Difference between revisions of "Template:Article of the week"
Shawndouglas (talk | contribs) (Updated article of the week text) |
Shawndouglas (talk | contribs) (Updated article of the week text) |
||
| Line 1: | Line 1: | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File: | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Parker DataSciJourn2019 18-1.png|240px]]</div> | ||
'''"[[Journal: | '''"[[Journal:Building infrastructure for African human genomic data management|Building infrastructure for African human genomic data management]]"''' | ||
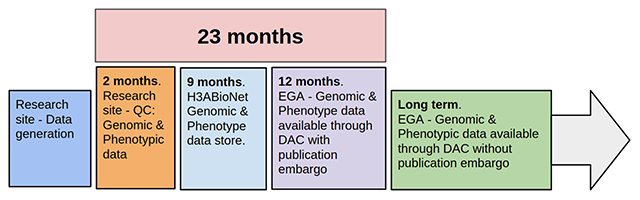
[[ | Human [[Genomics|genomic]] data are large and complex, and require adequate infrastructure for secure storage and transfer. The [[National Institutes of Health]] (NIH) and The Wellcome Trust have funded multiple projects on genomic research, including the Human Heredity and Health in Africa (H3Africa) initiative, and data are required to be deposited into the public domain. The European Genome-phenome Archive (EGA) is a repository for [[Sequencing|sequence]] and genotype data where data access is controlled by access committees. Access is determined by a formal application procedure for the purpose of secure storage and distribution, which must be in line with the informed consent of the study participants. H3Africa researchers based in Africa and generating their own data can benefit tremendously from the data sharing capabilities of the internet by using the appropriate technologies. The H3Africa Data Archive is an effort between the H3Africa data generating projects, H3ABioNet, and the EGA to store and submit genomic data to public repositories. ('''[[Journal:Building infrastructure for African human genomic data management|Full article...]]''')<br /> | ||
<br /> | <br /> | ||
''Recently featured'': | ''Recently featured'': | ||
: ▪ [[Journal:Process variation detection using missing data in a multihospital community practice anatomic pathology laboratory|Process variation detection using missing data in a multihospital community practice anatomic pathology laboratory]] | |||
: ▪ [[Journal:Development and validation of a fast gas chromatography–mass spectrometry method for the determination of cannabinoids in Cannabis sativa L|Development and validation of a fast gas chromatography–mass spectrometry method for the determination of cannabinoids in Cannabis sativa L]] | : ▪ [[Journal:Development and validation of a fast gas chromatography–mass spectrometry method for the determination of cannabinoids in Cannabis sativa L|Development and validation of a fast gas chromatography–mass spectrometry method for the determination of cannabinoids in Cannabis sativa L]] | ||
: ▪ [[Journal:Design and refinement of a data quality assessment workflow for a large pediatric research network|Design and refinement of a data quality assessment workflow for a large pediatric research network]] | : ▪ [[Journal:Design and refinement of a data quality assessment workflow for a large pediatric research network|Design and refinement of a data quality assessment workflow for a large pediatric research network]] | ||
Revision as of 19:26, 23 December 2019
"Building infrastructure for African human genomic data management"
Human genomic data are large and complex, and require adequate infrastructure for secure storage and transfer. The National Institutes of Health (NIH) and The Wellcome Trust have funded multiple projects on genomic research, including the Human Heredity and Health in Africa (H3Africa) initiative, and data are required to be deposited into the public domain. The European Genome-phenome Archive (EGA) is a repository for sequence and genotype data where data access is controlled by access committees. Access is determined by a formal application procedure for the purpose of secure storage and distribution, which must be in line with the informed consent of the study participants. H3Africa researchers based in Africa and generating their own data can benefit tremendously from the data sharing capabilities of the internet by using the appropriate technologies. The H3Africa Data Archive is an effort between the H3Africa data generating projects, H3ABioNet, and the EGA to store and submit genomic data to public repositories. (Full article...)
Recently featured:
- ▪ Process variation detection using missing data in a multihospital community practice anatomic pathology laboratory
- ▪ Development and validation of a fast gas chromatography–mass spectrometry method for the determination of cannabinoids in Cannabis sativa L
- ▪ Design and refinement of a data quality assessment workflow for a large pediatric research network