Journal:Handling metadata in a neurophysiology laboratory
| Full article title | Handling metadata in a neurophysiology laboratory |
|---|---|
| Journal | Frontiers in Neuroinformatics |
| Author(s) |
Zehl, Lyuba; Jaillet, Florent; Stoewer, Adrian; Grewe, Jan; Sobolev, Andrey; Wachtler, Thomas; Brochier, Thomas G.; Riehle, Alexa; Denker, Michael; Grün, Sonja |
| Author affiliation(s) |
Jülich Research Centre, Aix-Marseille Université, Ludwig-Maximilians-Universität München, Eberhard-Karls-Universität Tübingen, Aachen University |
| Primary contact | Email: l dot zehl at fz-juelich dot de |
| Editors | Luo, Qingming |
| Year published | 2016 |
| Volume and issue | 10 |
| Page(s) | 26 |
| DOI | 10.3389/fninf.2016.00026 |
| ISSN | 1662-5196 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://www.frontiersin.org/articles/10.3389/fninf.2016.00026/full |
| Download | https://www.frontiersin.org/articles/10.3389/fninf.2016.00026/pdf (PDF) |
|
|
This article should not be considered complete until this message box has been removed. This is a work in progress. |
Abstract
To date, non-reproducibility of neurophysiological research is a matter of intense discussion in the scientific community. A crucial component to enhance reproducibility is to comprehensively collect and store metadata, that is, all information about the experiment, the data, and the applied preprocessing steps on the data, such that they can be accessed and shared in a consistent and simple manner. However, the complexity of experiments, the highly specialized analysis workflows, and a lack of knowledge on how to make use of supporting software tools often overburden researchers to perform such a detailed documentation. For this reason, the collected metadata are often incomplete, incomprehensible for outsiders, or ambiguous. Based on our research experience in dealing with diverse datasets, we here provide conceptual and technical guidance to overcome the challenges associated with the collection, organization, and storage of metadata in a neurophysiology laboratory. Through the concrete example of managing the metadata of a complex experiment that yields multi-channel recordings from monkeys performing a behavioral motor task, we practically demonstrate the implementation of these approaches and solutions, with the intention that they may be generalized to other projects. Moreover, we detail five use cases that demonstrate the resulting benefits of constructing a well-organized metadata collection when processing or analyzing the recorded data, in particular when these are shared between laboratories in a modern scientific collaboration. Finally, we suggest an adaptable workflow to accumulate, structure and store metadata from different sources using, by way of example, the odML metadata framework.
Introduction
Technological advances in neuroscience during the last decades have led to methods that nowadays enable researchers to simultaneously record the activity from tens to hundreds of neurons simultaneously, in vitro or in vivo, using a variety of techniques[1][2][3] in combination with sophisticated stimulation methods, such as optogenetics.[4][5] In addition, recordings can be performed in parallel from multiple brain areas, together with behavioral measures such as eye or limb movements.[6][7] Such recordings enable researchers to study network interactions and cross-area coupling, and to relate neuronal processing to the behavioral performance of the subject.[8][9][10] These approaches lead to increasingly complex experimental designs that are difficult to parameterize, e.g., due to multidimensional characterization of natural stimuli[11] or high-dimensional movement parameters for almost freely behaving subjects.[12] It is a serious challenge for researchers to keep track of the overwhelming amount of metadata generated at each experimental step and to precisely extract all the information relevant for data analysis and interpretation of results. Various aspects such as the parametrization of the experimental task, filter settings and sampling rates of the setup, the quality of the recorded data, broken electrodes, preprocessing steps (e.g., spike sorting), or the condition of the subject need to be considered. Nevertheless, the organization of these metadata is of utmost importance for conducting research in a reproducible manner, i.e., the ability to faithfully reproduce the experimental procedures and subsequent analysis steps.[13][14][15] Moreover, detailed knowledge of the complete recording and analysis processes is crucial for the correct interpretation of results, and is a minimal requirement to enable researchers to verify published results and build their own research on the previous findings.
To achieve reproducibility, experimenters have typically developed their own lab procedures and practices for performing experiments and their documentation. Within the lab, crucial information about the experiment is often transmitted by personal communication, through handwritten laboratory notebooks, or implicitly by trained experimental procedures. However, at least when it comes to data sharing across labs, essential information is often missed in the exchange.[16][17] Moreover, if collaborating groups have different scientific backgrounds, for example experimenters and theoreticians, implicit domain-specific knowledge is often not communicated or is communicated in an ambiguous fashion that leads to misunderstandings. To avoid such scenarios, the general principle should be to keep as much information about an experiment as possible from the beginning on, even if information seems to be trivial or irrelevant at the time. Furthermore, one should annotate the data with these metadata in a clear and concise fashion.
