Difference between revisions of "Template:Article of the week"
Shawndouglas (talk | contribs) (Updated article of the week text.) |
Shawndouglas (talk | contribs) (Updated article of the week text.) |
||
| Line 1: | Line 1: | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File: | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig2 Tsai JofPersMed2016 6-1.png|80px]]</div> | ||
'''"[[Journal: | '''"[[Journal:Bioinformatics workflow for clinical whole genome sequencing at Partners HealthCare Personalized Medicine|Bioinformatics workflow for clinical whole genome sequencing at Partners HealthCare Personalized Medicine]]"''' | ||
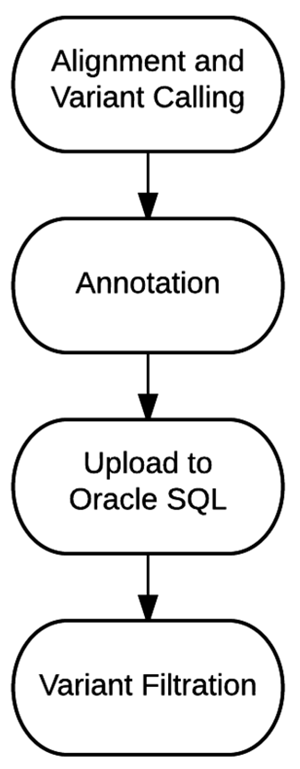
Effective implementation of precision medicine will be enhanced by a thorough understanding of each patient’s genetic composition to better treat his or her presenting symptoms or mitigate the onset of disease. This ideally includes the sequence [[information]] of a complete genome for each individual. At Partners HealthCare Personalized Medicine, we have developed a clinical process for whole genome sequencing (WGS) with application in both healthy individuals and those with disease. In this manuscript, we will describe our [[bioinformatics]] strategy to efficiently process and deliver [[Genomics|genomic]] data to geneticists for clinical interpretation. We describe the handling of data from FASTQ to the final variant list for clinical review for the final report. We will also discuss our methodology for validating this workflow and the cost implications of running WGS. ('''[[Journal:Bioinformatics workflow for clinical whole genome sequencing at Partners HealthCare Personalized Medicine|Full article...]]''')<br /> | |||
<br /> | <br /> | ||
''Recently featured'': | ''Recently featured'': | ||
: ▪ [[Journal:Pathology report data extraction from relational database using R, with extraction from reports on melanoma of skin as an example|Pathology report data extraction from relational database using R, with extraction from reports on melanoma of skin as an example]] | |||
: ▪ [[Journal:A robust, format-agnostic scientific data transfer framework|A robust, format-agnostic scientific data transfer framework]] | : ▪ [[Journal:A robust, format-agnostic scientific data transfer framework|A robust, format-agnostic scientific data transfer framework]] | ||
: ▪ [[Journal:A benchmarking analysis of open-source business intelligence tools in healthcare environments|A benchmarking analysis of open-source business intelligence tools in healthcare environments]] | : ▪ [[Journal:A benchmarking analysis of open-source business intelligence tools in healthcare environments|A benchmarking analysis of open-source business intelligence tools in healthcare environments]] | ||
Revision as of 17:27, 30 January 2017
Effective implementation of precision medicine will be enhanced by a thorough understanding of each patient’s genetic composition to better treat his or her presenting symptoms or mitigate the onset of disease. This ideally includes the sequence information of a complete genome for each individual. At Partners HealthCare Personalized Medicine, we have developed a clinical process for whole genome sequencing (WGS) with application in both healthy individuals and those with disease. In this manuscript, we will describe our bioinformatics strategy to efficiently process and deliver genomic data to geneticists for clinical interpretation. We describe the handling of data from FASTQ to the final variant list for clinical review for the final report. We will also discuss our methodology for validating this workflow and the cost implications of running WGS. (Full article...)
Recently featured: