Journal:Laboratory testing for coronavirus disease (COVID-19) in suspected human cases
| Full article title | Laboratory testing for coronavirus disease (COVID-19) in suspected human cases: Interim guidance, 19 March 2020 |
|---|---|
| Author(s) |
World Health Organization; Leitmeyer, Katrin; Zambon, Maria; Drosten, Christian; Koopmans, Marion; Poon, Leo; Gao, George; Nahapetyan, Karen; Inbanathan, Francis; Pereyaslov, Dmitriy; Uhlenhaut, Christine; Grabovac, Varja; Vandemaele, Katelijn; Samaan, Magdi; Fuster, Christian; Zhang, Wenqing; Stevens, Lisa; Oxenford, Chris; Cognat, Sebastian; Kojima, Kazunobu; Dolea, Carmen; Brown, Caroline; Barnadas, Céline; Van Kerkhove, Maria; Carter, Lisa; Perkins, Mark D.; von Eije, Karin |
| Author affiliation(s) |
European Center for Disease Control, Public Health England, Charité - Universitätsmedizin Berlin, Erasmus University Medical Center, Hong Kong University, Chinese Center for Disease Control and Prevention, World Health Organization |
| Primary contact | Email: WHElab at who dot int |
| Year published | 2020 |
| Volume and issue | 2020.5 |
| Page(s) | 1–7 |
| Distribution license | Attribution-NonCommercial-ShareAlike 3.0 IGO |
| Website | https://apps.who.int/iris/handle/10665/331501 |
| Download | https://apps.who.int/iris/bitstream/handle/10665/331501/WHO-COVID-19-laboratory-2020.5-eng.pdf (PDF) |
Background
|
|
This LIMSwiki article is out-of-date as of September 2020. The WHO document Diagnostic testing for SARS-CoV-2: Interim guidance, 11 September 2020 supersedes this article's guidance. Please consult the latest WHO guidance for more information. |
This document provides interim guidance to laboratories and stakeholders involved in COVID-19 virus laboratory testing of patients. It is based in part on the interim guidance on laboratory testing for Middle East respiratory syndrome (MERS) coronavirus.[1][2][3][4][5][6] Information on human infection with the COVID-19 virus is evolving and the World Health Organization (WHO) continues to monitor developments and revise recommendations as necessary. This document will be revised as new information becomes available. Feedback is welcome and can be sent to WHElab@who.int.
The virus has now been named SARS-CoV-2 by the International Committee of Taxonomy of Viruses (ICTV)(2).[7] This virus can cause the disease named coronavirus disease 2019 (COVID-19). WHO refers to the virus as COVID-19 virus in its current documentation.
Laboratory testing guiding pricinciples for patients who meet the suspect case definition
The decision to test should be based on clinical and epidemiological factors and linked to an assessment of the likelihood of infection. Polymerase chain reaction (PCR) testing of asymptomatic or mildly symptomatic contacts can be considered in the assessment of individuals who have had contact with a COVID-19 case. Screening protocols should be adapted to the local situation. The case definitions are being regularly reviewed and updated as new information becomes available. For the WHO suspected case definition see Global surveillance for COVID-19 caused by human infection with COVID-19 virus.[8]
Rapid collection and testing of appropriate specimens from patients meeting the suspected case definition for COVID-19 is a priority for clinical management and outbreak control and should be guided by a laboratory expert. Suspected cases should be screened for the virus with nucleic acid amplification tests (NAAT), such as Reverse transcription polymerase chain reaction (RT-PCR).
If testing for COVID-19 is not yet available nationally, specimens should be referred. A list of WHO reference laboratories providing confirmatory testing for COVID-19 and shipment instructions can be found at the bottom of the WHO's national laboratories technical guidance page.
If case management requires it, patients should be tested for other respiratory pathogens using routine laboratory procedures, as recommended in local management guidelines for community-acquired pneumonia. Additional testing should not delay testing for COVID-19. As co-infections can occur, all patients who meet the suspected case definition should be tested for COVID-19 virus regardless of whether another respiratory pathogen is found.
In an early study in Wuhan, the mean incubation period for COVID-19 was 5.2 days among 425 cases, though it varies widely among individuals.[9][10] Virus shedding patterns are not yet well understood, and further investigations are needed to better understand the timing, compartmentalization, and quantity of viral shedding to inform optimal specimen collection. Although respiratory samples have the greatest yield, the virus can be detected in other specimens, including stool and blood.[11][12] Local guidelines on informed consent should be followed for specimen collection, testing, and potentially any future research.
Specimen collection and shipment
Safety procedures during specimen collection
Ensure that adequate standard operating procedures (SOPs) are in use and that staff are trained for appropriate specimen collection, storage, packaging, and transport. All specimens collected for laboratory investigations should be regarded as potentially infectious.
Ensure that health care workers who collect specimens adhere rigorously to infection prevention and control guidelines. Specific WHO interim guidance has been published.[13] Also see Box 1, below.
|
Specimens to be collected
At a minimum, respiratory material should be collected. This includes:
- upper respiratory specimens, using a nasopharyngeal and oropharyngeal swab or wash in ambulatory patients; and/or,
- lower respiratory specimens such as sputum (if produced), or use endotracheal aspiration or bronchoalveolar lavage procedures in patients with more severe respiratory disease (note the high risk of aerosolization; adhere strictly to infection prevention and control procedures).
Additional clinical specimens may be collected. For example, COVID-19 virus has been detected in blood and stool, as had the coronaviruses responsible for SARS (severe acute respiratory syndrome) and MERS (Middle East respiratory syndrome).[11][12][15][16][17] The duration and frequency of shedding of COVID-19 virus in stool and potentially in urine is unknown. In case of patients who are deceased, consider autopsy material, including lung tissue. In surviving patients, paired serum (acute and convalescent) can be useful to retrospectively define cases as serological assays become available.
Further recommendations on materials to collect, including the testing of asymptomatic individuals, can be found in Table 1.
| |||||||||||||||||||||||
Packaging and shipment of clinical specimens
Specimens for virus detection should reach the laboratory as soon as possible after collection. Correct handling of specimens during transportation is essential. Specimens that can be delivered promptly to the laboratory can be stored and shipped at 2-8°C. When there is likely to be a delay in specimens reaching the laboratory, the use of viral transport medium (VTM) is strongly recommended. Specimens may be frozen to -20°C or ideally -70°C and shipped on dry ice if further delays are expected (see Table 2). It is important to avoid repeated freezing and thawing of specimens.
| ||||||||||||||||||||||||||||||||||||||||||||||||
Transport of specimens within national borders should comply with applicable national regulations. International transport of specimens potentially containing the COVID-19 virus should follow the United Nations Model Regulations, and any other applicable regulations depending on the mode of transport being used. More information may be found in the WHO's Guidance on Regulations for the Transport of Infectious Substances 2019-2020[20][21] and WHO interim guidance for laboratory biosafety related to coronavirus disease.[13]
Ensure good communication with the laboratory and provide needed information
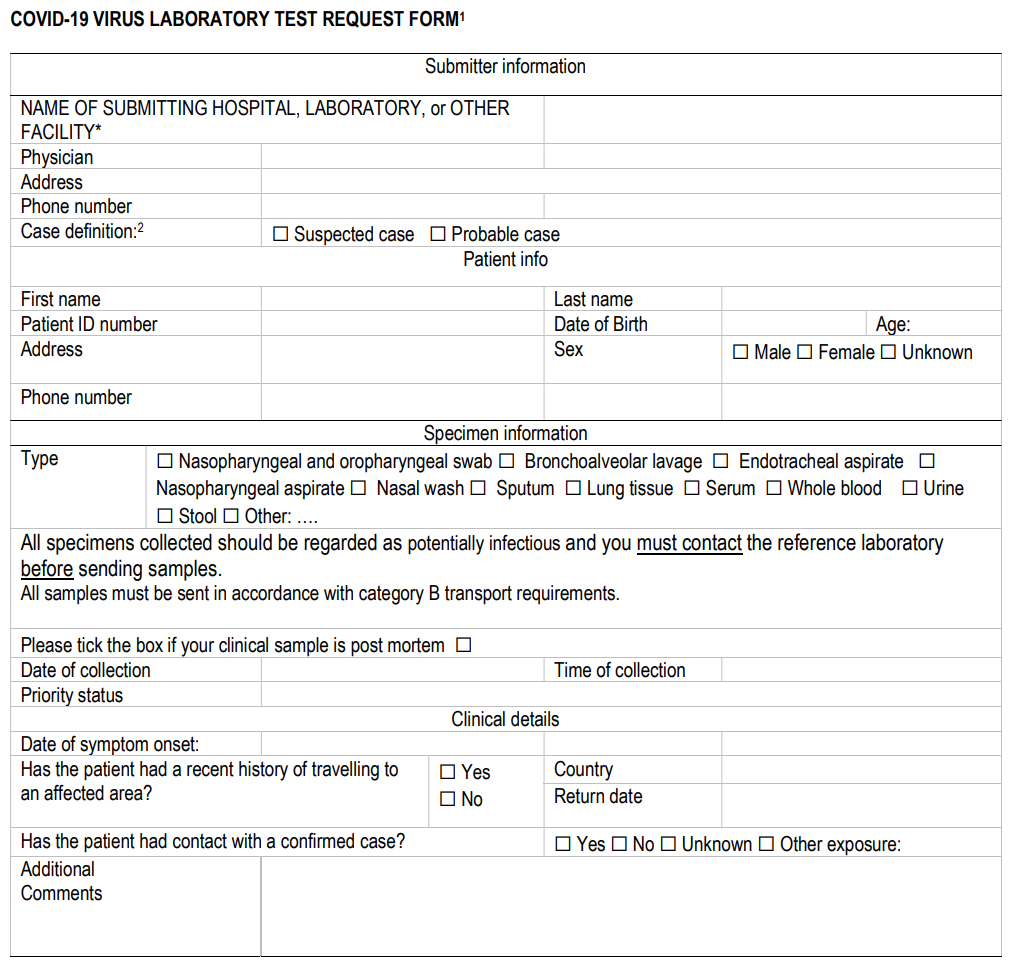
Alerting the laboratory before sending specimens encourages their proper and timely processing and reporting. Specimens should be correctly labelled and accompanied by a diagnostic request form (template provided in Annex I).
Laboratory testing for COVID-19 virus
Laboratories undertaking testing for COVID-19 virus should adhere strictly to appropriate biosafety practices.
Nucleic acid amplification tests (NAAT) for COVID-19 virus
Routine confirmation of cases of COVID-19 is based on detection of unique sequences of virus RNA by NAAT using methods such as real-time reverse-transcription polymerase chain reaction (rRT-PCR), with confirmation by nucleic acid sequencing when necessary. The viral genes targeted so far include the N, E, S, and RdRP genes. Examples of protocols used may be found here. RNA extraction should be done in a biosafety cabinet in a BSL-2 or equivalent facility. Heat treatment of samples before RNA extraction is not recommended.
Laboratory confirmation of cases by NAAT in areas with no known COVID-19 virus circulation
To consider a case as laboratory-confirmed by NAAT in an area with no COVID-19 virus circulation, one of the following conditions need to be met:
- Testing demonstrates a positive NAAT result for at least two different targets on the COVID-19 virus genome, of which at least one target is preferably specific for COVID-19 virus using a validated assay. (As at present no other SARS-like coronaviruses are circulating in the human population, it can be debated whether it must be COVID-19 or SARS-like coronavirus-specific.); OR,
- Testing demonstrates one positive NAAT result for the presence of betacoronavirus, with the COVID-19 virus further identified by sequencing a partial or whole genome of the virus, as long as the sequence target is larger or different from the amplicon probed in the NAAT assay used.
When there are discordant results, the patient should be resampled and, if appropriate, sequencing of the virus from the original specimen or of an amplicon generated from an appropriate NAAT assay, different from the NAAT assay initially used, should be obtained to provide a reliable test result. Laboratories are urged to seek confirmation of any surprising results in an international reference laboratory.
Laboratory-confirmed case by NAAT in areas with established COVID-19 virus circulation
In areas where COVID-19 virus is widely spread, a simpler algorithm might be adopted in which, for example, screening by rRT-PCR of a single discriminatory target is considered sufficient.