In order to provide metadata in an organized, easily accessible, but also machine-readable way, Grewe et al.[18] introduced odML (open metadata Markup Language) as a simple file format in analogy to SBML in systems biology[19], or NeuroML in neuroscientific simulation studies.[20][21] However, lacking to date is a detailed investigation on how to incorporate metadata management in the daily lab routine in terms of (i) organizing the metadata in a comprehensive collection, (ii) practically gathering and entering the metadata, and (iii) profiting from the resulting comprehensive metadata collection in the process of analyzing the data. Here we address these points, both conceptually and practically, in the context of a complex behavioral experiment that involves neuronal recordings from a large number of electrodes that yield massively parallel spike and local field potential (LFP) data.[22] To illustrate how to organize a comprehensive metadata collection (i), we introduce in the next section the concept of metadata and demonstrate the rich diversity of metadata that arise in the context of the example experiment. To demonstrate why the effort of creating a comprehensive metadata collection is time well spent, we next describe five use cases that summarize where the access to metadata becomes relevant when working with the data (iii). Afterwards we provide detailed guidelines and assistance on how to create, structure, and hierarchically organize comprehensive metadata collections (i and ii). Complementing these guidelines, we provide in the supplementary material section a thorough practical introduction on how to embed a metadata management tool, such as the odML library, into the experimental and analysis workflow. Finally, we close by critically contrasting the importance of proper metadata handling against its difficulties and deriving future challenges.
Organizing metadata in neurophysiology
Metadata are generally defined as data describing data.[23][24] More specifically, metadata are information that describe the conditions under which a certain dataset has been recorded.[18] Ideally all metadata would be machine-readable and available at a single location that is linked to the corresponding recorded dataset. The fact that such central, comprehensive metadata collections are not common practice already today is by no means a sign of negligence on the part of the scientists, but is explained by the fact that in the absence of conceptual guidelines and software support, such a situation is extremely difficult to achieve given the high complexity of the task. Already, the fact that an electrophysiological setup is composed of several hardware and software components, often from different vendors, imposes the need to handle multiple files of different formats. Some files may even contain metadata that are not machine-readable and -interpretable. Furthermore, performing an experiment requires the full attention of the experimenters, which limits the amount of metadata that can be manually captured online. Metadata that arise unexpectedly during an experiment, e.g., the cause of a sudden noise artifact, are commonly documented as handwritten notes in the laboratory notebook. In fact, handwritten notes are often unavoidable, because the legal regulations of some countries, e.g., France, require the documentation of experiments in the form of a handwritten laboratory notebook.
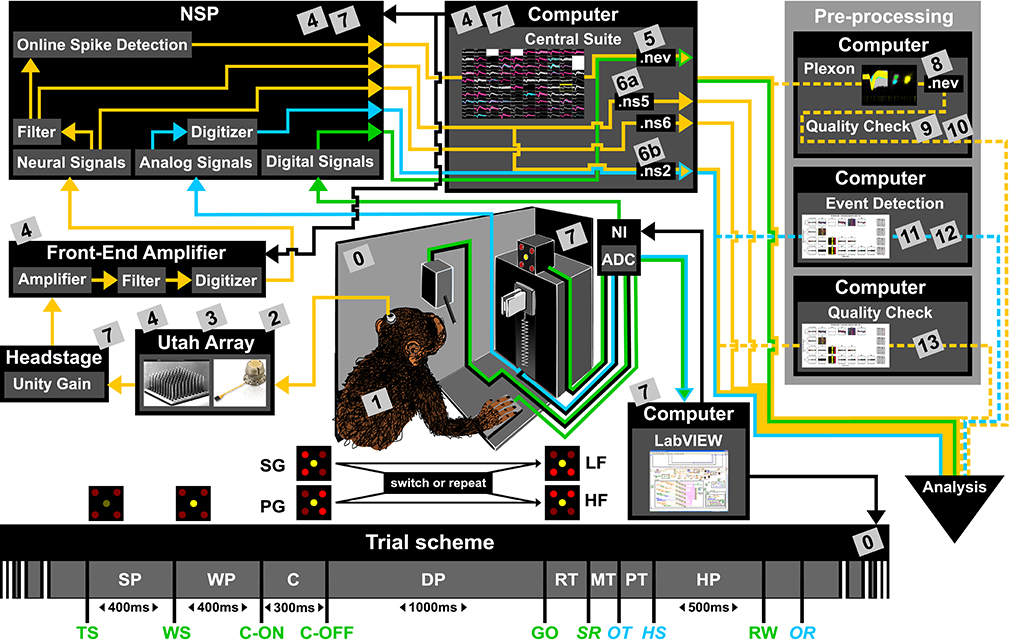
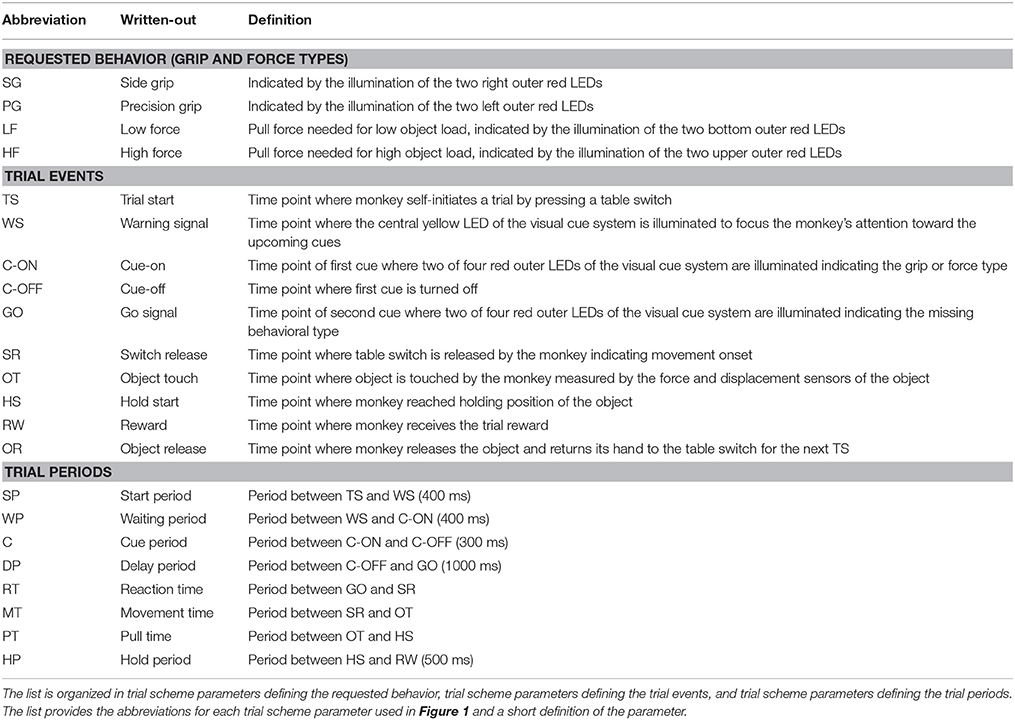
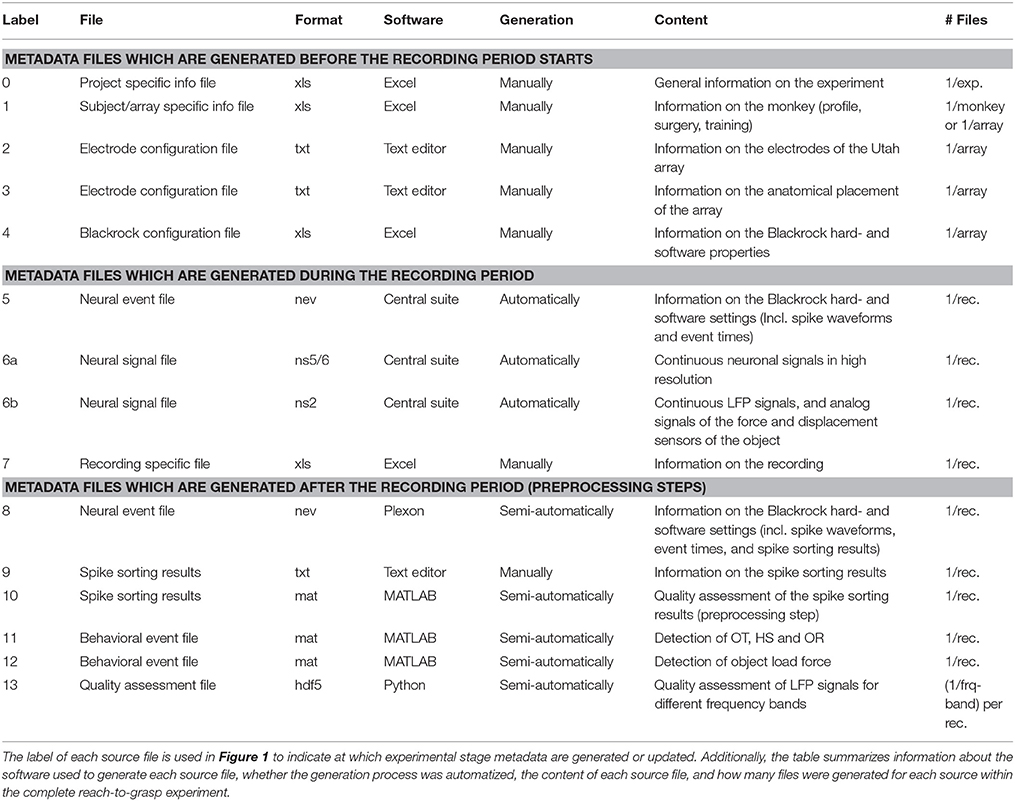
To present the concept of metadata management in a practical context, we introduce in the following a selected electrophysiological experiment. For clarity, a graphical summary of the behavioral task and the recording setup is provided in Figure 1, and a list of task-related abbreviations (e.g., behavioral conditions and events) is listed in Table 1. First described in Riehle et al.[22] and Milekovic et al.[25], the experiment employs high density multi-electrode recordings to study the modulation of spiking and LFP activities in the motor/pre-motor cortex of monkeys performing an instructed delayed reach-to-grasp task. In total, three monkeys (Macaca mulatta; two females, L, T; one male, N) were trained to grasp an object using one of two different grip types (side grip, SG, or precision grip, PG) and to pull it against one of two possible loads requiring either a high (HF) or low (LF) pulling force (cf. Figure 1). In each trial, instructions for the requested behavior were provided to the monkeys through two consecutive visual cues (C and GO) which were separated by a one-second delay and generated by the illumination of specific combinations of five LEDs positioned above the object (cf. Figure 1). The complexity of this study makes it particularly suitable to expose the difficulty of collecting, organizing, and storing metadata. Moreover, metadata were first organized according to the laboratory internal procedures and practices for documentation, while the organization of the metadata into a comprehensive machine-readable format was done a-posteriori, after the experiment was completed. To work with the data and metadata of this experiment, one has to handle on average 300 recording sessions per monkey, with data distributed over three files per session, metadata distributed over five files per implant, at least 10 files per recording, and one file for general information of the experiment (cf. Table 2). This situation imposed an additional complexity to reorganize the various metadata sources.
