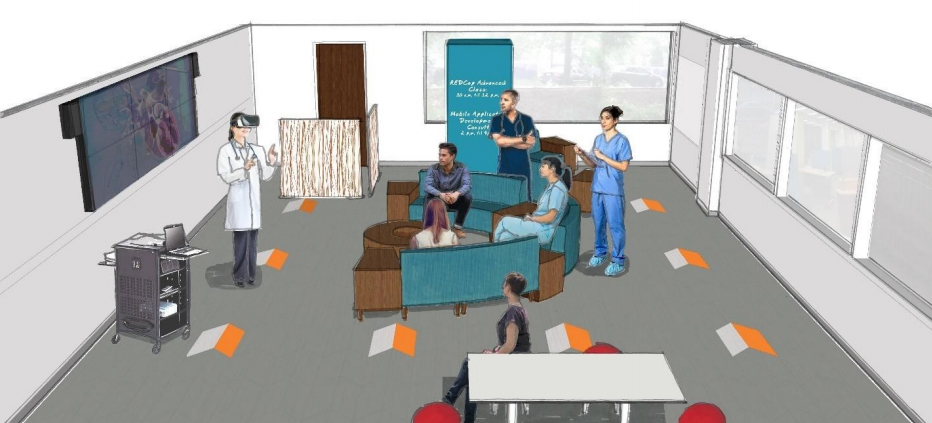
Featured article of the week: April 15–21:"Semantics for an integrative and immersive pipeline combining visualization and analysis of molecular data"
The advances made in recent years in the field of structural biology significantly increased the throughput and complexity of data that scientists have to deal with. Combining and analyzing such heterogeneous amounts of data became a crucial time consumer in the daily tasks of scientists. However, only few efforts have been made to offer scientists an alternative to the standard compartmentalized tools they use to explore their data and that involve a regular back and forth between them. We propose here an integrated pipeline especially designed for immersive environments, promoting direct interactions on semantically linked 2D and 3D heterogeneous data, displayed in a common working space. The creation of a semantic definition describing the content and the context of a molecular scene leads to the creation of an intelligent system where data are (1) combined through pre-existing or inferred links present in our hierarchical definition of the concepts, (2) enriched with suitable and adaptive analyses proposed to the user with respect to the current task and (3) interactively presented in a unique working environment to be explored. (Full article...)
Featured article of the week: April 8–14:
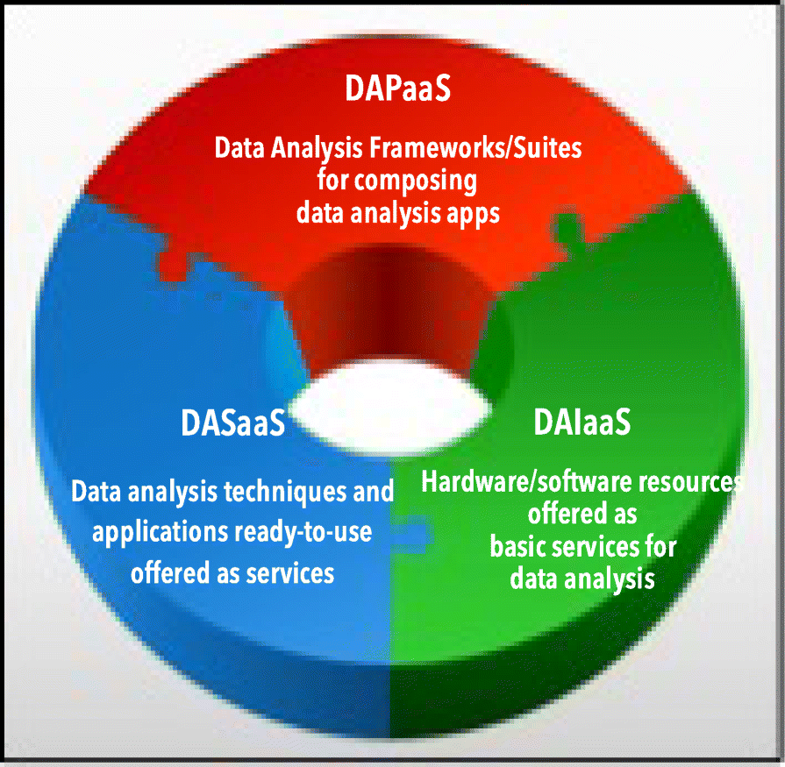
"A view of programming scalable data analysis: From clouds to exascale"
Scalability is a key feature for big data analysis and machine learning frameworks and for applications that need to analyze very large and real-time data available from data repositories, social media, sensor networks, smartphones, and the internet. Scalable big data analysis today can be achieved by parallel implementations that are able to exploit the computing and storage facilities of high-performance computing (HPC) systems and cloud computing systems, whereas in the near future exascale systems will be used to implement extreme-scale data analysis. Here is discussed how cloud computing currently supports the development of scalable data mining solutions and what the main challenges to be addressed and solved for implementing innovative data analysis applications on exascale systems currently are. (Full article...)
