Difference between revisions of "Journal:Rapid development of entity-based data models for bioinformatics with persistence object-oriented design and structured interfaces"
Shawndouglas (talk | contribs) (Saving and adding more.) |
Shawndouglas (talk | contribs) (Saving and adding more.) |
||
| Line 41: | Line 41: | ||
==Implementation== | ==Implementation== | ||
===Framework=== | |||
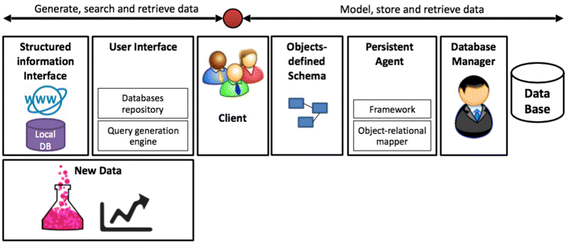
Here, a simple framework for rapid development of specialized databases is proposed. This framework integrates several database technologies into a unified platform which allows the user to search and retrieve data from existing sources, incorporate established data with newly discovered information in a single data model, define object-based data architecture and persist it to memory for future inquiries. The framework consists of three main data streams: existing data sources to the user (search and retrieve data), new information sources such as experiments and data analytics to the user (generate data), as well as the individual user to a self-curated specialized data-base (model, store and retrieve data) (Fig. 1). The platform enables the user to search and retrieve data from local databases using structured information interfaces, and from online databases using structured URL interfaces. The user interface consists of a database repository which defines the differently structured accesses to existing resources, and a query generation engine which provides advanced searching mechanisms such as field-oriented search and the use of logical relations among search terms. The user designs their own data architecture using object-oriented programming (OOP), within which new data and the database-derived information are encapsulated to class-defined entities, an action which forms the database schema. Entities are persisted to memory with a persistent agent (PA). The PA uses an object relational mapper to map the object-based schema to a set of interconnected tables which resemble a typical relational database. This will allow the user to query the database with SQL-like queries. The relational model is transferred to a database manager that stores the data in memory. | |||
[[File:Fig1 Tsur BioDataMining2017 10.gif|567px]] | |||
{{clear}} | |||
{| | |||
| STYLE="vertical-align:top;"| | |||
{| border="0" cellpadding="5" cellspacing="0" width="567px" | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;"| <blockquote>'''Figure 1.''' Framework schematics. Our framework is composed of structured information interfaces, used to search and fetch information from local and online repositories, incorporation of user generated new data, query generation interface and data source selection, a persistent agent which persists user-defined object schema to memory and finally, a database manager. This framework supports generation, search and retrieval of data, as well as modeling, storing and searching persisted schema.</blockquote> | |||
|- | |||
|} | |||
|} | |||
This framework can be implemented in various ways using different programming languages and libraries providers. For example, Southern and colleagues developed a Java API, which maps entities to NCBI's PubChem schema and provides wrapper functions for calling NCBI eUtilities and PubChem web services.<ref name="SouthernAJava11">{{cite journal |title=A Java API for working with PubChem datasets |journal=Bioinformatics |author=Southern, M.R.; Griffin, P.R. |volume=27 |issue=5 |pages=741—2 |year=2011 |doi=10.1093/bioinformatics/btq715 |pmid=21216779 |pmc=PMC3105478}}</ref> Another implementation is the BioPython project, which provides modules to access NCBI's databases from within Python.<ref name="CockBiopython09">{{cite journal |title=Biopython: Freely available Python tools for computational molecular biology and bioinformatics |journal=Bioinformatics |author=Cock, P.J.; Antao, T.; Chang, J.T. et al. |volume=25 |issue=11 |pages=1422-3 |year=2009 |doi=10.1093/bioinformatics/btp163 |pmid=19304878 |pmc=PMC2682512}}</ref> Object persistence can be achieved both in Python using its standard library, which support a family of hash-based file formats and objects serialization, and in JAVA using the Java Persistence API (JPA). Most JPA persistence providers offer the option to automatically create the database schema based on metadata. Popular implementations of JPA are Hibernate, EclipseLink and Apache OpenJPA. Common database managers include Apache Derby, Firebird, SQLite, and HSQL. | |||
===Software description=== | |||
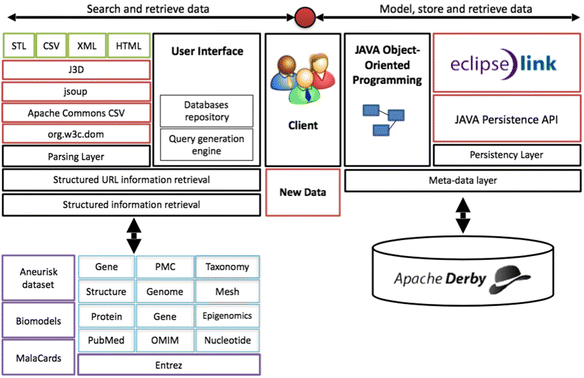
The proposed framework for the development of specialized databases can be implemented using different resources. Here, various open-source and free resources were utilized for implementation. Java was chosen as the development environment, EclipseLink as the JPA provider and Apache Derby as the database manager. Java was used to create interfaces to online databases such as MalaCards, Biomodels and NCBI's databases. The structured information interface was used to derive data from local repositories such as Aneurisk, which was downloaded from the Aneurisk web dataset. By integrating these tools with a series of data parsers, a powerful framework for the curation of specialized databases is provided, which would incorporate new data and database-derived information into a user-defined database architecture. A schematic of the implementation is presented in Fig. 2. | |||
[[File:Fig2 Tsur BioDataMining2017 10.gif|584px]] | |||
{{clear}} | |||
{| | |||
| STYLE="vertical-align:top;"| | |||
{| border="0" cellpadding="5" cellspacing="0" width="584px" | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;"| <blockquote>'''Figure 2.''' Implemented framework. Our implementation of the proposed framework utilizes Java as the development environment, EclipseLink as the JPA provider and Apache Derby as the database manager. Parsing layers were based on J3D, jsoup, Apache Commons and Org.w3c libraries. We implemented structured interfaces to various databases including MalaCards, Biomodels and several NCBI’s data sets.</blockquote> | |||
|- | |||
|} | |||
|} | |||
The main data stream from established online databases was implemented using a structured URL interface. Databases often use a fixed URL syntax which translates a standard set of input parameters into the values necessary to search and retrieve requested data. For example, Entrez Programming Utilities provide a structured URL interface to the Entrez system, which currently includes 38 databases covering a variety of biomedical data, including nucleotide and protein sequences, gene records, three-dimensional molecular structures, and associated biomedical literature.<ref name="NCBIEntrez">{{cite web |url=https://www.ncbi.nlm.nih.gov/books/NBK25501/ |title=Entrez Programming Utilities Help |author=National Center for Biotechnology Information; U.S. National Library of Medicine |date=2017}}</ref> A series of data processing tools were utilized to implement parsers for syntactic analysis of the retrieved data. The w3c.dom package provides the Document Object Model (DOM) interfaces, which were used as the API for XML processing. The Apache Commons libraries, the jsoup library and the org.j3d library of the Java 3D Community were utilized for CSV, HTML and STL parsing, respectively. An API with which the user can specify the required database from which she wants to search and retrieve data was developed. This API contains an expandable database repository and a simple query generation engine which can be used to search a specified database. The user uses Java OOP to encapsulate the retrieved data and integrate it with her own data. The mapping of Java objects and database tables is defined via persistence metadata, which is used by the JPA provider — EclipseLink — to execute database SQL-like operations for static and dynamic queries. EclipseLink defines the metadata via annotations in the Java class and with XML. The open-source relational database Apache Derby — part of the Apache DB Project — was used as the database manager. Derby is written in Java and is suitable for embedding due to its limited footprint and ease-of-use. Derby supports SQL data storing and querying in a client/server operation mode, commonly used by specialized databases. | |||
==References== | ==References== | ||
Revision as of 22:45, 15 August 2017
| Full article title | Rapid development of entity-based data models for bioinformatics with persistence object-oriented design and structured interfaces |
|---|---|
| Journal | BioData Mining |
| Author(s) | Tsur, Elishai Ezra |
| Author affiliation(s) | Jerusalem College of Technology |
| Primary contact | Email: elishai85 at gmail dot com |
| Year published | 2017 |
| Volume and issue | 10 |
| Page(s) | 11 |
| DOI | 10.1186/s13040-017-0130-z |
| ISSN | 1756-0381 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://biodatamining.biomedcentral.com/articles/10.1186/s13040-017-0130-z |
| Download | https://biodatamining.biomedcentral.com/track/pdf/10.1186/s13040-017-0130-z (PDF) |
|
|
This article should not be considered complete until this message box has been removed. This is a work in progress. |
