A.; McShane, L.M. et al. (2013). "Biospecimen Reporting for Improved Study Quality (BRISQ)". Transfusion 53 (7): e1. doi:10.1111/trf.12281. PMID 23844646.
- ↑ Simeon-Dubach, D.; Burt, A.D.; Hall, P.A. (2012). "Quality Really Matters: The Need to Improve Specimen Quality in Biomedical Research". Journal of Pathology 228 (4): 431–3. doi:10.1002/path.4117. PMID 23023660.
- ↑ Murphy, S.; Churchill, S.; Bry, L. et al. (2009). "Instrumenting the Health Care Enterprise for Discovery Research in the Genomic Era". Genome Research 19 (9): 1675–81. doi:10.1101/gr.094615.109. PMC PMC2752136. PMID 19602638. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2752136.
- ↑ 9.0 9.1 McPadden, J.; Durant, T.J.S.; Bunch, D.R. et al. (2019). "Health Care and Precision Medicine Research: Analysis of a Scalable Data Science Platform". Journal of Medical Internet Research 21 (4): e13043. doi:10.2196/13043. PMC PMC6477571. PMID 30964441. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6477571.
- ↑ Wang-Sattler, R.; Yu, Z.; Herder, C. et al. (2012). "Novel Biomarkers for Pre-Diabetes Identified by Metabolomics". Molecular Systems Biology 8: 615. doi:10.1038/msb.2012.43. PMC PMC3472689. PMID 23010998. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3472689.
- ↑ Guasch-Ferré, M.; Hruby, A.; Toledo, E. et al. (2016). "Metabolomics in Prediabetes and Diabetes: A Systematic Review and Meta-analysis". Diabetes Care 39 (5): 833-46. doi:10.2337/dc15-2251. PMC PMC4839172. PMID 27208380. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4839172.
- ↑ Schulz, W.L.; Durant, T.J. S.; Siddon, A.J. et al. (2016). "Use of Application Containers and Workflows for Genomic Data Analysis". Journal of Pathology Informatics 7: 53. doi:10.4103/2153-3539.197197. PMC PMC5248400. PMID 28163975. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5248400.
- ↑ Eiseman, E.; Haga, S.B. (1999). Handbook of Human Tissue Sources: A National Resource of Human Tissue Samples. RAND Corporation. ISBN 0833027662.
- ↑ Gee, S.; Georghiou, L.; Oliver, R. et al. (June 2013). "Financing UK biobanks: Rationale for a National Biobanking Research Infrastructure". Manchester Business School, Faculty of Humanities. https://www.escholar.manchester.ac.uk/uk-ac-man-scw:199203.
- ↑ Simeon-Dubach, D.; Watson, P. (2014). "Biobanking 3.0: Evidence Based and Customer Focused Biobanking". Clinical Biochemistry 47 (4–5): 300-8. doi:10.1016/j.clinbiochem.2013.12.018. PMID 24406300.
- ↑ Simeon-Dubach, D.; Henderson, M.K. (2014). "Sustainability in Biobanking". Biopreservation and Biobanking 12 (5): 287–91. doi:10.1089/bio.2014.1251. PMID 25314050.
- ↑ 17.0 17.1 Vaught, J. (2016). "Biobanking Comes of Age: The Transition to Biospecimen Science". Annual Review of Pharmacology and Toxicology 56: 211–28. doi:10.1146/annurev-pharmtox-010715-103246. PMID 26514206.
- ↑ Betsou, F.; Barnes, R.; Burke, T. et al. (2009). "Human Biospecimen Research: Experimental Protocol and Quality Control Tools". Cancer Epidemiology, Biomarkers & Prevention 18 (4): 1017-25. doi:10.1158/1055-9965.EPI-08-1231. PMID 19336543.
- ↑ Vaught, J.; Rogers, J.; Myers, K. et al. (2011). "An NCI Perspective on Creating Sustainable Biospecimen Resources". Journal of the National Cancer Institute Monographs 2011 (42): 1–7. doi:10.1093/jncimonographs/lgr006. PMID 21672889.
- ↑ El Messaoudi, S.; Rolet, F.; Mouliere, F. et al. (2013). "Circulating Cell Free DNA: Preanalytical Considerations". Clinica Chimica Acta 424: 222–30. doi:10.1016/j.cca.2013.05.022. PMID 23727028.
- ↑ 21.0 21.1 All of Us Research Program Investigators; Denny, J.C.; Rutter, J.L. et al. (2019). "The "All of Us" Research Program". New England Journal of Medicine 381 (7): 668-676. doi:10.1056/NEJMsr1809937. PMID 31412182.
- ↑ Bycroft, C.; Freeman, C.; Petkova, D. et al. (2018). "The UK Biobank Resource With Deep Phenotyping and Genomic Data". Nature 562 (7726): 203–9. doi:10.1038/s41586-018-0579-z. PMC PMC6786975. PMID 30305743. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6786975.
- ↑ Chen, Z.; Chen, J.; Collins, R. et al. (2011). "China Kadoorie Biobank of 0.5 Million People: Survey Methods, Baseline Characteristics and Long-Term Follow-Up". International Journal of Epidemiology 40 (6): 1652-66. doi:10.1093/ije/dyr120. PMC PMC3235021. PMID 22158673. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3235021.
- ↑ Gaziano, J.M.; Concato, J.; Brophy, M. et al. (2016). "Million Veteran Program: A Mega-Biobank to Study Genetic Influences on Health and Disease". Journal of Clinical Epidemiology 70: 214–23. doi:10.1016/j.jclinepi.2015.09.016. PMID 26441289.
- ↑ Buckley, J.M.; Coopey, S.B.; Sharko, J. et al. (2012). "The Feasibility of Using Natural Language Processing to Extract Clinical Information From Breast Pathology Reports". Journal of Pathology Informatics 3: 23. doi:10.4103/2153-3539.97788. PMC PMC3424662. PMID 22934236. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3424662.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. Grammar was cleaned up for smoother reading. In some cases important information was missing from the references, and that information was added. As the time of loading this article, the link to Additional file 1 was broken on the original site. Unfortunately, it can not be included here.