Difference between revisions of "Template:Article of the week"
Shawndouglas (talk | contribs) (Updated article of the week text.) |
Shawndouglas (talk | contribs) (Updated article of the week text.) |
||
| Line 1: | Line 1: | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Bianchi FrontinGenetics2016 7.jpg|240px]]</div> | ||
'''"[[Journal: | '''"[[Journal:Integrated systems for NGS data management and analysis: Open issues and available solutions|Integrated systems for NGS data management and analysis: Open issues and available solutions]]"''' | ||
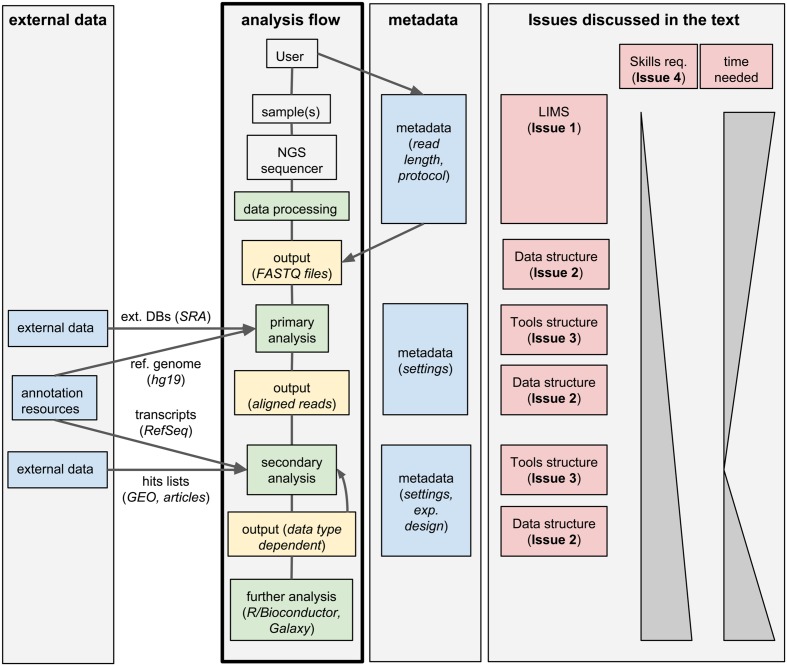
Next-generation sequencing (NGS) technologies have deeply changed our understanding of cellular processes by delivering an astonishing amount of data at affordable prices; nowadays, many biology [[laboratory|laboratories]] have already accumulated a large number of sequenced samples. However, managing and analyzing these data poses new challenges, which may easily be underestimated by research groups devoid of IT and quantitative skills. In this perspective, we identify five issues that should be carefully addressed by research groups approaching NGS technologies. In particular, the five key issues to be considered concern: (1) adopting a [[laboratory information management system]] (LIMS) and safeguard the resulting raw data structure in downstream analyses; (2) monitoring the flow of the data and standardizing input and output directories and file names, even when multiple analysis protocols are used on the same data; (3) ensuring complete traceability of the analysis performed; (4) enabling non-experienced users to run analyses through a graphical user interface (GUI) acting as a front-end for the pipelines; (5) relying on standard metadata to annotate the datasets, and when possible using controlled vocabularies, ideally derived from biomedical ontologies. ('''[[Journal:Integrated systems for NGS data management and analysis: Open issues and available solutions|Full article...]]''')<br /> | |||
<br /> | <br /> | ||
''Recently featured'': | ''Recently featured'': | ||
: ▪ [[Journal:Practical approaches for mining frequent patterns in molecular datasets|Practical approaches for mining frequent patterns in molecular datasets]] | |||
: ▪ [[Journal:Improving the creation and reporting of structured findings during digital pathology review|Improving the creation and reporting of structured findings during digital pathology review]] | : ▪ [[Journal:Improving the creation and reporting of structured findings during digital pathology review|Improving the creation and reporting of structured findings during digital pathology review]] | ||
: ▪ [[Journal:The challenges of data quality and data quality assessment in the big data era|The challenges of data quality and data quality assessment in the big data era]] | : ▪ [[Journal:The challenges of data quality and data quality assessment in the big data era|The challenges of data quality and data quality assessment in the big data era]] | ||
Revision as of 16:23, 31 October 2016
"Integrated systems for NGS data management and analysis: Open issues and available solutions"
Next-generation sequencing (NGS) technologies have deeply changed our understanding of cellular processes by delivering an astonishing amount of data at affordable prices; nowadays, many biology laboratories have already accumulated a large number of sequenced samples. However, managing and analyzing these data poses new challenges, which may easily be underestimated by research groups devoid of IT and quantitative skills. In this perspective, we identify five issues that should be carefully addressed by research groups approaching NGS technologies. In particular, the five key issues to be considered concern: (1) adopting a laboratory information management system (LIMS) and safeguard the resulting raw data structure in downstream analyses; (2) monitoring the flow of the data and standardizing input and output directories and file names, even when multiple analysis protocols are used on the same data; (3) ensuring complete traceability of the analysis performed; (4) enabling non-experienced users to run analyses through a graphical user interface (GUI) acting as a front-end for the pipelines; (5) relying on standard metadata to annotate the datasets, and when possible using controlled vocabularies, ideally derived from biomedical ontologies. (Full article...)
Recently featured: