Difference between revisions of "Main Page/Featured article of the week/2021"
Shawndouglas (talk | contribs) (Added prior week's article of the week) |
Shawndouglas (talk | contribs) (Added last week's article of the week) |
||
| (7 intermediate revisions by the same user not shown) | |||
| Line 17: | Line 17: | ||
<!-- Below this line begin pasting previous news --> | <!-- Below this line begin pasting previous news --> | ||
<h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: July 19–25:</h2> | <h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: September 13–19:</h2> | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Navale F1000Research2020 8.gif|240px]]</div> | |||
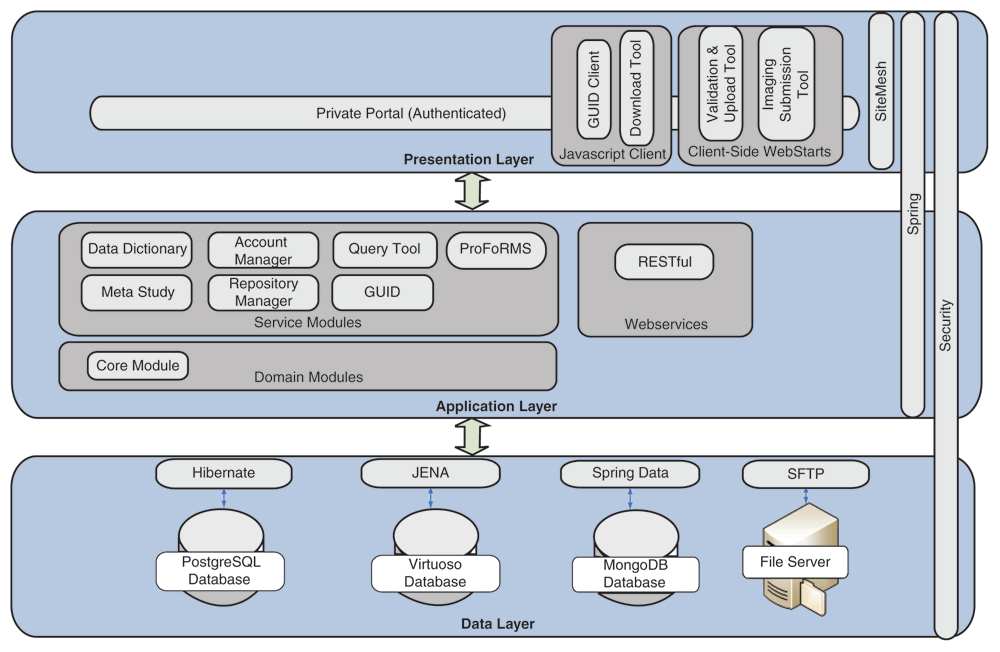
'''"[[Journal:Development of an informatics system for accelerating biomedical research|Development of an informatics system for accelerating biomedical research]]"''' | |||
The Biomedical Research Informatics Computing System (BRICS) was developed to support multiple disease-focused research programs. Seven service modules are integrated together to provide a collaborative and extensible web-based environment. The modules—Data Dictionary, Account Management, Query Tool, Protocol and Form Research Management System, Meta Study, Data Repository, and Globally Unique Identifier—facilitate the management of research protocols, including the submission, processing, curation, access, and storage of clinical, imaging, and derived [[genomics]] data within the associated data repositories. Multiple instances of BRICS are deployed to support various biomedical research communities focused on accelerating discoveries for rare diseases, traumatic brain injuries, Parkinson’s disease, inherited eye diseases, and symptom science research. No personally identifiable [[information]] is stored within the data repositories. Digital object identifiers (DOIs) are associated with the research studies. ('''[[Journal:Development of an informatics system for accelerating biomedical research|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: September 6–12:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Cicek FrontPubHealth2020 8.jpg|240px]]</div> | |||
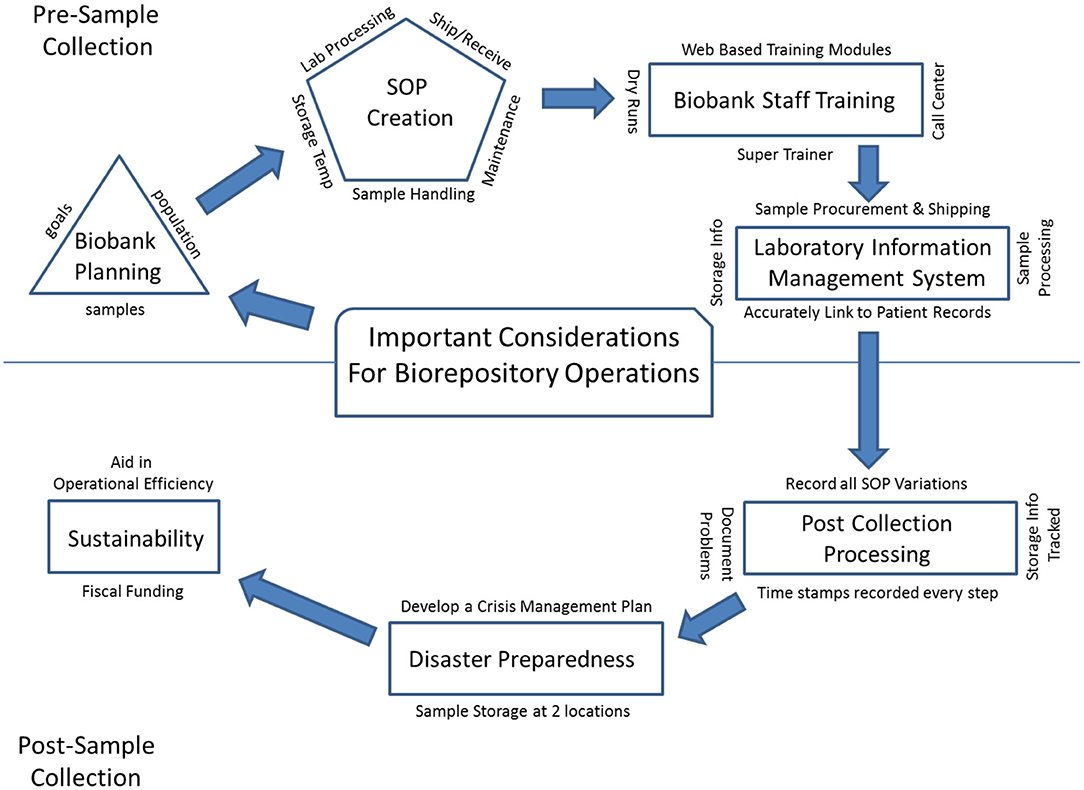
'''"[[Journal:Mini-review of laboratory operations in biobanking: Building biobanking resources for translational research|Mini-review of laboratory operations in biobanking: Building biobanking resources for translational research]]"''' | |||
[[Biobank]]s have become integral to improving [[Public health|population health]]. We are in a new era in medicine as patients, health professionals, and researchers increasingly collaborate to gain new knowledge and explore new paradigms for diagnosing and treating disease. Many large-scale biobanking efforts are underway worldwide at the institutional, national, and even international level. When linked with subject data from questionnaires and medical records, biobanks serve as valuable resources in [[translational research]]. A biobank must have high-quality biospecimens that meet researcher's needs. Biobank [[laboratory]] operations require an enormous amount of support, from lab and storage space, information technology expertise, and a [[laboratory information management system]] to logistics for biospecimen tracking, [[quality management system]]s, and appropriate facilities. ('''[[Journal:Mini-review of laboratory operations in biobanking: Building biobanking resources for translational research|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: August 30–September 5:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig3 Dixon BMJHealthCareInfo2020 27-1.png|240px]]</div> | |||
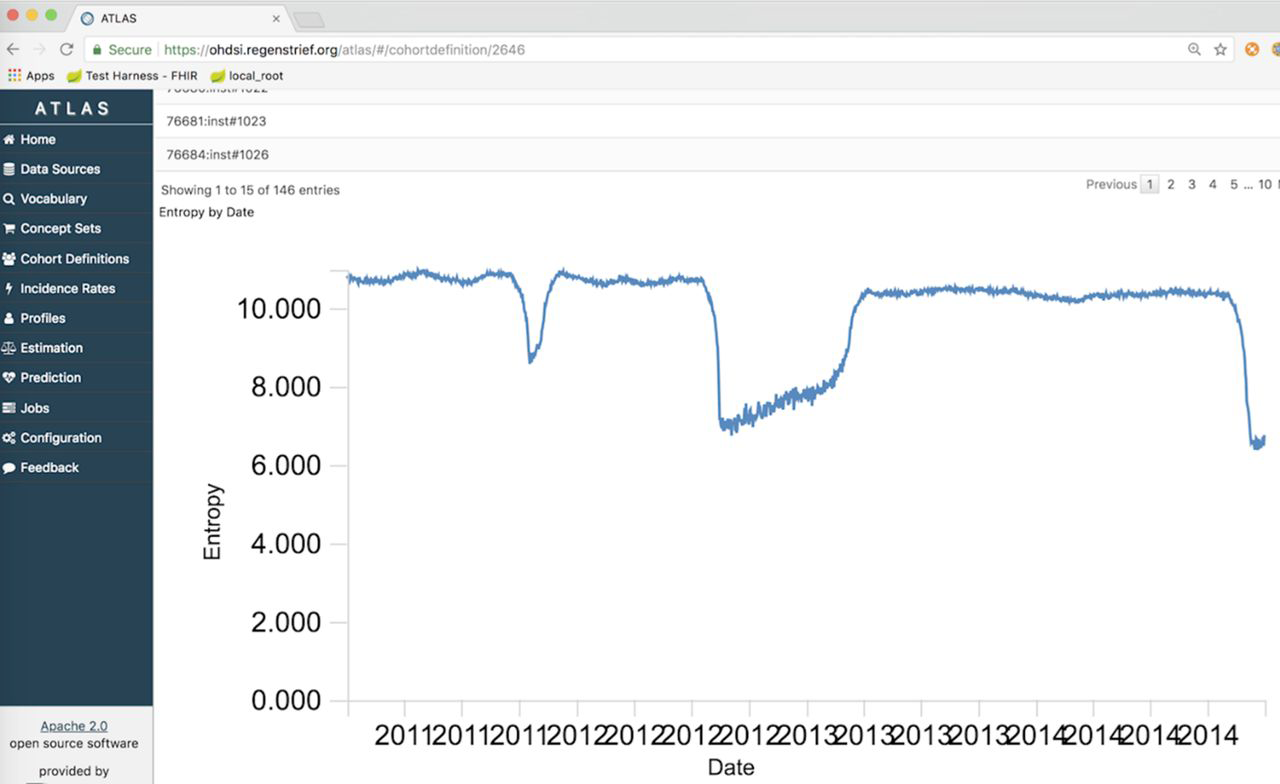
'''"[[Journal:Extending an open-source tool to measure data quality: Case report on Observational Health Data Science and Informatics (OHDSI)|Extending an open-source tool to measure data quality: Case report on Observational Health Data Science and Informatics (OHDSI)]]"''' | |||
As the health system seeks to leverage large-scale data to inform population outcomes, the [[Informatics (academic field)|informatics]] community is developing tools for analyzing these data. To support [[data quality]] assessment within such a tool, we extended the open-source software Observational Health Data Sciences and Informatics (OHDSI) to incorporate new functions useful for population health. We developed and tested methods to measure the completeness, timeliness, and entropy of [[information]]. The new data quality methods were applied to over 100 million clinical messages received from emergency department information systems for use in [[Public health informatics|public health syndromic surveillance systems]]. ('''[[Journal:Extending an open-source tool to measure data quality: Case report on Observational Health Data Science and Informatics (OHDSI)|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: August 23–29:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Raymond BMCMedInfoDecMak2020 20.png|240px]]</div> | |||
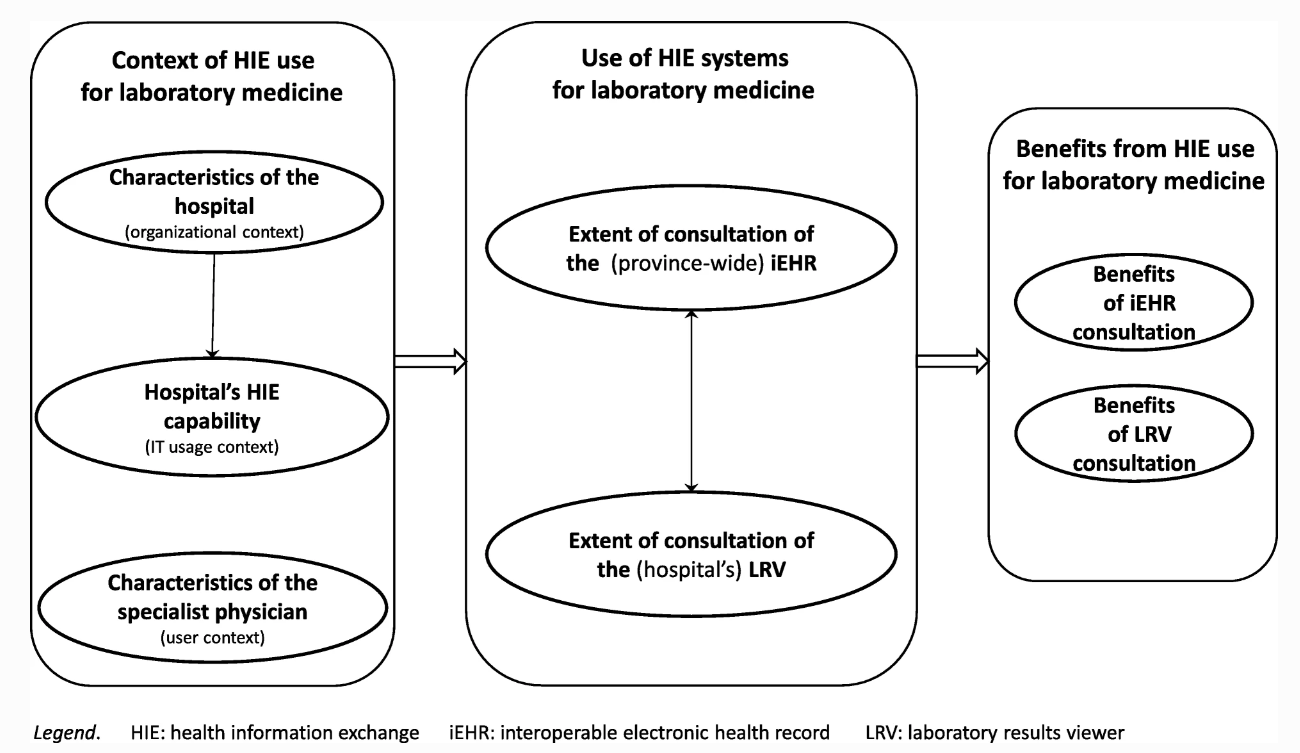
'''"[[Journal:Advancing laboratory medicine in hospitals through health information exchange: A survey of specialist physicians in Canada|Advancing laboratory medicine in hospitals through health information exchange: A survey of specialist physicians in Canada]]"''' | |||
[[Laboratory]] testing occupies a prominent place in healthcare. Information technology systems have the potential to empower laboratory experts and to enhance the interpretation of test results in order to better support physicians in their quest for better and safer patient care. This study sought to develop a better understanding of which laboratory information exchange (LIE) systems and features specialist physicians are using in [[hospital]] settings to consult their patients’ laboratory test results, and what benefit they derive from such use. As part of a broader research program on the use of [[health information exchange]] systems for laboratory medicine in Quebec, Canada, this study was designed as on online survey. Our sample is composed of 566 specialist physicians working in hospital settings, out of the 1,512 physicians who responded to the survey (response rate of 17%). Respondents are representative of the targeted population of specialist physicians in terms of gender, age, and hospital location. ('''[[Journal:Advancing laboratory medicine in hospitals through health information exchange: A survey of specialist physicians in Canada|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: August 16–22:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig3 Krill Metabolites2020 10-7.png|240px]]</div> | |||
'''"[[Journal:A high-throughput method for the comprehensive analysis of terpenes and terpenoids in medicinal cannabis biomass|A high-throughput method for the comprehensive analysis of terpenes and terpenoids in medicinal cannabis biomass]]"''' | |||
[[wikipedia:Cannabis|Cannabis]] and its secondary metabolite content have recently seen a surge in research interest. Cannabis [[wikipedia:Terpene|terpenes]] and [[wikipedia:Terpenoid|terpenoids]] in particular are increasingly the focus of research efforts due to the possibility of their contribution to the overall therapeutic effect of [[wikipedia:Medical cannabis|medicinal cannabis]]. Current methodology to quantify terpenes in cannabis biomass mostly relies on large quantities of biomass, long extraction protocols, and long [[gas chromatography]] (GC) gradient times, often exceeding 60 minutes. They are therefore not easily applicable in the high-throughput environment of a cannabis breeding program. ('''[[Journal:A high-throughput method for the comprehensive analysis of terpenes and terpenoids in medicinal cannabis biomass|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: August 9–15:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Arendt ClinEpidem2020 12.jpg|240px]]</div> | |||
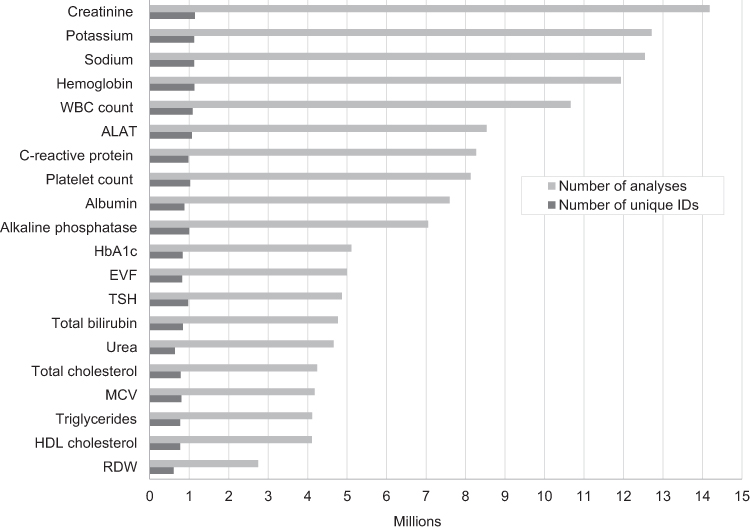
'''"[[Journal:Existing data sources in clinical epidemiology: Laboratory information system databases in Denmark|Existing data sources in clinical epidemiology: Laboratory information system databases in Denmark]]"''' | |||
Routine [[biomarker]] results from [[hospital]] [[laboratory information system]]s (LIS)—covering hospitals and general practitioners—in Denmark are available to researchers through access to the regional Clinical Laboratory Information System Research Database at Aarhus University and the nationwide Register of Laboratory Results for Research. This review describes these two data sources. The [[laboratory]] databases have different geographical and temporal coverage. They both include individual-level biomarker results that are electronically transferred from LISs. The biomarker results can be linked to all other Danish registries at the individual level using the unique identifier, the CPR number. ('''[[Journal:Existing data sources in clinical epidemiology: Laboratory information system databases in Denmark|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: August 2–8:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig2 Heinen BMCBioinfo2020 21.png|240px]]</div> | |||
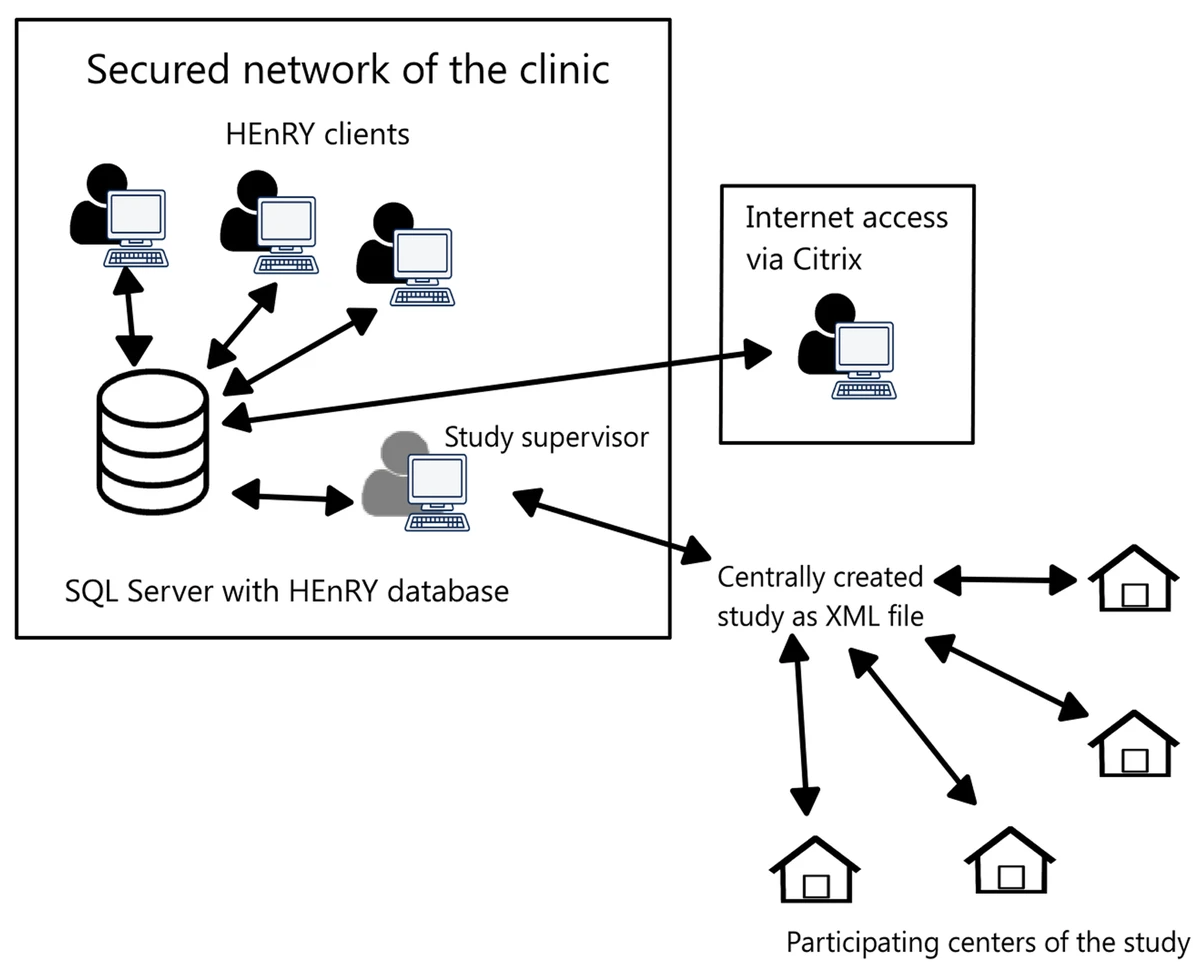
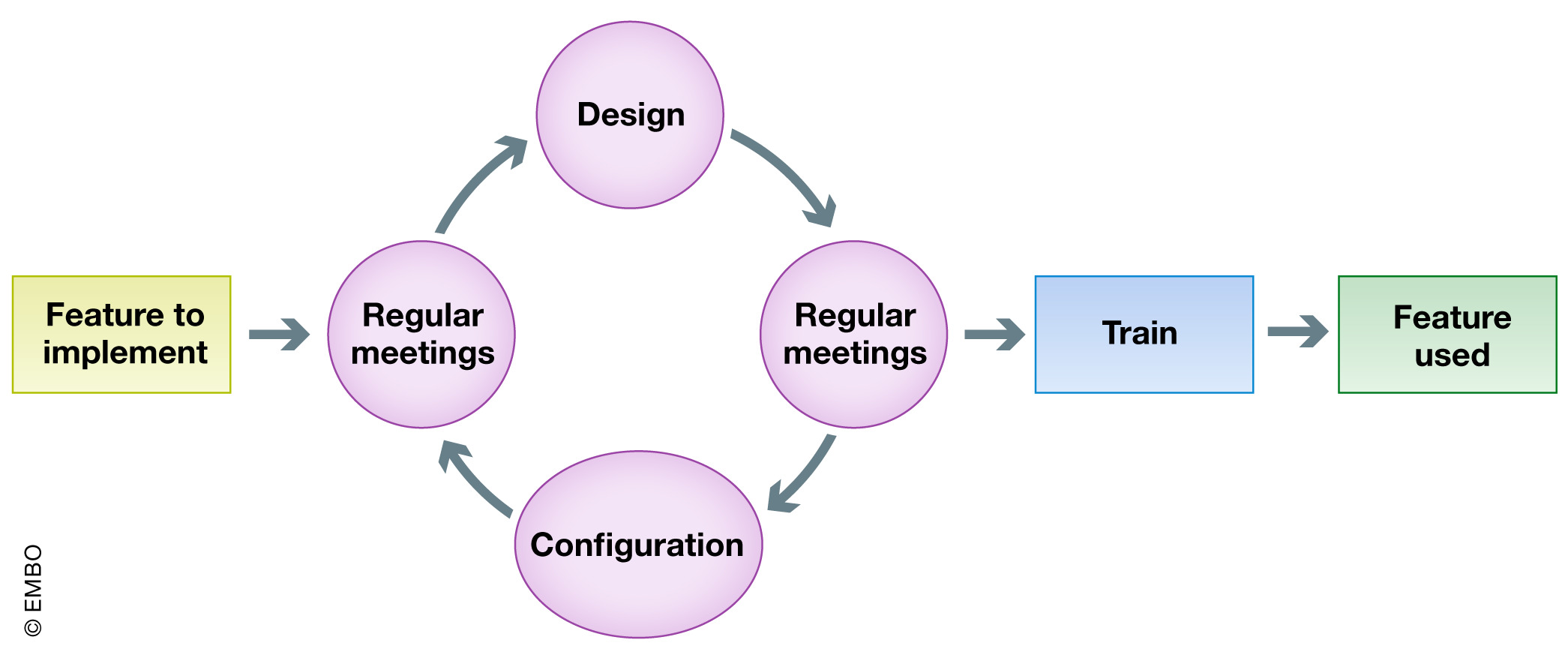
'''"[[Journal:HEnRY: A DZIF LIMS tool for the collection and documentation of biospecimens in multicentre studies|HEnRY: A DZIF LIMS tool for the collection and documentation of biospecimens in multicentre studies]]"''' | |||
Well-characterized biological specimens (biospecimens) of high quality have great potential for the acceleration of and quality improvement in [[Translational research|translational biomedical research]]. To improve accessibility of local [[Sample (material)|specimen]] collections, efforts have been made to create central repositories ([[biobank]]s) and catalogues. Available technical solutions for creating professional local specimen catalogues and connecting them to central systems are cost intensive and/or technically complex to implement. Therefore, the HIV-focused Thematic Translational Unit (TTU) of the German Center for Infection Research (DZIF) developed a [[laboratory information management system]] (LIMS) called HIV Engaged Research Technology (HEnRY) for implementation into the HIV Translational Platform (TP-HIV) at the DZIF and other research networks. ('''[[Journal:HEnRY: A DZIF LIMS tool for the collection and documentation of biospecimens in multicentre studies|Full article...]]''')<br /> | |||
<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: July 26–August 1:</h2> | |||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Tab1 Comess FrontArtInt2020 3.jpg|240px]]</div> | |||