|
Featured article of the week: April 1–7:
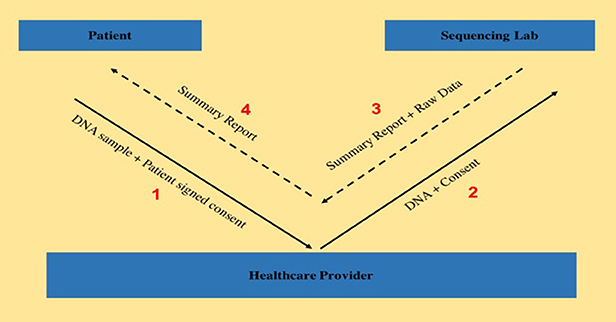
"Transferring exome sequencing data from clinical laboratories to healthcare providers: Lessons learned at a pediatric hospital"
The adoption rate of genome sequencing for clinical diagnostics has been steadily increasing, leading to the possibility of improvement in diagnostic yields. Although laboratories generate a summary clinical report, sharing raw genomic data with healthcare providers is equally important, both for secondary research studies as well as for a deeper analysis of the data itself, as seen by the efforts from organizations such as American College of Medical Genetics and Genomics, as well as Global Alliance for Genomics and Health. Here, we aim to describe the existing protocol of genomic data sharing between a certified clinical laboratory and a healthcare provider and highlight some of the lessons learned. This study tracked and subsequently evaluated the data transfer workflow for 19 patients, all of whom consented to be part of this research study and visited the genetics clinic at a tertiary pediatric hospital between April 2016 and December 2016. (Full article...)
|
Featured article of the week: March 25–31:
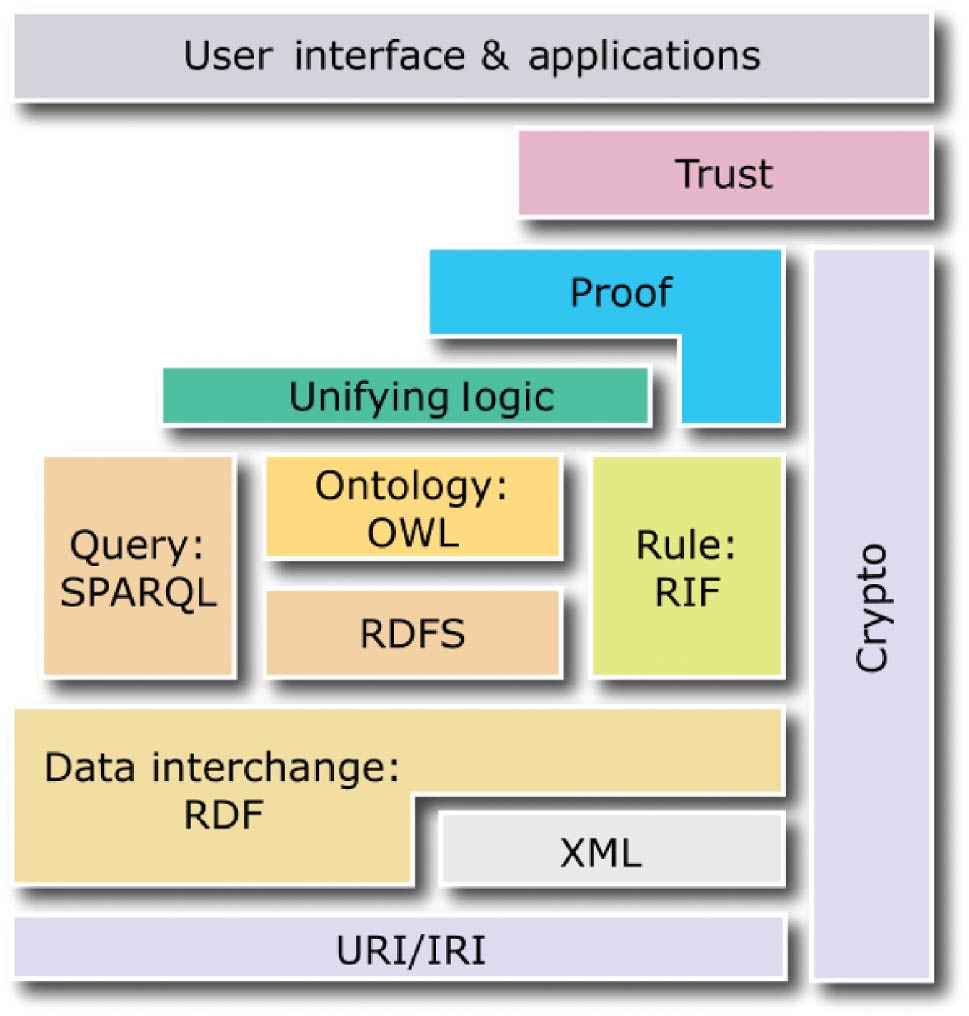
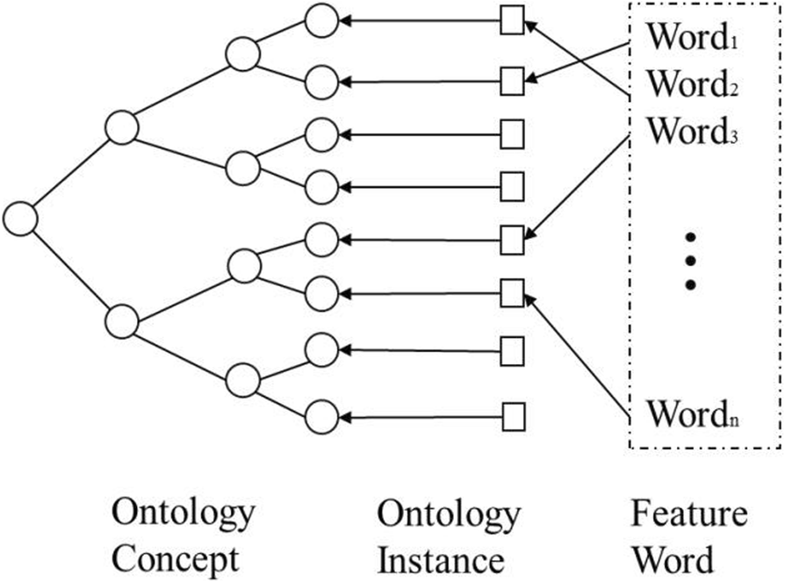
"Research on information retrieval model based on ontology"
An information retrieval system not only occupies an important position in the network information platform, but also plays an important role in information acquisition, query processing, and wireless sensor networks. It is a procedure to help researchers extract documents from data sets as document retrieval tools. The classic keyword-based information retrieval models neglect the semantic information which is not able to represent the user’s needs. Therefore, how to efficiently acquire personalized information that users need is of concern. The ontology-based systems lack an expert list to obtain accurate index term frequency. In this paper, a domain ontology model with document processing and document retrieval is proposed, and the feasibility and superiority of the domain ontology model are proved by the method of experiment. (Full article...)
