Journal:systemPipeR: NGS workflow and report generation environment
| Full article title | systemPipeR: NGS workflow and report generation environment |
|---|---|
| Journal | BMC Bioinformatics |
| Author(s) | Backman, Tyler W.H.; Girke, Thomas |
| Author affiliation(s) | University of California, Riverside |
| Primary contact | Email: thomas dot girke at ucr dot edu |
| Year published | 2016 |
| Volume and issue | 17 |
| Page(s) | 388 |
| DOI | 10.1186/s12859-016-1241-0 |
| ISSN | 1471-2105 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-016-1241-0 |
| Download | https://bmcbioinformatics.biomedcentral.com/track/pdf/10.1186/s12859-016-1241-0 (PDF) |
Abstract
Background: Next-generation sequencing (NGS) has revolutionized how research is carried out in many areas of biology and medicine. However, the analysis of NGS data remains a major obstacle to the efficient utilization of the technology, as it requires complex multi-step processing of big data, demanding considerable computational expertise from users. While substantial effort has been invested on the development of software dedicated to the individual analysis steps of NGS experiments, insufficient resources are currently available for integrating the individual software components within the widely used R/Bioconductor environment into automated workflows capable of running the analysis of most types of NGS applications from start-to-finish in a time-efficient and reproducible manner.
Results: To address this need, we have developed the R/Bioconductor package systemPipeR. It is an extensible environment for both building and running end-to-end analysis workflows with automated report generation for a wide range of NGS applications. Its unique features include a uniform workflow interface across different NGS applications, automated report generation, and support for running both R and command-line software on local computers and computer clusters. A flexible sample annotation infrastructure efficiently handles complex sample sets and experimental designs. To simplify the analysis of widely used NGS applications, the package provides pre-configured workflows and reporting templates for RNA-Seq, ChIP-Seq, VAR-Seq, and Ribo-Seq. Additional workflow templates will be provided in the future.
Conclusions: systemPipeR accelerates the extraction of reproducible analysis results from NGS experiments. By combining the capabilities of many R/Bioconductor and command-line tools, it makes efficient use of existing software resources without limiting the user to a set of predefined methods or environments. systemPipeR is freely available for all common operating systems from Bioconductor (http://bioconductor.org/packages/devel/systemPipeR).
Keywords: analysis workflow, next generation sequencing (NGS), Ribo-Seq, ChIP-Seq, RNA-Seq, VAR-Seq
Background
By allowing scientists to rapidly sequence and quantify DNA and RNA molecules, next-generation sequencing (NGS) technology has transformed biology into one of the most data intensive research disciplines. In the past, experiments have been performed on a gene-by-gene basis, while NGS has introduced an age where it is has become a routine to sequence entire transcriptomes, genomes, or epigenomes rather than their isolated parts of interest. It will soon be possible to conduct these experiments on large numbers of single cell samples[1][2] for a wide range of time points, treatments, and genetic backgrounds to study biological systems with greater resolution and precision. Sequencing the genetic material of each individual within entire populations of organisms of the same species or genus will enable the study of adaptation processes[3], disease progression, and micro-evolution in real time.[4] This technological shift empowers researchers to address questions at a genome-wide scale, for example by profiling the mRNA, miRNA, and DNA methylation states of a large set of biological samples in parallel.[5]
The success of NGS-driven research has led to a data explosion of increasing size and complexity, making it now more time-consuming and challenging for researchers to extract knowledge from their experiments. Rapid processing of the results is essential to test, refine, and formulate new hypotheses for designing follow-up experiments. As a result, biologists have to dedicate nowadays substantial time to data analysis tasks while training themselves effectively as genome data scientists rather than focusing on experimentation as they used to in the past.
In recent years, a considerable number of algorithms, statistical methods, and software tools has been developed to perform the individual analysis steps of different NGS applications. These include short read pre-processors, aligners, variant and peak callers, as well as statistical methods for the analysis of genomic regions that are differentially expressed[6][7], bound[8], or methylated.[9][10] Also essential are tools for processing short read alignments[11], genomic intervals[12][13], and annotations.[14] However, most data analysis routines of NGS applications are very complex, involving multiple software tools for their many processing steps. As a result, there is a great need for flexible software environments connecting the individual software components to automated workflows in order to perform complex genome-wide analyses in an efficient and reproducible manner. While many workflow management resources exist[15][16][17][18][19][20][21][22][23][24] for a variety of data analysis programming languages (for details see below), only insufficient general purpose NGS workflow solutions are currently available for the popular R programming language. R and the affiliated Bioconductor environment provide a substantial number of widely used tools with a large user base in this area.[10] Thus, a workflow framework for federating NGS applications from within R will have many benefits for experimental and computational scientists who use R for NGS data analysis.
To address this need, we designed systemPipeR as a Bioconductor package for building and running workflows for most NGS applications, with support for integrating a wide array of command-line and R/Bioconductor software.
Implementation
Environment
systemPipeR has been implemented as an open-source Bioconductor package using the R programming language for statistical computing and graphics. R was chosen as the core development platform for systemPipeR because of the following reasons. (i) R is currently one of the most popular statistical data analysis and programming environments in bioinformatics. (ii) Its external language bindings support the implementation of computationally time-consuming analysis steps in high-performance languages such as C/C++. (iii) It supports advanced parallel computation on multi-core machines and computer clusters. (iv) A well developed infrastructure interfaces R with several other popular programing languages such as Python. (v) R provides advanced graphical and visualization utilities for scientific computing. (vi) It offers access to a vast landscape of statistical and machine learning tools. (vii) Its integration with the Bioconductor project promotes reusability of genomics software components, while also making efficient use of a large number of existing NGS packages that are well tested and widely used by the community. To support long-term reproducibility of analysis outcomes, systemPipeR is also distributed as a Docker image of Bioconductor’s sequencing division. Docker containers provide an efficient solution for packaging complex software together with all its system dependencies to ensure it will run the same in the future across different environments, including different operating systems and cloud-based solutions.
Workflow design
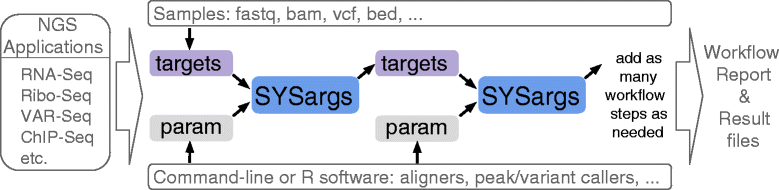
systemPipeR workflows (Fig. 1) can be run from start-to-finish with a single command, or stepwise in interactive mode from the R console. New workflows are constructed, or existing ones modified, by connecting so-called SYSargs workflow control modules (R S4 class). Each SYSargs module contains instructions needed for processing a set of input files with a specific command-line or R software; as well as the paths to the corresponding outputs generated by a specific NGS tool such as a read preprocessor (trimmed/filtered FASTQ files), aligner (SAM/BAM files), read counter, variant caller (VCF/BCF files), peak caller (BED/WIG files), or statistical function. Typically, the only input the user needs to provide for running workflows is a single tabular targets file containing the paths to the initial sample input files (e.g. FASTQ) along with sample labels, and if appropriate biological replicate and contrast information for controlling differential abundance analyses (e.g., gene expression). Downstream derivatives of these targets files along with the corresponding SYSargs instances (see Fig. 1) are created automatically within each workflow.
|
The parameters required for running command-line software are provided by parameter (param) files, described below. For R-based workflow steps, param files are not required but can be useful for operations importing and/or exporting sample-level files. This modular design has several advantages. First, it provides a high level of flexibility for designing workflows, such as allowing the user to start workflows from the very beginning or anywhere in-between (e.g. FASTQ or BAM level). Second, it is straightforward to add custom workflow steps without requiring computational expert knowledge from users. Workflows can also have any number of steps including branch points. Lastly, it also minimizes errors as all input and output files are registered, and sample labels specified in the initial targets file will be consistently used throughout all workflow results, including plots, tables, and workflow reports.
Command-line software support
An important feature of systemPipeR is support for running command-line software directly from R on both single machines or computer clusters. This offers several advantages, such as seamless integration of most command-line software available in the NGS field with the extensive genome analysis resources provided by R/Bioconductor. The user interface for running command-line software has been generalized as a single function for ease of use, while only one additional command will run the same tool in parallel mode on a computer cluster (see below). Examples of command-line software used by systemPipeR’s preconfigured workflow templates (see below) include the aligners BWA-MEM[25], Bowtie2[26], TopHat2[27], HISAT2[28], as well as the peak/variant callers MACS[29], GATK[30], and BCFtools.[11] Support for additional command-line NGS software can be added by simply providing the argument settings of a chosen software in a tabular param file. If appropriate, new param files can be permanently included in the package to share them with the community. Functionality for creating param files automatically will be provided in the future. This will allow users to create new param instances simply by providing an example of the command-line syntax of a chosen software tool.
Major advantages of running command-line software from within systemPipeR include: a uniform sample management infrastructure within and across workflows; integration of BatchJobs’[31] efficient error management infrastructure for job submissions on computer clusters; the simplicity of restarting failed processes; as well as seamless addition of new samples (e.g., FASTQ or BAM files). In case of a restart, the system will skip the analysis steps of already completed samples and only perform the analysis of the missing ones. If required, any workflow step can be rerun on demand for all or a subset of samples. When submitting command-line software to computer clusters, BatchJobs monitors the status of job submissions and alerts users of exceptions, while recording warning and error messages for each process in a log directory with a database-like structure that is accessible from within R or the command-line. This organization helps to diagnose and resolve errors.
Parallel evaluation
The processing time for NGS experiments can be greatly reduced by making use of parallel evaluation across several CPU cores on single machines, or multiple nodes of computer clusters and cloud-based systems. systemPipeR simplifies these parallelization tasks without creating any limitations for users who do not have access to high-performance computer (HPC) resources by providing the option to run workflows in serial or parallel mode. The parallelization functionalities available in systemPipeR are largely based on existing and well maintained R packages, mainly BatchJobs and BiocParallel.[31] By making use of cluster template files, most schedulers and queuing systems are also supported (e.g., Torque, Sun Grid Engine, Slurm). If required, entire workflows can be executed in parallel mode by issuing a single command, while simultaneously generating a detailed analysis report (for details see below). If sufficient parallel computer resources are available, systemPipeR can complete the entire analysis workflow of several complex NGS experiments, each containing large numbers of FASTQ files, within hours rather than days or weeks, as can be the case for non-parallelized workflows.
Automated analysis reports
systemPipeR generates automated analysis reports with knitr and R markdown.[32] These modern reporting environments integrate R code with LaTeX or Markdown. During the evaluation of the R code, reports are dynamically generated in PDF or HTML format. A caching system allows to re-execute selected workflow reporting steps without repeating unnecessary components. This way one can generate reports that resemble a research paper where user generated text is combined with analysis results. This includes support for citations, autogenerated bibliographies, code chunks with syntax highlighting, and inline evaluation of variables to update text content. Data components in a report such as tables and figures are updated automatically when rebuilding the document and/or rerunning workflows partially or entirely.
Results and discussion
Overview
systemPipeR provides utilities for building and running NGS analysis workflows. To adapt to community standards, widely used R/Bioconductor packages are integrated where possible. This includes the Bioconductor packages ShortRead, Biostrings, and Rsamtools for processing sequence and alignment files[33]; GenomicRanges, GenomicAlignments, and GenomicFeatures for handling genomic range operations, read counting, and annotation data[12]; edgeR and DESeq2 for differential abundance analysis[6][7]; and VariantTools and VariantAnnotation for filtering and annotating genome variants.[34] If necessary, one can substitute most of these packages with alternative R or command-line tools.
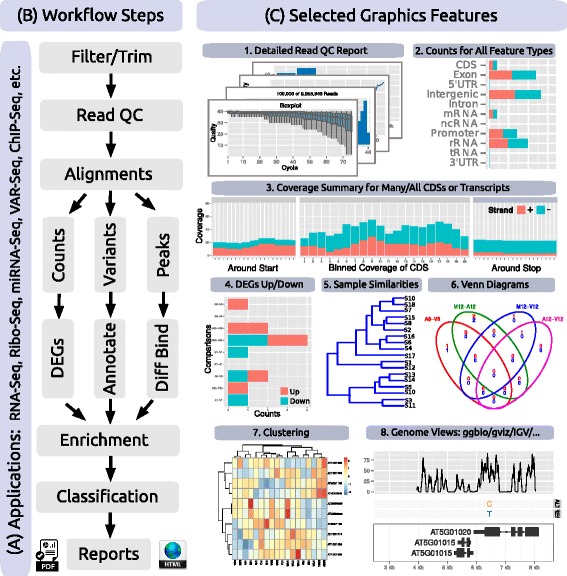
Because many NGS applications share overlapping analysis needs (Fig. 2 a), certain workflow steps are conceptualized in systemPipeR by a single generic function, with support for application-specific parameter settings (Table 1). For instance, most NGS applications involve a short read alignment step (see Fig. 2 b), but with very distinct mapping requirements, such as splice junction awareness and variant tolerance for RNA-Seq and VAR-Seq, respectively. To simplify their execution for the user, the different aligners can be run with the same runCommandline function where the software, and its parameter settings are specified in the corresponding SYSargs instance (see above and Fig. 1).
|
| ||||||||||||||||||||||||||||||||||||
Workflow templates
systemPipeR also provides end-to-end workflow templates for RNA-Seq, Ribo-Seq, ChIP-Seq, and VAR-Seq analysis. A detailed vignette (manual) is provided for each workflow, while an overview vignette introduces the general design concepts. Templates for additional NGS applications will be made available in the future. To test workflows quickly or design new ones from existing templates, users can generate with a single command (genWorkenvir) workflow instances fully populated with sample data and parameter files required for running a chosen workflow. The corresponding sample data are provided by the affiliated data package systemPipeRdata, also available from Bioconductor. To illustrates the utilities of systemPipeR’s workflow templates, a case study has been included as Additional file 1 that guides the reader through the most important steps of a sample workflow. A typical gene-level RNA-Seq analysis was chosen here because it is currently one of the most widely used applications in the NGS field.
Add-on tools
In addition to providing a framework for running NGS analysis workflows, systemPipeR includes many functions and methods that expand and enhance its workflows. The following gives selected examples of these utilities (also illustrated in Fig. 2 c and Table 1). A read pre-processor function (preprocessReads) addresses the often very sophisticated quality filtering and adaptor trimming needs of specialized NGS applications such as Ribo-Seq or smallRNA-Seq. The functions seeFastq and seeFastqPlot generate and plot detailed quality reports for FASTQ files (Fig. 2 c1). These reports are easy to generate and designed to facilitate the visual inspection of large numbers of FASTQ files in a single report. The featuretypeCounts function computes and plots the distribution of reads across all features available in a genome annotation rather than just a single one (Fig. 2 c2). The featureCoverage function generates from genome-level alignments read depth coverage summaries for all or a subset of transcripts with nucleotide resolution upstream/downstream of their start and stop codons, as well as binned coverage for their coding regions (Fig. 2 c3). Additional utilities include functions to automate the analysis of differentially expressed genes (DEGs) with edgeR or DESeq2 (Fig. 2 c4), to compute Venn intersects for large numbers of sample sets (e.g. 2-20 or as many as available memory allows) with plotting functionalities for 2-5 way Venn diagrams (Fig. 2 c6), and to run gene set enrichment analyses in batch mode on large numbers of gene sets. The modular design of the systemPipeR environment allows users to easily substitute any of these built-in tools with alternative R-based or command-line software, such as using FastQC[35], FASTX-Toolkit[36], or MultiQC[37] for quality reporting, read trimming or result aggregation, respectively.