One or more negative results do not rule out the possibility of COVID-19 virus infection. A number of factors could lead to a negative result in an infected individual, including:
- The specimen quality is poor, containing little patient material (as a control, consider determining whether there is adequate human DNA in the sample by including a human target in the PCR testing).
- The specimen was collected late or very early in the infection.
- The specimen was not handled and shipped appropriately.
- Technical aspects inherent in the test, e.g., virus mutation or PCR inhibition, were at fault.
If a negative result is obtained from a patient with a high index of suspicion for COVID-19 virus infection—particularly when only upper respiratory tract specimens were collected—additional specimens, including from the lower respiratory tract if possible, should be collected and tested.
Each NAAT run should include both external and internal controls, and laboratories are encouraged to participate in external quality assessment schemes when they become available. Laboratories that order their own primers and probes are also recommended to perform entry testing/validation on functionality and potential contaminants.
Serological testing
Serological surveys can aid investigation of an ongoing outbreak and retrospective assessment of the attack rate or extent of an outbreak. In cases where NAAT assays are negative and there is a strong epidemiological link to COVID-19 infection, paired serum samples (in the acute and convalescent phase) could support diagnosis once validated serology tests are available. Serum samples can be stored for these purposes.
Cross reactivity to other coronaviruses can be challenging[22], but commercial and non-commercial serological tests are currently under development. Some studies with COVID-19 serological data on clinical samples have been published.[23][24]
Viral sequencing
In addition to providing confirmation of the presence of the virus, regular sequencing of a percentage of specimens from clinical cases can be useful to monitor for viral genome mutations that might affect the performance of medical countermeasures, including diagnostic tests. Virus whole genome sequencing can also inform molecular epidemiology studies. Many public-access databases for deposition of genetic sequence data are available, including GISAID, which is intended to protect the rights of the submitting party.[25]
Viral culture
Virus isolation is not recommended as a routine diagnostic procedure.
Reporting of cases and test results
Laboratories should follow national reporting requirements. In general, all test results, positive or negative, should be immediately reported to national authorities. States Parties to the International Health Regulations are reminded of their obligations to share with WHO relevant public health information for events for which they notified WHO, using the decision instrument in Annex 1 of the IHR (2005).[26]
Research toward improved detection of COVID-19 virus
Many aspects of the virus and disease are still not understood. A better understanding will be needed to provide improved guidance, particularly in regard to viral dynamics. For example:
- optimal timing and type of clinical material to sample for molecular testing
- dynamic of immunological response
- disease severity in various populations, e.g., by age
- the relationship between viral concentration and disease severity
- the duration of shedding and its relation to the clinical picture (e.g., Does clinical recovery occurs with viral clearing? Does shedding persists despite clinical improvement?)
- development and validation of useful serological assays
- comparative studies of available molecular and serological assays
- optimal percentage of positive cases that requires sequencing to monitor mutations that might affect the performance of molecular test
WHO encourages the sharing of data to better understand and thus manage the COVID-19 outbreak, and to develop countermeasures.
Annex I
|
Acknowledgements
The following people contributed to the drafting of the evolving versions of this guidance document: Katrin Leitmeyer, European Center for Disease Control; Maria Zambon, Public Health England, UK; Christian Drosten, Charité - Universitätsmedizin Berlin, Germany; Marion Koopmans, Erasmus MC, Rotterdam, The Netherlands; Leo Poon, Hong Kong University, China, Hong Kong SAR; George Gao, Chinese CDC, China. From WHO: Karen Nahapetyan, Francis Inbanathan, Dmitriy Pereyaslov, Christine Uhlenhaut, Varja Grabovac, Katelijn Vandemaele, Magdi Samaan, Christian Fuster, Wenqing Zhang, Lisa Stevens, Chris Oxenford, Sebastian Cognat, Kazunobu Kojima, Carmen Dolea, Caroline Brown, Céline Barnadas, Maria Van Kerkhove, Lisa Carter, Mark D Perkins and Karin von Eije.
WHO continues to monitor the situation closely for any changes that may affect this interim guidance. Should any factors change, WHO will issue a further update. Otherwise, this interim guidance document will expire two years after the date of publication.
References
- ↑ World Health Organization (January 2018). "Laboratory testing for Middle East Respiratory Syndrome coronavirus, interim guidance (revised), January 2018". WHO/MERS/LAB/15.1/Rev1/2018. World Health Organization. https://www.who.int/csr/disease/coronavirus_infections/mers-laboratory-testing/en/.
- ↑ World Health Organization (2018). Managing epidemics: Key facts about major deadly diseases. World Health Organization. pp. 257. ISBN 9789241565530. https://apps.who.int/iris/handle/10665/272442.
- ↑ World Health Organization (2011). WHO Global Influenza Surveillance Network: Manual for the laboratory diagnosis and virological surveillance of influenza. World Health Organization. pp. 140. ISBN 9789241548090. https://apps.who.int/iris/handle/10665/44518.
- ↑ 4.0 4.1 World Health Organization (October 2018). "Protocol to investigate non-seasonal influenza and other emerging acute respiratory diseases". WHO/WHE/IHM/GIP/2018.2. World Health Organization. https://www.who.int/influenza/resources/publications/outbreak_investigation_protocol/en/.
- ↑ World Health Organization (October 1999). "WHO Recommended Surveillance Standards, Second Edition". WHO/CDS/CSR/ISR/99.2. World Health Organization. https://apps.who.int/iris/handle/10665/65517.
- ↑ World Health Organization (2000). "Guidelines for the Collection of Clinical Specimens During Field Investigation of Outbreaks". WHO/CDS/CSR/EDC/2000.4. World Health Organization. https://apps.who.int/iris/handle/10665/66348.
- ↑ Gorbalenya, A.E.; Baker, S.C.; Baric, R.S. et al. (2020). "Severe acute respiratory syndrome-related coronavirus: The species and its viruses – A statement of the Coronavirus Study Group". bioRxiv: 937862. doi:10.1101/2020.02.07.937862.
- ↑ 8.0 8.1 World Health Organization (20 March 2020). "Global surveillance for COVID-19 caused by human infection with COVID-19 virus: interim guidance, 20 March 2020". WHO/2019-nCoV/SurveillanceGuidance/2020.6. World Health Organization. https://apps.who.int/iris/handle/10665/331506.
- ↑ Li, Q.; Guan, X.; Wu, P. et al. (2020). "Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus–Infected Pneumonia". The New England Journal of Medicine. doi:10.1056/NEJMoa2001316. PMID 31995857.
- ↑ Guan, W.; Ni, Z.; Hu, Y. et al. (2020). "Clinical Characteristics of Coronavirus Disease 2019 in China". The New England Journal of Medicine. doi:10.1056/NEJMoa2002032. PMID 32109013.
- ↑ 11.0 11.1 Linton, N.M.; Kobayashi, T.; Yang, Y. et al. (2020). "Incubation Period and Other Epidemiological Characteristics of 2019 Novel Coronavirus Infections with Right Truncation: A Statistical Analysis of Publicly Available Case Data". Journal of Clinical Medicine 9 (2): 538. doi:10.3390/jcm9020538. PMC PMC7074197. PMID 32079150. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7074197.
- ↑ 12.0 12.1 Zhang, W.; Du, R.-H.; Li, B. et al. (2020). "Molecular and serological investigation of 2019-nCoV infected patients: Implication of multiple shedding routes". Emerging Microbes & Infections 9 (1): 386–89. doi:10.1080/22221751.2020.1729071. PMC PMC7048229. PMID 32065057. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7048229.
- ↑ 13.0 13.1 World Health Organization (19 March 2020). "Laboratory biosafety guidance related to coronavirus disease (COVID-19): Interim guidance, 19 March 2020". WHO/WPE/GIH/2020.2. World Health Organization. https://apps.who.int/iris/handle/10665/331500.
- ↑ Kojima, K. (7 February 2019). "WHO Laboratory Biosafety Manual, Revision Update". SlideShare. World Health Organization. https://www.slideshare.net/MicrobiologySection/who-laboratory-biosafety-manual-revision-update.
- ↑ Shi, X.; Gong, E.; Gao, D. et al. (2005). "Severe acute respiratory syndrome associated coronavirus is detected in intestinal tissues of fatal cases". American Journal of Gastroenterology 100 (1): 169–76. doi:10.1111/j.1572-0241.2005.40377.x. PMID 15654797.
- ↑ Zhou, J.; Li, C.; Zhao, G. et al. (2017). "Human intestinal tract serves as an alternative infection route for Middle East respiratory syndrome coronavirus". Science Advances 3 (11): eaao4966. doi:10.1126/sciadv.aao4966. PMC PMC5687858. PMID 29152574. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5687858.
- ↑ Ding, Y.; He, L.; Zhang, Q. et al. (2004). "Organ distribution of severe acute respiratory syndrome (SARS) associated coronavirus (SARS-CoV) in SARS patients: Implications for pathogenesis and virus transmission pathways". Journal of Pathology 203 (2): 622-30. doi:10.1002/path.1560. PMID 15141376.
- ↑ World Health Organization (October 2018). "Protocol to investigate non-seasonal influenza and other emerging acute respiratory diseases". WHO/WHE/IHM/GIP/2018.2. World Health Organization.
- ↑ Druce, J.; Garcia, K. Tran, T. et al. (2012). "Evaluation of swabs, transport media, and specimen transport conditions for optimal detection of viruses by PCR". Journal of Clinical Microbiology 50 (3): 1064–5. doi:10.1128/JCM.06551-11. PMC PMC3295134. PMID 22205810. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3295134.
- ↑ World Health Organization (January 2019). "Guidance on Regulations for the Transport of Infectious Substances 2019-2020". WHO/WHE/CPI/2019.20. World Health Organization. https://www.who.int/ihr/publications/WHO-WHE-CPI-2019.20/en/.
- ↑ World Health Organization (2018). "Guidance to Minimize Risks for Facilities Collecting, Handling or Storing Materials Potentially Infectious for Polioviruses". WHO/POLIO/18.03. World Health Organization. https://apps.who.int/iris/handle/10665/276199.
- ↑ Meyer, B.; Drosten, C.; Müller, M.A. (2014). "Serological assays for emerging coronaviruses: Challenges and pitfalls". Virus Research 194: 175–83. doi:10.1016/j.virusres.2014.03.018. PMID 24670324.
- ↑ Bai, S.L.; Wang, J.Y.; Zhou, Y.Q. et al. (2020). "Analysis of the first cluster of cases in a family of novel coronavirus pneumonia in Gansu Province". Chinese Journal of Preventive Medicine 54: E005. doi:10.3760/cma.j.issn.0253-9624.2020.0005. PMID 32064855.
- ↑ Xiao, S.Y.; Wu, Y.; Liu, H. (2020). "Evolving status of the 2019 novel coronavirus infection: Proposal of conventional serologic assays for disease diagnosis and infection monitoring". Journal of Medical Virology 92 (5): 464–7. doi:10.1002/jmv.25702. PMID 32031264.
- ↑ "GISAID". Freunde von GISAID e.V. https://www.gisaid.org/. Retrieved 19 February 2020.
- ↑ World Health Organization (2016). International Health Regulations (2005) (3rd ed.). World Health Organization. pp. 84. ISBN 9789241580496. https://www.who.int/ihr/publications/9789241580496/en/.
Notes
This presentation is faithful to the original, with some changes to presentation, grammar, and punctuation. In some cases important information was missing from the references, and that information was added. Their citation #8 referencing the Laboratory Biosafety Manual appears misplaced in the original document; it is appended in where it presumably should go in this version. Their citation #9 is placed where #8 was in the original, as it makes the most sense. The original lists a citation #13, but no in-line citation could be found in the text body; as such, citation #13 is omitted from this version. The original lists a citation #15, but it appears that document no longer exists and was likely superceded by citation #16; as such, citation #15 is omitted from this version. In cases where the document names have changed recently, the correct document name was substituted. An incorrect citation number (22) is used in the packaging and shipping section; presumably citations #20 and #21 were intended, and they were added in place of #22 for this version. A citation concerning the fourth edition of WHO's Laboratory Biosafety Manual was added for this version. Per the WHO's licensing agreement, this reproduction acknowledges the World Health Organization as the source, licensed under the CC BY-NC-SA 3.0 IGO license (see the infobox at top for full details).