|
Featured article of the week: March 18–24:
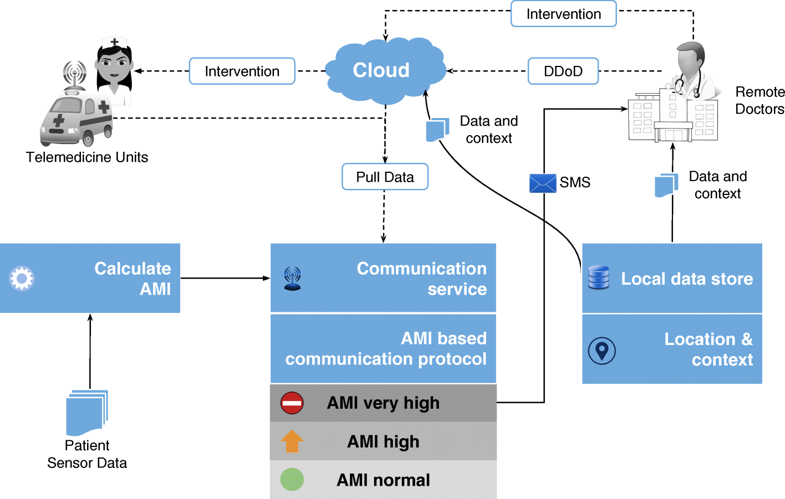
"Data to diagnosis in global health: A 3P approach"
With connected medical devices fast becoming ubiquitous in healthcare monitoring, there is a deluge of data coming from multiple body-attached sensors. Transforming this flood of data into effective and efficient diagnosis is a major challenge. To address this challenge, we present a "3P" approach: personalized patient monitoring, precision diagnostics, and preventive criticality alerts. In a collaborative work with doctors, we present the design, development, and testing of a healthcare data analytics and communication framework that we call RASPRO (Rapid Active Summarization for effective PROgnosis). The heart of RASPRO is "physician assist filters" (PAF) that 1. transform unwieldy multi-sensor time series data into summarized patient/disease-specific trends in steps of progressive precision as demanded by the doctor for a patient’s personalized condition, and 2. help in identifying and subsequently predictively alerting the onset of critical conditions. (Full article...)
|
Featured article of the week: March 11–17:
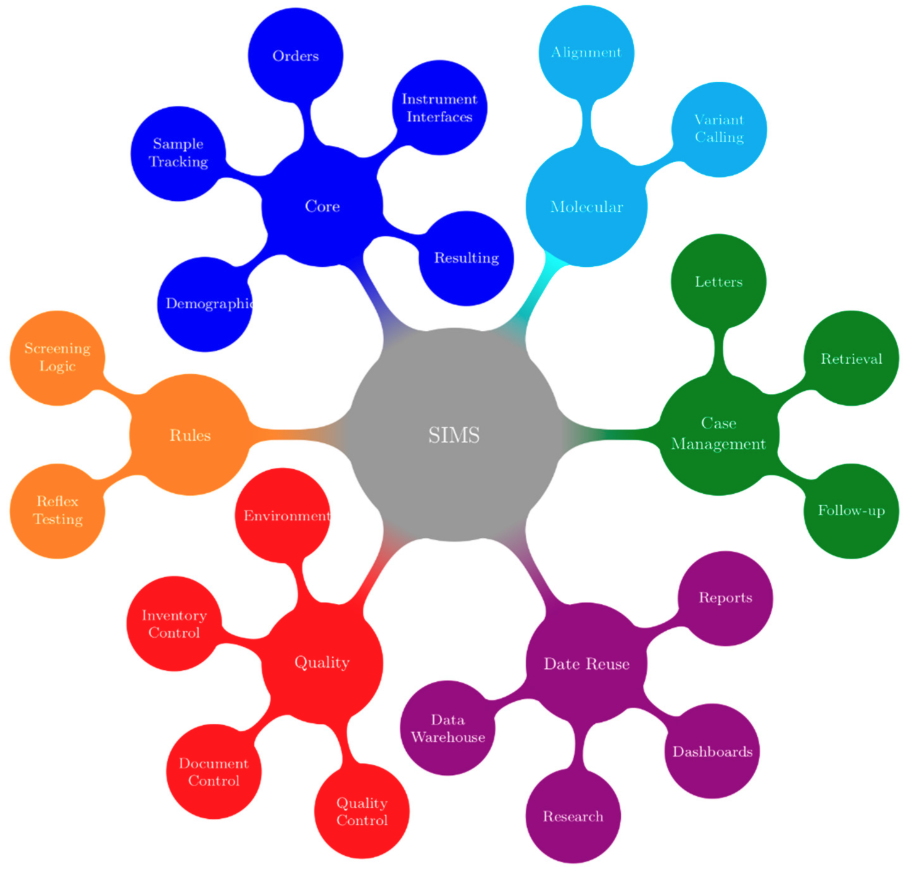
"Building a newborn screening information management system from theory to practice"
Information management systems are the central process management and communication hub for many newborn screening programs. In late 2014, Newborn Screening Ontario (NSO) undertook an end-to-end assessment of its information management needs, which resulted in a project to develop a flexible information systems (IS) ecosystem and related process changes. This enabled NSO to better manage its current and future workflow and communication needs. An idealized vision of a screening information management system (SIMS) was developed that was refined into enterprise and functional architectures. This was followed by the development of technical specifications, user requirements, and procurement. In undertaking a holistic full product lifecycle redesign approach, a number of change management challenges were faced by NSO across the entire program. Strong leadership support and full program engagement were key for overall project success. It is anticipated that improvements in program flexibility and the ability to innovate will outweigh the efforts and costs. (Full article...)
|
Featured article of the week: March 04–10:
"Adapting data management education to support clinical research projects in an academic medical center"
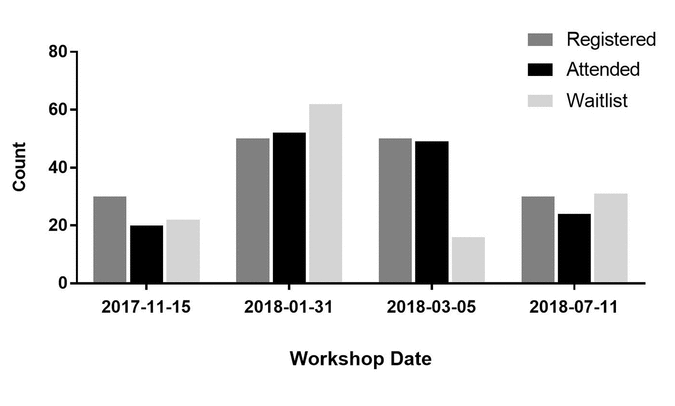
Librarians and researchers alike have long identified research data management (RDM) training as a need in biomedical research. Despite the wealth of libraries offering RDM education to their communities, clinical research is an area that has not been targeted. Clinical RDM (CRDM) is seen by its community as an essential part of the research process where established guidelines exist, yet educational initiatives in this area are unknown.
Leveraging my academic library’s experience supporting CRDM through informationist grants and REDCap training in our medical center, I developed a 1.5 hour CRDM workshop. This workshop was designed to use established CRDM guidelines in clinical research and address common questions asked by our community through the library’s existing data support program. The workshop was offered to the entire medical center four times between November 2017 and July 2018. This case study describes the development, implementation, and evaluation of this workshop. (Full article...)
|
Featured article of the week: February 25–March 03:
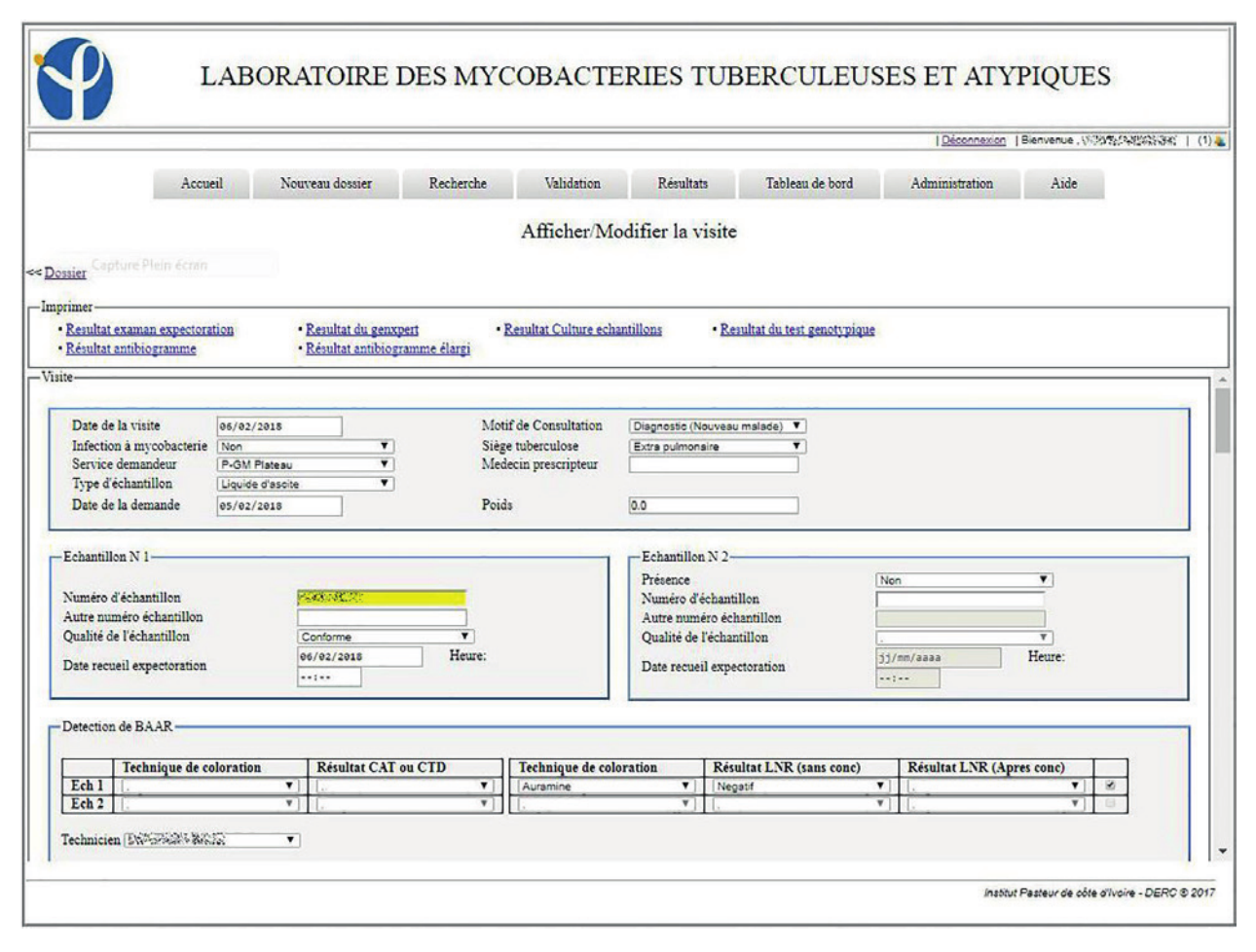
"Development of an electronic information system for the management of laboratory data of tuberculosis and atypical mycobacteria at the Pasteur Institute in Côte d’Ivoire"