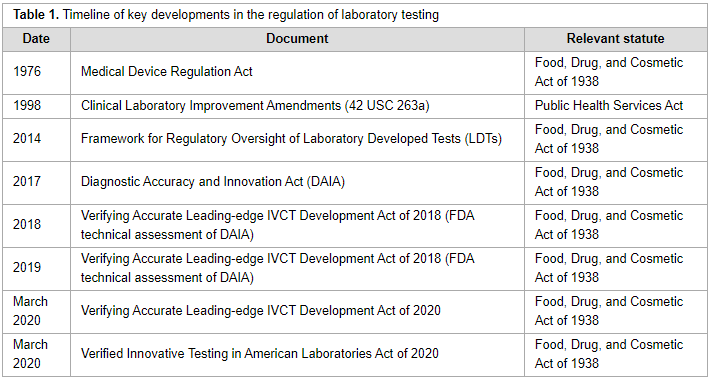
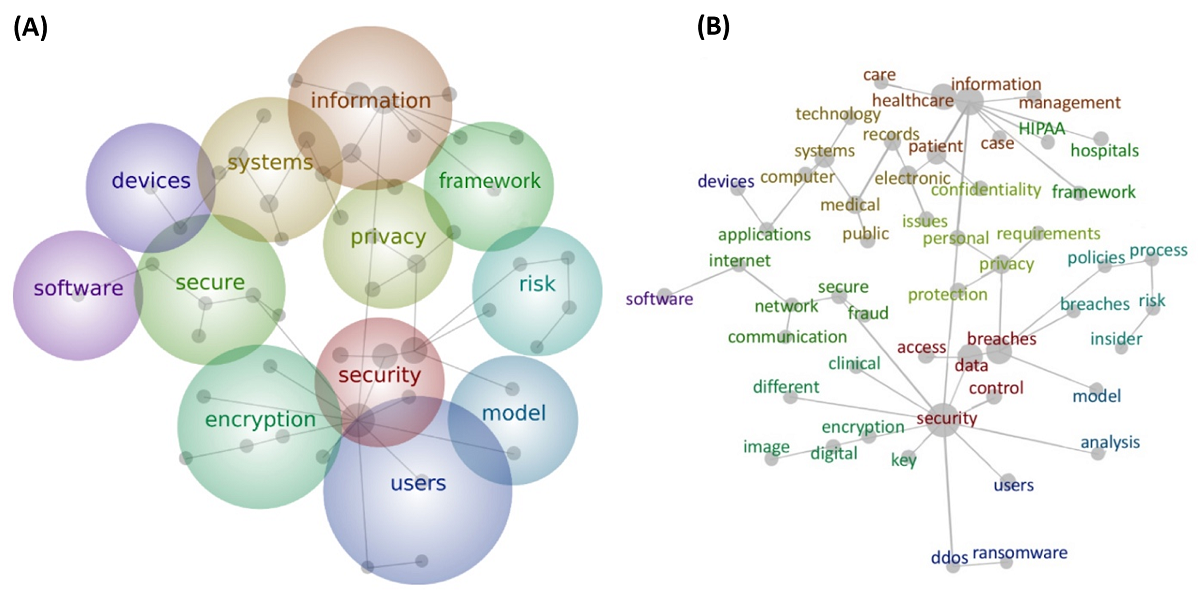
'''"[[Journal:Bringing big data to bear in environmental public health: Challenges and recommendations|Bringing big data to bear in environmental public health: Challenges and recommendations]]"''' | |||
Understanding the role that the environment plays in influencing [[public health]] often involves collecting and studying large, complex data sets. There have been a number of private and public efforts to gather sufficient [[information]] and confront significant unknowns in the field of [[Environmental health|environmental public health]], yet there is a persistent and largely unmet need for findable, accessible, interoperable, and reusable [[Journal:The FAIR Guiding Principles for scientific data management and stewardship|(FAIR) data]]. Even when data are readily available, the ability to create, [[Data analysis|analyze]], and draw conclusions from these data using emerging computational tools, such as augmented intelligence, [[artificial intelligence]] (AI), and [[machine learning]], requires technical skills not currently implemented on a programmatic level across research hubs and academic institutions. ('''[[Journal:Bringing big data to bear in environmental public health: Challenges and recommendations|Full article...]]''')<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: July 19–25:</h2> | |||
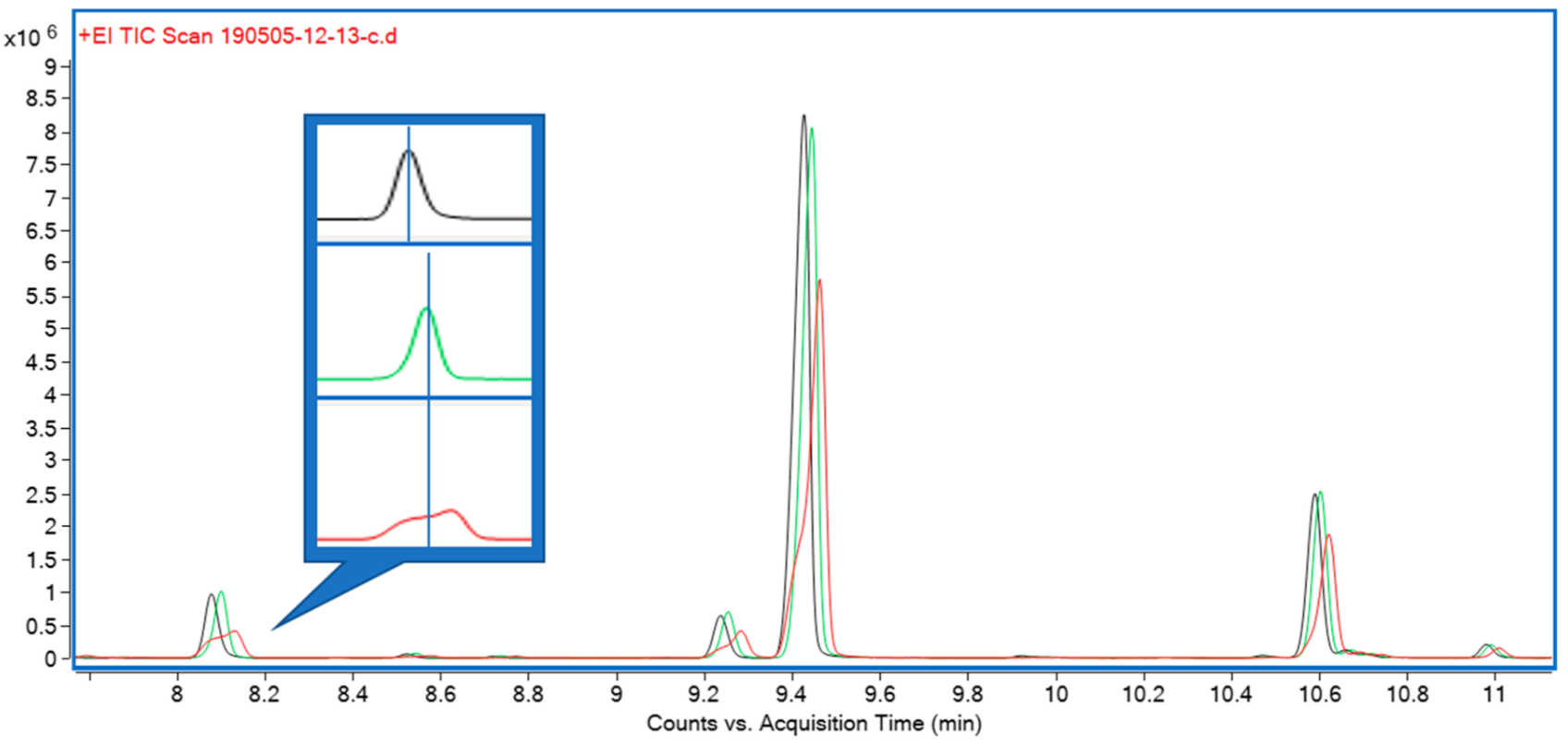
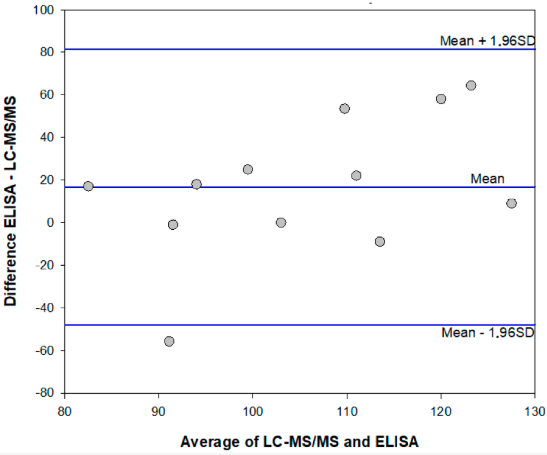
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig2 DiNardo Toxins2020 12-4.png|240px]]</div> | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig2 DiNardo Toxins2020 12-4.png|240px]]</div> | ||