Databases are imperative for research in bioinformatics and computational biology. Current challenges in database design include data heterogeneity and context-dependent interconnections between data entities. These challenges drove the development of unified data interfaces and specialized databases. The curation of specialized databases is an ever-growing challenge due to the introduction of new data sources and the emergence of new relational connections between established datasets. Here, an open-source framework for the curation of specialized databases is proposed. The framework supports user-designed models of data encapsulation, object persistence and structured interfaces to local and external data sources such as MalaCards, Biomodels and the National Center for Biotechnology Information (NCBI) databases. The proposed framework was implemented using Java as the development environment, EclipseLink as the data persistence agent and Apache Derby as the database manager. Syntactic analysis was based on J3D, jsoup, Apache Commons and w3c.dom open libraries. Finally, a construction of a specialized database for aneurysm-associated vascular diseases is demonstrated. This database contains three-dimensional geometries of aneurysms, patients' clinical information, articles, biological models, related diseases and our recently published model of aneurysms’ risk of rapture. The framework is available at: http://nbel-lab.com.
Keywords: specialized databases, object-relational databases, EclipseLink, Apache Derby, object-oriented programming
Background
In the last few decades the intersection of computer science and biology has evolved to the point at which answers to fundamental biological questions have emerged.[1] Some of the most important cross-talks between biology and computer science lie within the data-intensive nature of modern biology.[2] It is currently evident that fields such as computational biology and bioinformatics are practically fueled by the increasing computational resources available and the development of software encapsulation and abstraction layers.[3] An important corner stone of the computer-science/biology interface is object-centered reductionism where relations between discrete biological entities such as DNA, protein and RNA are investigated.[1] Data regarding biological entities is stored in databases, which have become the most important corner stone for research in computational biology and bioinformatics.
Biological database designers currently face two main challenges: data heterogeneity and the emergence of new relational connections between data entities. Today, biological data is not limited to sequential information, which is typically stored in primary databases such as NCBI's Nucleotide and Protein data sets. Biological data also encompass graphs[4], statistical models[5], geometric information[6], vector fields[7], patterns[8], images[9], computational models[10] and others. A recent important advance regarding data heterogeneity was developed by Allan and colleagues, who have developed OMERO, an open-source software platform which uses a server-based middleware application to provide a unified interface for images, matrices and tables.[9] However, while OMERO provides a unified interface for file types, it is currently limited to microscopy images. Another important effort is the development of Semantic Web languages (SWLs), which promote web-based standardization of data formats by utilizing Extensible Markup Language (XML) and Resource Description Framework (RDF). SWLs have been implemented by many biological portals such as MGED Ontology, which provides terms for annotating microarray experiments; BioPAX, which provides an exchange format for biological pathway data; and Gene Ontology (GO), which describes biological processes, molecular functions and cellular components of gene products.[11]
The management of relational connections between biological data entities is a great challenge due to the variety of contexts in which data can be related. The vast spectrum of possible relations between biological entities drove the momentum for the curation of specialized databases. Specialized databases include organism-centered datasets such as Flybase (Drosophila)[12], WormBase (Nematode)[13], AceDB (C. elegans)[14], and TAIR (Arabidopsis)[15]; biological pathways databases such as MetaCyc and Biocyc[16]; and disease databases such as NCBI's OMIM database. Today, specialized databases are often curated to serve consortiums and single laboratories that define their own data relations architecture with their own data format. Specialized database curation is an ever-growing need since new data sources are constantly evolving due to rapidly advancing biological research: new experimental techniques produce types of data greater in both variety and number, requiring database structures to change accordingly. Moreover, most specialized databases contain both new data and data that were derived from established datasets. This hybrid approach of the new and the old presents a major challenge to specialized database designers, which should query, acquire and parse data from existing databases, as well as integrate it into their own database architecture.