Tuberculosis remains a public health problem despite all the efforts made to eradicate it. To strengthen the surveillance system for this condition, it is necessary to have a good data management system. Indeed, the use of electronic information systems in data management can improve the quality of data. The objective of this project was to set up a laboratory-specific electronic information system for mycobacteria and atypical tuberculosis.
The design of this laboratory information system required a general understanding of the workflow and the implementation processes in order to generate a realistic model. For the implementation of the system, Java technology was used to develop a web application compatible with the intranet of the company. (Full article...)
|
Featured article of the week: February 18–24:
"Codesign of the Population Health Information Management System to measure reach and practice change of childhood obesity programs"
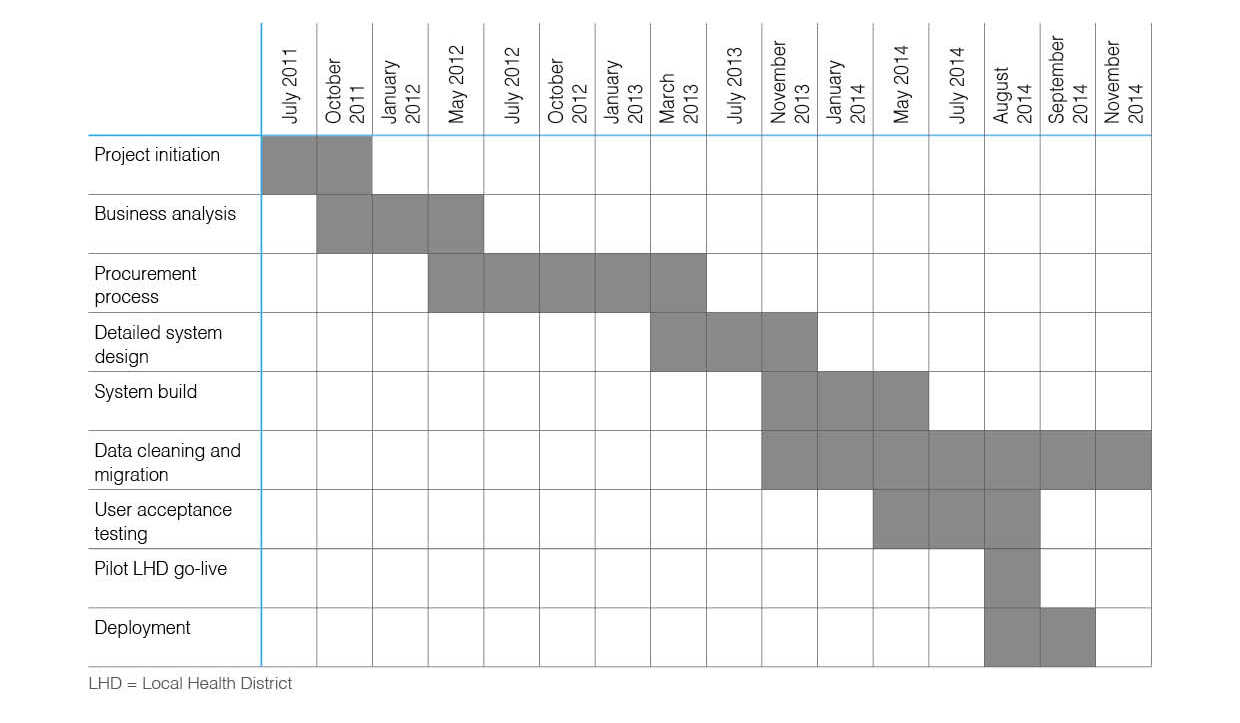
Childhood obesity prevalence is an issue of international public health concern, and governments have a significant role to play in its reduction. The Healthy Children Initiative (HCI) has been delivered in New South Wales (NSW), Australia, since 2011 to support implementation of childhood obesity prevention programs at scale. Consequently, a system to support local implementation and data collection, analysis, and reporting at local and state levels was necessary. The Population Health Information Management System (PHIMS) was developed to meet this need.
A collaborative and iterative process was applied to the design and development of the system. The process comprised identifying technical requirements, building system infrastructure, delivering training, deploying the system, and implementing quality measures. (Full article...)
|
Featured article of the week: February 11–17:
"Open data in scientific communication"
The development of information technology makes it possible to collect and analyze a growing number of data resources. The results of research, regardless of the discipline, constitute one of the main sources of data. Currently, research results are increasingly being published in the open access model. The open access concept has been accepted and recommended worldwide by many institutions financing and implementing research. Initially, the idea of openness concerned only the results of research and scientific publications; at present, more attention is paid to the problem of sharing scientific data, including raw data. Proceedings towards open data are intricate, as data specificity requires the development of an appropriate legal, technical and organizational model, followed by the implementation of data management policies at both the institutional and national levels. (Full article...)
|
Featured article of the week: February 4–10:
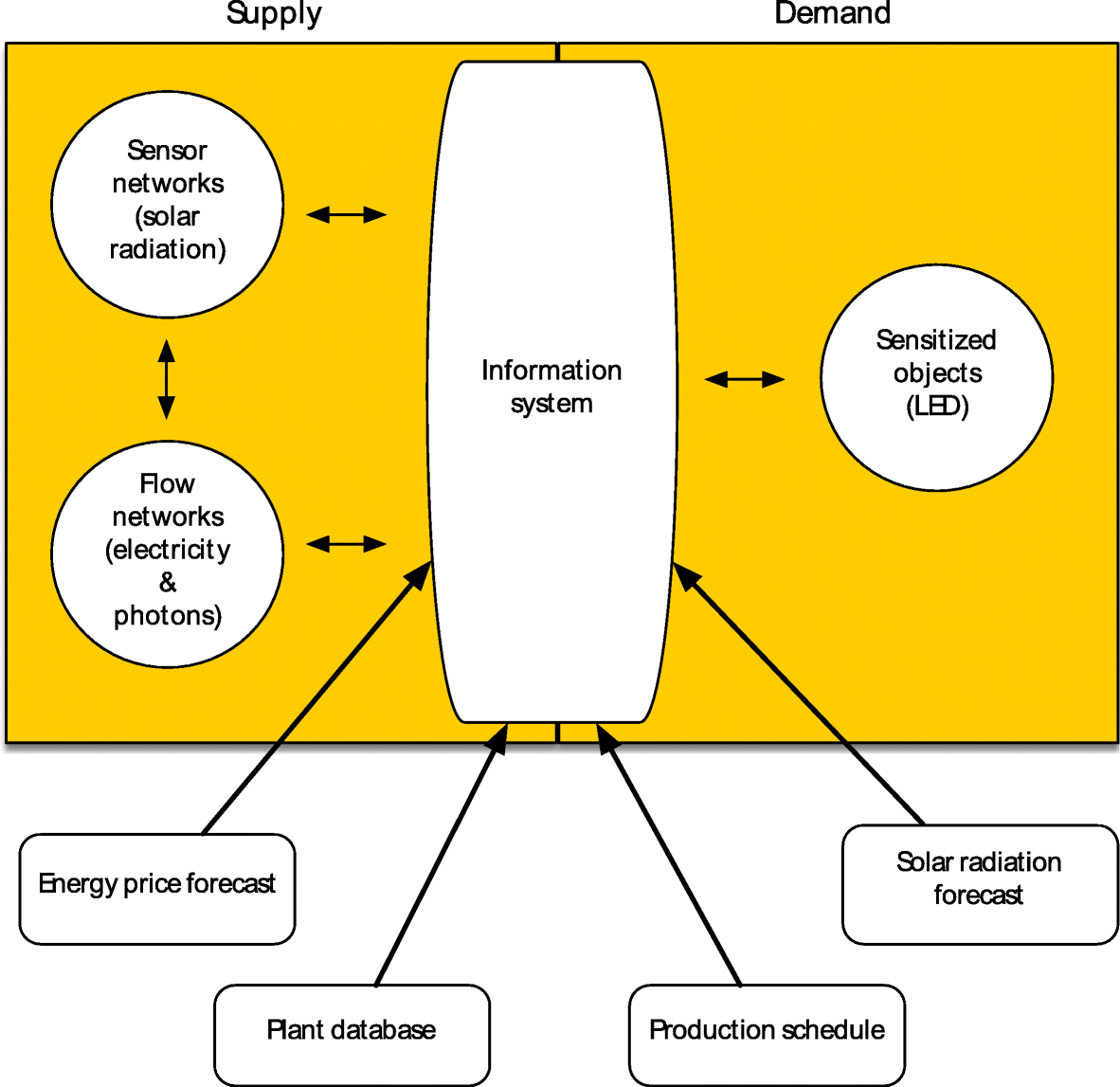
"Simulation of greenhouse energy use: An application of energy informatics"
Greenhouse agriculture is a highly efficient method of food production that can greatly benefit from supplemental electric lighting. The needed electricity associated with greenhouse lighting amounts to about 30% of its operating costs. As the light level of LED lighting can be easily controlled, it offers the potential to reduce energy costs by precisely matching the amount of supplemental light provided to current weather conditions and a crop’s light needs. Three simulations of LED lighting for growing lettuce in the Southeast U.S. using historical solar radiation data for the area were conducted. Lighting costs can be potentially reduced by approximately 60%. (Full article...)
|
Featured article of the week: January 28-February 3:
"Learning health systems need to bridge the "two cultures" of clinical informatics and data science"
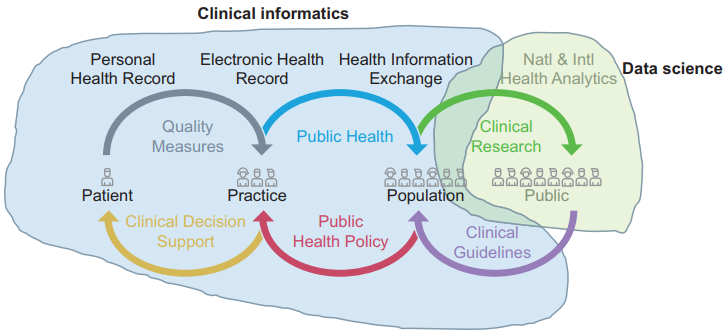
United Kingdom (U.K.) health research policy and plans for population health management are predicated upon transformative knowledge discovery from operational "big data." Learning health systems require not only data but also feedback loops of knowledge into changed practice. This depends on knowledge management and application, which in turn depends upon effective system design and implementation. Biomedical informatics is the interdisciplinary field at the intersection of health science, social science, and information science and technology that spans this entire scope.
In the U.K., the separate worlds of health data science (bioinformatics, big data) and effective healthcare system design and implementation (clinical informatics, "digital health") have operated as "two cultures." Much National Health Service and social care data is of very poor quality. Substantial research funding is wasted on data cleansing or by producing very weak evidence. There is not yet a sufficiently powerful professional community or evidence base of best practice to influence the practitioner community or the digital health industry. (Full article...)
|
Featured article of the week: January 21-27:
"The problem with dates: Applying ISO 8601 to research data management"
Dates appear regularly in research data and metadata but are a problematic data type to normalize due to a variety of potential formats. This suggests an opportunity for data librarians to assist with formatting dates, yet there are frequent examples of data librarians using diverse strategies for this purpose. Instead, data librarians should adopt the international date standard ISO 8601. This standard provides needed consistency in date formatting, allows for inclusion of several types of date-time information, and can sort dates chronologically. As regular advocates for standardization in research data, data librarians must adopt ISO 8601 and push for its use as a data management best practice.(Full article...)
|
Featured article of the week: January 14–20:
"Health sciences libraries advancing collaborative clinical research data management in universities"
Medical libraries need to actively review their service models and explore partnerships with other campus entities to provide better-coordinated clinical research management services to faculty and researchers. TRAIL (Translational Research and Information Lab), a five-partner initiative at the University of Washington (UW), explores how best to leverage existing expertise and space to deliver clinical research data management (CRDM) services and emerging technology support to clinical researchers at UW and collaborating institutions in the Pacific Northwest. The initiative offers 14 services and a technology-enhanced innovation lab located in the Health Sciences Library (HSL) to support the University of Washington clinical and research enterprise. Sharing of staff and resources merges library and non-library workflows, better coordinating data and innovation services to clinical researchers. Librarians have adopted new roles in CRDM, such as providing user support and training for UW’s Research Electronic Data Capture (REDCap) instance. (Full article...)
|
Featured article of the week: January 7–13:
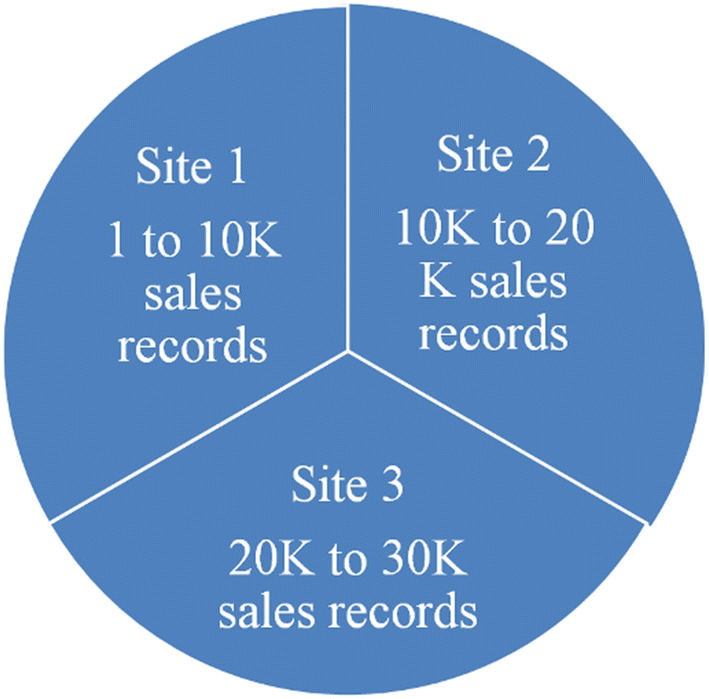
"Privacy preservation techniques in big data analytics: A survey"
Incredible amounts of data are being generated by various organizations like hospitals, banks, e-commerce, retail and supply chain, etc. by virtue of digital technology. Not only humans but also machines contribute to data streams in the form of closed circuit television (CCTV) streaming, web site logs, etc. Tons of data is generated every minute by social media and smart phones. The voluminous data generated from the various sources can be processed and analyzed to support decision making. However data analytics is prone to privacy violations. One of the applications of data analytics is recommendation systems, which are widely used by e-commerce sites like Amazon and Flipkart for suggesting products to customers based on their buying habits, leading to inference attacks. Although data analytics is useful in decision making, it will lead to serious privacy concerns. Hence privacy preserving data analytics became very important. This paper examines various privacy threats, privacy preservation techniques, and models with their limitations. The authors then propose a data lake-based modernistic privacy preservation technique to handle privacy preservation in unstructured data. (Full article...)