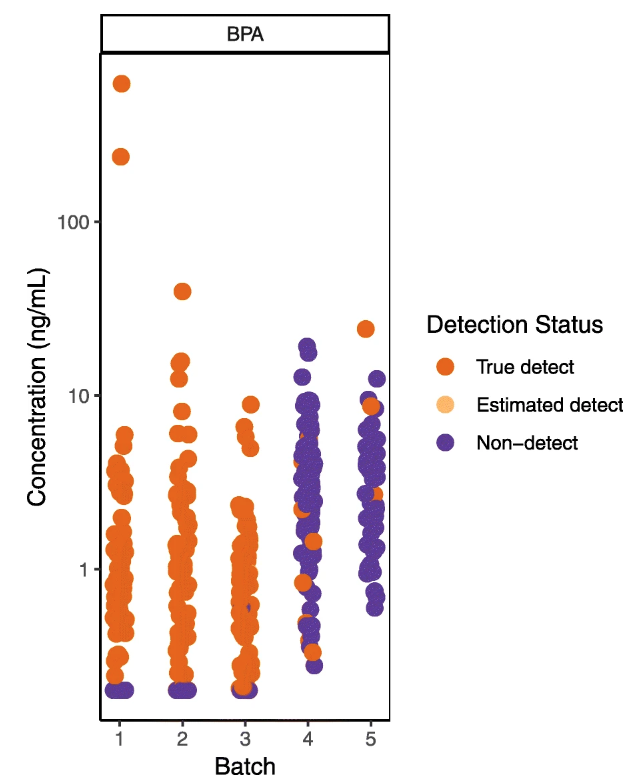
'''"[[Journal:Enzyme immunoassay for measuring aflatoxin B1 in legal cannabis|Enzyme immunoassay for measuring aflatoxin B1 in legal cannabis]]"''' | '''"[[Journal:Enzyme immunoassay for measuring aflatoxin B1 in legal cannabis|Enzyme immunoassay for measuring aflatoxin B1 in legal cannabis]]"''' | ||
Revision as of 17:50, 21 September 2021
|
|
If you're looking for other "Article of the Week" archives: 2014 - 2015 - 2016 - 2017 - 2018 - 2019 - 2020 - 2021 |
Featured article of the week archive - 2021
Welcome to the LIMSwiki 2021 archive for the Featured Article of the Week.
Featured article of the week: September 13–19:"Development of an informatics system for accelerating biomedical research" The Biomedical Research Informatics Computing System (BRICS) was developed to support multiple disease-focused research programs. Seven service modules are integrated together to provide a collaborative and extensible web-based environment. The modules—Data Dictionary, Account Management, Query Tool, Protocol and Form Research Management System, Meta Study, Data Repository, and Globally Unique Identifier—facilitate the management of research protocols, including the submission, processing, curation, access, and storage of clinical, imaging, and derived genomics data within the associated data repositories. Multiple instances of BRICS are deployed to support various biomedical research communities focused on accelerating discoveries for rare diseases, traumatic brain injuries, Parkinson’s disease, inherited eye diseases, and symptom science research. No personally identifiable information is stored within the data repositories. Digital object identifiers (DOIs) are associated with the research studies. (Full article...)
|