The relational model, in which sets of tables are used to organize data, was first introduced by Edgar Codd in 1970 and is currently the most widely used model for data representation.[17] Although relational database management systems (DBMSs) have often been used for biological data management, they are in many ways inadequate. One of the main reasons for this inadequacy is that a relational data model cannot accurately encapsulate important biological data structures such as pedigrees, taxonomies, maps, networks, cascade processes, etc. Moreover, while application development techniques and programming languages have evolved significantly over the past decades, the relational database technology has remained relatively unchanged, frequently causing discrepancies between the object-oriented model used by many modern applications and the relational model.[18] However, while relational databases may be inconvenient to consume by modern programming languages, they are still the main choice for many applications due to their maturity and reliability. One of the main alternatives to the relational data model is an object-based representation of information, in which entities are defined with a set of properties and connected as attributes. Object-oriented database management systems (OODBMSs), popularized as NoSQL, allows the encapsulation of internal details of the data associated with the heterogeneity of the underlying data sources, extending object-oriented programming with data persistence. Data persistence allow objects to be created, stored and used directly with no need to explicitly serialize objects to or from a database. Examples of OODBMSs include hbase and DocDb. Importantly, a technique called object-relational mapping (ORM) allows the user to use SQL-like queries while dealing directly with objects, creating a hybrid of both object and relational approaches.
Here, an ORM-based open-source platform is proposed. The platform provides an efficient way for researchers to curate their own database using their own model of encapsulation, while providing direct access to external biological databases such as the National Center for Biotechnology Information (NCBI) databases, MalaCards, Biomodels and others using structured interfaces.
Implementation
Framework
Here, a simple framework for rapid development of specialized databases is proposed. This framework integrates several database technologies into a unified platform which allows the user to search and retrieve data from existing sources, incorporate established data with newly discovered information in a single data model, define object-based data architecture and persist it to memory for future inquiries. The framework consists of three main data streams: existing data sources to the user (search and retrieve data), new information sources such as experiments and data analytics to the user (generate data), as well as the individual user to a self-curated specialized data-base (model, store and retrieve data) (Fig. 1). The platform enables the user to search and retrieve data from local databases using structured information interfaces, and from online databases using structured URL interfaces. The user interface consists of a database repository which defines the differently structured accesses to existing resources, and a query generation engine which provides advanced searching mechanisms such as field-oriented search and the use of logical relations among search terms. The user designs their own data architecture using object-oriented programming (OOP), within which new data and the database-derived information are encapsulated to class-defined entities, an action which forms the database schema. Entities are persisted to memory with a persistent agent (PA). The PA uses an object relational mapper to map the object-based schema to a set of interconnected tables which resemble a typical relational database. This will allow the user to query the database with SQL-like queries. The relational model is transferred to a database manager that stores the data in memory.
|
This framework can be implemented in various ways using different programming languages and libraries providers. For example, Southern and colleagues developed a Java API, which maps entities to NCBI's PubChem schema and provides wrapper functions for calling NCBI eUtilities and PubChem web services.[19] Another implementation is the BioPython project, which provides modules to access NCBI's databases from within Python.[20] Object persistence can be achieved both in Python using its standard library, which support a family of hash-based file formats and objects serialization, and in JAVA using the Java Persistence API (JPA). Most JPA persistence providers offer the option to automatically create the database schema based on metadata. Popular implementations of JPA are Hibernate, EclipseLink and Apache OpenJPA. Common database managers include Apache Derby, Firebird, SQLite, and HSQL.