|
Featured article of the week: January 1–6:
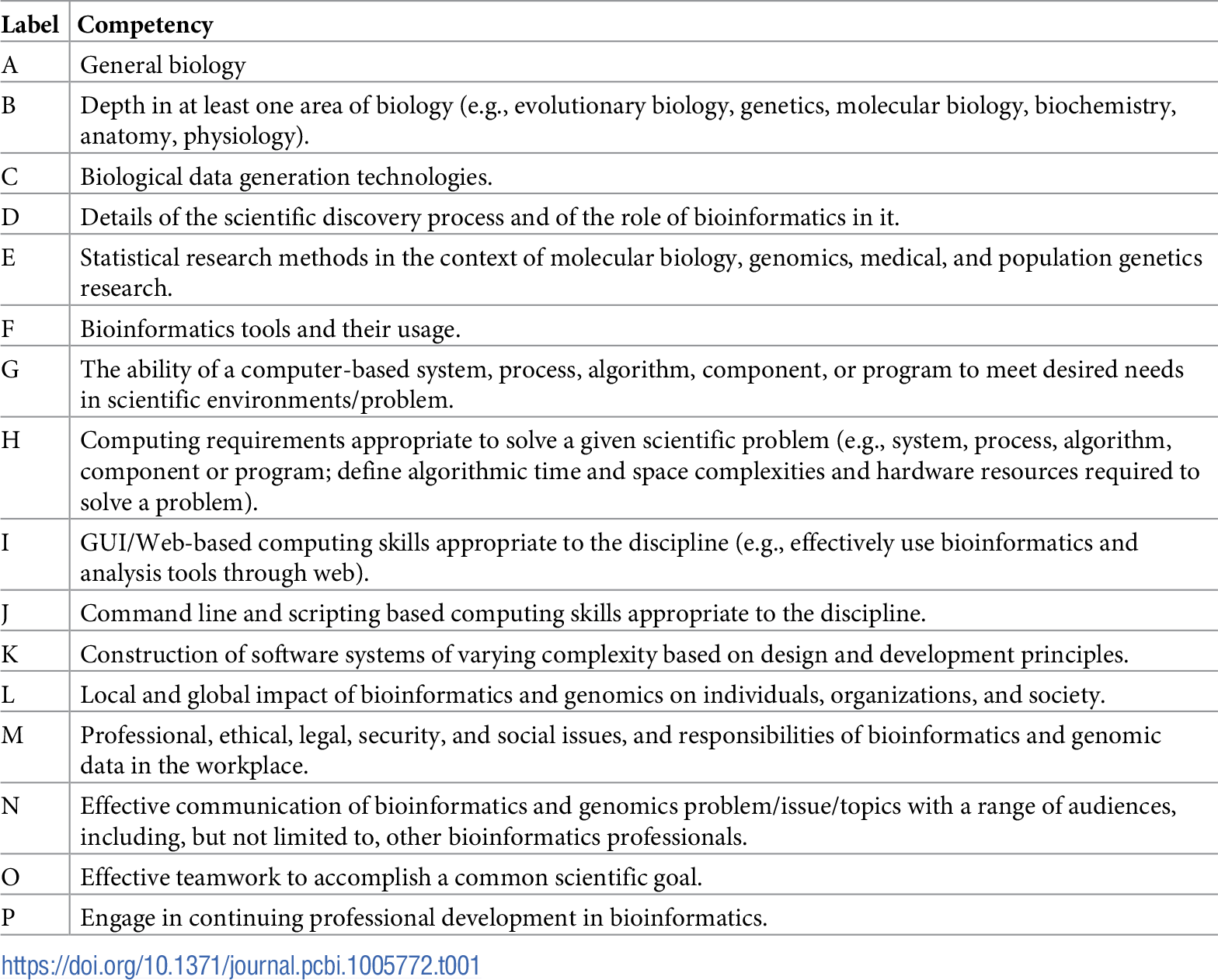
"The development and application of bioinformatics core competencies to improve bioinformatics training and education"
Bioinformatics is recognized as part of the essential knowledge base of numerous career paths in biomedical research and healthcare. However, there is little agreement in the field over what that knowledge entails or how best to provide it. These disagreements are compounded by the wide range of populations in need of bioinformatics training, with divergent prior backgrounds and intended application areas. The Curriculum Task Force of the International Society of Computational Biology (ISCB) Education Committee has sought to provide a framework for training needs and curricula in terms of a set of bioinformatics core competencies that cut across many user personas and training programs. The initial competencies developed based on surveys of employers and training programs have since been refined through a multiyear process of community engagement. This report describes the current status of the competencies and presents a series of use cases illustrating how they are being applied in diverse training contexts. (Full article...)
|
|