Performance and scalability
systemPipeR has been optimized to run workflows in a time and memory efficient manner even on very large read sets from complex genomes (e.g., mammalian genomes). This is achieved by making heavy use of indexing, file streaming, and parallelization functionalities. For instance, users can limit the RAM requirements of several workflow steps by specifying the maximum number of reads or alignments to stream into memory at any time. This enables analysis of very large files with tens of GBs of storage space on systems with limited RAM resources, making it possible to run systemPipeR workflows even on laptops or smaller workstations, provided they have the required software installed and enough disk space available for storing large NGS input and result files. The processing time of non-parallelized analysis steps depends on the time performance of a specific software tool chosen for a workflow step. For instance, in the RNA-Seq workflow described under Additional file 1 the alignment step will run on a single sample (FASTQ file) with the native time performance of the chosen aligner Bowtie2/Tophat2. Using the much faster HISAT2 aligner instead would accelerate the alignment step proportionally to the time improvements provided by this aligner without the need of additional parallel computer resources.[28]
On a computer cluster, parallelized systemPipeR workflows scale nearly linearly in time with the number of sample files (i.e., FASTQ files) since every step can be parallelized at the sample level. In practice this means, the runtime of an analysis of 100 FASTQ files can be accelerated by 10 or 100 fold when using instead of a single CPU core 10 or 100 CPU cores, respectively. For example, the RNA-Seq workflow in Additional file 1 can process 100 FASTQ files, each with 30–40 million reads from a mammalian genome, in six to eight hours using 100 CPU cores (CPU Model: AMD 6376, 2.3 GHz) and a maximum RAM requirement of less than 10 GB per node. Since the alignment step with Bowtie2/Tophat2 accounts for most of the compute time of the entire workflow, the use of faster RNA-Seq aligners, such as Rsubread or HISAT2, can reduce the compute time to less than three hours. With comparable parallel computer resources available, one can complete with systemPipeR the end-to-end analysis of several complex NGS experiments each containing 50–100 FASTQ files in less than a day rather than many days or weeks as is common in non-parallelized workflows.
Need for an R-based NGS workflow environment
Several related software tools with NGS workflow functionality are available. These include Galaxy[15][38], Snakemake[16], Taverna[17], BioBlend[39], bcbio-nextgen[18], Knime[19], Ruffus[20], Kepler[21], Wasp[22], ViennaNGS[23], Mercury[24], RAP[40], and LONI[41] among others. Additionally, general purpose utilities for workflow management and design are provided by Rabix[42] and WDL.[43]
These tools provide infrastructure for streamlining the analysis of NGS data in a variety of data analysis environments and computer languages. However, only limited resources are available for designing and running analysis workflows for a wide range of NGS applications directly from within R as is possible with systemPipeR. One of the few exceptions is QuasR.[44] This Bioconductor package supports the initial analysis steps of several NGS applications, but it lacks an interface to integrate external command-line software and functionalities to build new workflows. Other existing R/Bioconductor resources for analyzing NGS data address the needs in this area only partially. For instance, many of them are limited to certain NGS applications, or cover only a subset of the processing steps required for complete workflows; do not support command-line software; or lack workflow design functionalities for different NGS applications. systemPipeR has been designed to address these requirements. However, it is important to mention here that well established community workflow environments like Galaxy provide several additional features not available in systemPipeR. A small sub-selection of them includes: (i) a web interface to support non-expert users who are not familiar with data analysis programming environments like R; (ii) support for a wider range of data types outside of the NGS field; (iii) a well-established infrastructure and community for archiving and sharing workflow protocols; or (iv) support for additional reporting technologies such as iPython notebooks. To take advantage of this powerful infrastructure, Galaxy compatible versions of systemPipeR’s NGS workflows will be released in the future. This will allow biologists to run them from an easy-to-use web interface, while also being able to access additional functionalities available in Galaxy’s large ecosystem of analysis tools.
Conclusion
The systemPipeR package unites R/Bioconductor resources with external command-line software to standardize and automate the analysis of a wide range of NGS applications. Its functionalities reduce the complexity and time required to translate NGS data into interpretable research results, while a built-in reporting feature improves reproducibility. The environment provides sufficient flexibility to choose the optimal software for each step in complex NGS workflows, customize workflows, and design new workflows. Pre-configured workflow templates are included for several NGS applications. Templates for additional NGS applications are under development and will be added to the package in the near future.
Availability and requirements
Project name: systemPipeR workflow environment
Project home page: https://bioconductor.org/packages/systemPipeR/
Archived version: systemPipeR
Operating system(s): Platform-independent
Programming language: R
Other requirements: R version ≥3.2, Bioconductor version ≥3.2
License: Artistic-2-0
Any restrictions to use by non-academics: none
Abbreviations
BAM: Binary version of sequence alignment map format
ChIP-Seq: Chromatin immunoprecipitation sequencing
DEG: Differentially expressed genes
FASTQ: short read sequence file format
NGS: Next generation sequencing
Ribo-Seq: NGS profiling of mRNA populations bound to ribosomes
RNA-Seq: NGS profiling of mRNA
SAM: Sequence alignment map format
VAR-Seq: NGS-based variant detection
Declarations
Acknowledgements
We acknowledge the Bioconductor core team and community for providing valuable input for developing systemPipeR.
Funding
This work was supported by grants from the National Science Foundation (ABI-0957099, MCB-1021969, IOS-1546879), the National Institutes of Health (U24AG051129, R01-AI36959), and the National Institute of Food and Agriculture (2011-68004-30154).
Authors’ contributions
TB and TG conceived the idea for systemPipeR. TG developed the methods, implemented the R package, and wrote the article. Both authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Additional files
Additional file 1: RNA-Seq Workflow Example. Case study to illustrate the utilities of systemPipeR using an RNA-Seq workflow as example. (PDF 89 kb)
References
- ↑ Kalisky, T.; Quake, S.R. (2011). "Single-cell genomics". Nature Methods 8 (4): 311–4. doi:10.1038/nmeth0411-311. PMID 21451520.
- ↑ Trapnell, C.; Cacchiarelli, D.; Grimsby, J. et al. (2014). "The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells". Nature Biotechnology 32 (4): 381–86. doi:10.1038/nbt.2859. PMC PMC4122333. PMID 24658644. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4122333.
- ↑ Lindblad-Toh, K.; Garber, M.; Zuk, O. et al. (2011). "A high-resolution map of human evolutionary constraint using 29 mammals". Nature 478 (7370): 476–82. doi:10.1038/nature10530. PMC PMC3207357. PMID 21993624. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3207357.
- ↑ Kato-Maeda, M.; Ho, C.; Passarelli, B. et al. (2013). "Use of whole genome sequencing to determine the microevolution of Mycobacterium tuberculosis during an outbreak". PLoS One 8 (3): e58235. doi:10.1371/journal.pone.0058235. PMC PMC3589338. PMID 23472164. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3589338.
- ↑ Holt, R.A.; Jones, S.J. (2008). "The new paradigm of flow cell sequencing". Genome Research 18 (6): 839-46. doi:10.1101/gr.073262.107. PMID 18519653.
- ↑ 6.0 6.1 Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. (2010). "edgeR: A Bioconductor package for differential expression analysis of digital gene expression data". Bioinformatics 26 (1): 139–40. doi:10.1093/bioinformatics/btp616. PMC PMC2796818. PMID 19910308. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2796818.
- ↑ 7.0 7.1 Love, M.I.; Huber, W.; Anders, S. (2014). "Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2". Genome Biology 15 (12): 550. doi:10.1186/s13059-014-0550-8. PMC PMC4302049. PMID 25516281. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4302049.
- ↑ Kharchenko, P.V.; Tolstorukov, M.Y.; Park, P.J. (2008). "Design and analysis of ChIP-seq experiments for DNA-binding proteins". Nature Biotechnology 26 (12): 1351–9. doi:10.1038/nbt.1508. PMC PMC2597701. PMID 19029915. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2597701.
- ↑ Akalin, A.; Kormaksson, M.; Li, S. et al. (2012). "methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles". Genome Biology 13 (10): R87. doi:10.1186/gb-2012-13-10-r87. PMC PMC3491415. PMID 23034086. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3491415.
- ↑ 10.0 10.1 Huber, W.; Carey, V.J.; Gentleman, R. et al. (2015). "Orchestrating high-throughput genomic analysis with Bioconductor". Nature Methods 12 (2): 115–21. doi:10.1038/nmeth.3252. PMC PMC4509590. PMID 25633503. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4509590.
- ↑ 11.0 11.1 Li, H.; Handsaker, B.; Wysoker, A. et al. (2009). "The Sequence Alignment/Map format and SAMtools". Bioinformatics 25 (16): 2078–9. doi:10.1093/bioinformatics/btp352. PMC PMC2723002. PMID 19505943. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2723002.
- ↑ 12.0 12.1 Lawrence, M.; Huber, W.; Pagès, H. et al. (2013). "Software for computing and annotating genomic ranges". PLoS Computational Biology 9 (8): e1003118. doi:10.1371/journal.pcbi.1003118. PMC PMC3738458. PMID 23950696. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3738458.
- ↑ Quinlan, A.R.; Hall, I.M. (2010). "BEDTools: a flexible suite of utilities for comparing genomic features". Bioinformatics 26 (6): 841-2. doi:10.1093/bioinformatics/btq033. PMC PMC2832824. PMID 20110278. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2832824.
- ↑ Durinck, S.; Moreau, Y.; Kasprzyk, A. (2005). "BioMart and Bioconductor: A powerful link between biological databases and microarray data analysis". Bioinformatics 21 (16): 3439-40. doi:10.1093/bioinformatics/bti525. PMID 16082012.
- ↑ 15.0 15.1 Goecks, Jeremey; Nekrutenko, Anton; Taylor, James; The Galaxy Team (2010). "Galaxy: A comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences". Genome Biology 11 (8): R86. doi:10.1186/gb-2010-11-8-r86. PMC PMC2945788. PMID 20738864. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2945788.
- ↑ 16.0 16.1 Köster, J.; Rahmann, S. (2012). "Snakemake: A scalable bioinformatics workflow engine". Bioinformatics 28 (19): 2520-2. doi:10.1093/bioinformatics/bts480. PMID 22908215.
- ↑ 17.0 17.1 Wolstencroft, K.; Haines, R.; Fellows, D. et al. (2013). "The Taverna workflow suite: Designing and executing workflows of Web Services on the desktop, web or in the cloud". Nucleic Acids Research 41 (W1): W557-W561. doi:10.1093/nar/gkt328. PMC PMC3692062. PMID 23640334. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3692062.
- ↑ 18.0 18.1 Guimera, R.V. (2012). "bcbio-nextgen: Automated, distributed next-gen sequencing pipeline". ENBnet Journal 17 (B): 30. doi:10.14806/ej.17.B.286.
- ↑ 19.0 19.1 Warr, W.A. (2012). "Scientific workflow systems: Pipeline Pilot and KNIME". Journal of Computer-aided Molecular Design 26 (7): 801–4. doi:10.1007/s10822-012-9577-7. PMC PMC3414708. PMID 22644661. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3414708.
- ↑ 20.0 20.1 Goodstadt, L. (2010). "Ruffus: A lightweight Python library for computational pipelines". Bioinformatics 26 (21): 2778-9. doi:10.1093/bioinformatics/btq524. PMID 20847218.
- ↑ 21.0 21.1 Stropp, T.; McPhillips, T.; Ludäscher, B.; Bieda, M. (2012). "Workflows for microarray data processing in the Kepler environment". BMC Bioinformatics 13: 102. doi:10.1186/1471-2105-13-102. PMC PMC3431220. PMID 22594911. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3431220.
- ↑ 22.0 22.1 McLellan, A.S.; Dubin, R.; Jing, Q. et al. (2012). "The Wasp System: An open source environment for managing and analyzing genomic data". Genomics 100 (6): 345-51. doi:10.1016/j.ygeno.2012.08.005. PMID 22944616.
- ↑ 23.0 23.1 Wolfinger, M.T.; Fallmann, J.; Eggenhofer, F.; Amman, F. (2015). "ViennaNGS: A toolbox for building efficient next-generation sequencing analysis pipelines". F1000Research 4: 50. doi:10.12688/f1000research.6157.2. PMC PMC4513691. PMID 26236465. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4513691.
- ↑ 24.0 24.1 Reid, J.G.; Carroll, A.; Veeraraghavan, N. et al. (2014). "Launching genomics into the cloud: Deployment of Mercury, a next generation sequence analysis pipeline". BMC Bioinformatics 15: 30. doi:10.1186/1471-2105-15-30. PMC PMC3922167. PMID 24475911. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3922167.
- ↑ Li, H. (26 May 2013). "Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM". arXiv.org. Cornell University Library. https://arxiv.org/abs/1303.3997.
- ↑ Langmead, B.; Salzberg, S.L. (2012). "Fast gapped-read alignment with Bowtie 2". Nature Methods 9 (4): 357-9. doi:10.1038/nmeth.1923. PMC PMC3322381. PMID 22388286. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3322381.
- ↑ Kim, D.; Pertea, G.; Trapnell, C. et al. (2013). "TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions". Genome Biology 14: R36. doi:10.1186/gb-2013-14-4-r36. PMC PMC4053844. PMID 23618408. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4053844.
- ↑ 28.0 28.1 Kim, D.; Langmead, B.; Salzberg, S.L. (2015). "HISAT: A fast spliced aligner with low memory requirements". Nature Methods 12 (4): 357-60. doi:10.1038/nmeth.3317. PMC PMC4655817. PMID 25751142. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4655817.
- ↑ Zhang, Y.; Liu, T.; Meyer, C.A. et al. (2008). "Model-based analysis of ChIP-Seq (MACS)". Genome Biology 9 (9): R137. doi:10.1186/gb-2008-9-9-r137. PMC PMC2592715. PMID 18798982. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2592715.
- ↑ McKenna, A.; Hanna, M.; Banks, E. et al. (2010). "The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data". Genome Research 20: 1297-1303. doi:10.1101/gr.107524.110.
- ↑ 31.0 31.1 Bischl, B.; Lang, M.; Mersmann, O. et al. (2012). "BatchJobs and BatchExperiments: Abstraction Mechanisms for Using R in Batch Environments". Journal of Statistical Software 64 (11): 1–25. doi:10.18637/jss.v064.i11.
- ↑ Xie, Y. (2013). Dynamic Documents with R and knitr (1st ed.). Chapman and Hall/CRC. pp. 216. ISBN 9781482203530.
- ↑ Morgan, M.; Anders, S.; Lawrence, M. et al. (2009). "ShortRead: A bioconductor package for input, quality assessment and exploration of high-throughput sequence data". Bioinformatics 25 (19): 2607-8. doi:10.1093/bioinformatics/btp450. PMC PMC2752612. PMID 19654119. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2752612.
- ↑ Obenchain, V.; Lawrence, M.; Carey, V. et al. (2014). "VariantAnnotation: a Bioconductor package for exploration and annotation of genetic variants". Bioinformatics 30 (14): 2076-8. doi:10.1093/bioinformatics/btu168. PMC PMC4080743. PMID 24681907. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4080743.
- ↑ "FastQC". Babraham Bioinformatics. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/. Retrieved 15 September 2015.
- ↑ "FASTX-Toolkit". Hannon Laboratory. http://hannonlab.cshl.edu/fastx_toolkit/index.html. Retrieved 17 September 2015.
- ↑ Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. (2016). "MultiQC: summarize analysis results for multiple tools and samples in a single report". Bioinformatics 32 (19): 3047–3048. doi:10.1093/bioinformatics/btw354. PMC PMC5039924. PMID 27312411. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5039924.
- ↑ Afgan, E.; Baker, D.; Coraor, N. et al. (2011). "Harnessing cloud computing with Galaxy Cloud". Nature Biotechnology 29 (11): 972-4. doi:10.1038/nbt.2028. PMC PMC3868438. PMID 22068528. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3868438.
- ↑ Sloggett, C.; Goonasekera, N.; Afgan, E. (2013). "BioBlend: automating pipeline analyses within Galaxy and CloudMan". Bioinformatics 29 (13): 1685-6. doi:10.1093/bioinformatics/btt199. PMC PMC4288140. PMID 23630176. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4288140.
- ↑ D'Antonio, M.; D'Onorio De Meo, P.; Pallocca, M. et al. (2015). "RAP: RNA-Seq Analysis Pipeline, a new cloud-based NGS web application". BMC Genomics 16: S3. doi:10.1186/1471-2164-16-S6-S3. PMC PMC4461013. PMID 26046471. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4461013.
- ↑ Torri, F.; Dinov, I.D.; Zamanyan, A. et al. (2012). "Next generation sequence analysis and computational genomics using graphical pipeline workflows". Genes 3 (3): 545–75. doi:10.3390/genes3030545. PMC PMC3490498. PMID 23139896. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3490498.
- ↑ "Rabix". Seven Bridges Genomics, Inc. http://rabix.io/.
- ↑ Broad Institute. "broadinstitute/wdl". GitHub. https://github.com/broadinstitute/wdl. Retrieved 16 September 2015.
- ↑ Gaidatzis, D.; Lerch, A.; Hahne, F. et al. (2015). "QuasR: Quantification and annotation of short reads in R". Bioinformatics 31: 7. doi:10.1093/bioinformatics/btu781. PMC PMC4382904. PMID 25417205. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4382904.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added. The original URL to Rabix was dead, and it was replaced with a current one for this version.