Software description
The proposed framework for the development of specialized databases can be implemented using different resources. Here, various open-source and free resources were utilized for implementation. Java was chosen as the development environment, EclipseLink as the JPA provider and Apache Derby as the database manager. Java was used to create interfaces to online databases such as MalaCards, Biomodels and NCBI's databases. The structured information interface was used to derive data from local repositories such as Aneurisk, which was downloaded from the Aneurisk web dataset. By integrating these tools with a series of data parsers, a powerful framework for the curation of specialized databases is provided, which would incorporate new data and database-derived information into a user-defined database architecture. A schematic of the implementation is presented in Fig. 2.
|
The main data stream from established online databases was implemented using a structured URL interface. Databases often use a fixed URL syntax which translates a standard set of input parameters into the values necessary to search and retrieve requested data. For example, Entrez Programming Utilities provide a structured URL interface to the Entrez system, which currently includes 38 databases covering a variety of biomedical data, including nucleotide and protein sequences, gene records, three-dimensional molecular structures, and associated biomedical literature.[21] A series of data processing tools were utilized to implement parsers for syntactic analysis of the retrieved data. The w3c.dom package provides the Document Object Model (DOM) interfaces, which were used as the API for XML processing. The Apache Commons libraries, the jsoup library and the org.j3d library of the Java 3D Community were utilized for CSV, HTML and STL parsing, respectively. An API with which the user can specify the required database from which she wants to search and retrieve data was developed. This API contains an expandable database repository and a simple query generation engine which can be used to search a specified database. The user uses Java OOP to encapsulate the retrieved data and integrate it with her own data. The mapping of Java objects and database tables is defined via persistence metadata, which is used by the JPA provider — EclipseLink — to execute database SQL-like operations for static and dynamic queries. EclipseLink defines the metadata via annotations in the Java class and with XML. The open-source relational database Apache Derby — part of the Apache DB Project — was used as the database manager. Derby is written in Java and is suitable for embedding due to its limited footprint and ease-of-use. Derby supports SQL data storing and querying in a client/server operation mode, commonly used by specialized databases.
References
- ↑ 1.0 1.1 Kitano, H. (2002). "Computational systems biology". Nature 420 (6912): 206–10. doi:10.1038/nature01254. PMID 12432404.
- ↑ Stein, L.D. (2003). "Integrating biological databases". Nature Reviews Genetics 4 (5): 337–45. doi:10.1038/nrg1065. PMID 12728276.
- ↑ Cannata, N.; Merelli, E.; Altman, R.B. (2005). "Time to organize the bioinformatics resourceome". PLOS Computational Biology 1 (7): e76. doi:10.1371/journal.pcbi.0010076. PMC PMC1323464. PMID 16738704. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1323464.
- ↑ Sharan, R.; Ideker, T. (2006). "Modeling cellular machinery through biological network comparison". Nature Biotechnology 24 (4): 427–33. doi:10.1038/nbt1196. PMID 16601728.
- ↑ Wilkinson, D.J. (2009). "Stochastic modelling for quantitative description of heterogeneous biological systems". Nature Reviews Genetics 10 (2): 122–33. doi:10.1038/nrg2509. PMID 19139763.
- ↑ Delp, S.L.; Ku, J.P.; Pande, V.S. et al. (2012). "Simbios: An NIH national center for physics-based simulation of biological structures". JAMIA 19 (2): 186–89. doi:10.1136/amiajnl-2011-000488. PMC PMC3277621. PMID 22081222. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3277621.
- ↑ Ezra, E.; Keinan, E.; Mandel, Y. et al. (2013). "Non-dimensional analysis of retinal microaneurysms: Critical threshold for treatment". Integrative Biology 5 (3): 474-80. doi:10.1039/c3ib20259c. PMC PMC3781337. PMID 23371018. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3781337.
- ↑ Naumova, O.Y.; Lee, M.; Koposov, R. et al. (2012). "Differential patterns of whole-genome DNA methylation in institutionalized children and children raised by their biological parents". Development and Psychopathology 24 (1): 143–55. doi:10.1017/S0954579411000605. PMC PMC3470853. PMID 22123582. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3470853.
- ↑ 9.0 9.1 Allan, C.; Burel, J.M.; Moore, J. et al. (2012). "OMERO: Flexible, model-driven data management for experimental biology". Nature Methods 9 (3): 245–53. doi:10.1038/nmeth.1896. PMC PMC3437820. PMID 22373911. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3437820.
- ↑ Chelliah, V.; Laibe, C.; Le Novère, N. et al. (2013). "BioModels Database: A repository of mathematical models of biological processes". Methods in Molecular Biology 1021: 189–99. doi:10.1007/978-1-62703-450-0_10. PMID 23715986.
- ↑ Pasquier, C. (2008). "Biological data integration using Semantic Web technologies". Biochimie 90 (4): 584–94. doi:10.1016/j.biochi.2008.02.007. PMID 18294970.
- ↑ FlyBase Consortium (2003). "The FlyBase database of the Drosophila genome projects and community literature". Nucleic Acids Research 31 (1): 172–5. PMC PMC165541. PMID 12519974. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC165541.
- ↑ Stein, L.; Sternberg, P.; Durbin, R. et al. (2001). "WormBase: Network access to the genome and biology of Caenorhabditis elegans". Nucleic Acids Research 29 (1): 82–6. PMC PMC29781. PMID 11125056. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC29781.
- ↑ Martinelli, S.D.; Brown, C.G.; Durbin, R. (1997). "Gene expression and development databases for C. elegans". Seminars in Cell & Developmental Biology 8 (5): 459–67. doi:10.1006/scdb.1997.0171.
- ↑ Poole, R.L. (2007). "The TAIR database". Methods in Molecular Biology 406: 179–212. PMID 18287693.
- ↑ Caspi, R.; Foerster, H.; Fulcher, C.A. et al. (2008). "The MetaCyc Database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases". Nucleic Acids Research 36 (DB1): D623–31. doi:10.1093/nar/gkm900. PMC PMC2238876. PMID 17965431. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2238876.
- ↑ Codd, E.F. (1970). "A relational model of data for large shared data banks". Communications of the ACM 13 (6): 377–87. doi:10.1145/362384.362685.
- ↑ Elmasri, R.; Navathe, S.B. (2008). Fundamentals of Database Systems (5th ed.). Pearson. ISBN 9780321369574.
- ↑ Southern, M.R.; Griffin, P.R. (2011). "A Java API for working with PubChem datasets". Bioinformatics 27 (5): 741—2. doi:10.1093/bioinformatics/btq715. PMC PMC3105478. PMID 21216779. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3105478.
- ↑ Cock, P.J.; Antao, T.; Chang, J.T. et al. (2009). "Biopython: Freely available Python tools for computational molecular biology and bioinformatics". Bioinformatics 25 (11): 1422-3. doi:10.1093/bioinformatics/btp163. PMC PMC2682512. PMID 19304878. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2682512.
- ↑ National Center for Biotechnology Information; U.S. National Library of Medicine (2017). "Entrez Programming Utilities Help". https://www.ncbi.nlm.nih.gov/books/NBK25501/.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added. Grammar and word use were updated to make the text easier to read.