|
|
|
To give a more concrete impression of the painstaking detail that needs to be considered while planning and organizing a comprehensive metadata collection, we provide an exhaustive description of the experiment in the supplementary material. To illustrate the level of complexity of metadata management in this example, Figure 1 outlines the different components of the experimental setup, the signal flow, the task, and the trial scheme. In addition, the heterogeneous pieces of metadata and the corresponding files that contain them in the absence of a comprehensive metadata collection are listed and described in Table 2. Here, all metadata source files are labeled by numbers and appear in Figure 1 wherever they were generated. If labels appear multiple times, they were iteratively enriched with information obtained from the corresponding components of the setup.
In such a complex experiment, the relevance of some metadata does not become immediately apparent and is sometimes underestimated. For example, the immediate relevance of each minute detail of the experimental setup are of little interest for the interpretation of a single recording, and only when one attempts to rebuild the experiment do these metadata become valuable. Apart from that, any piece of metadata can become highly relevant for screening and selecting datasets according to specific criteria at a later stage. Different experiments produce different sets of metadata, and it is therefore not possible to provide a detailed list of metadata that needs to be collected from any given type of experiment. Nevertheless, the general principle should be to keep as much information about an experiment as possible from the beginning on, even if information seems to be trivial or irrelevant at the time. Based on our experience gathered from various collaborations, we compiled a list of guiding questions to help scientists define their optimal collection of metadata for their use scenario, e.g., reproducing the experiment and/or the analysis of the data, or sharing the data:
- What metadata would be required to replicate the experiment elsewhere?
- Is the experiment sufficiently explained, such that someone else could continue the work?
- If the recordings exhibit spurious data, is the signal flow completely reconstructable to find the cause?
- Are metadata provided which may explain variability (e.g., between subjects or recordings) in the recorded data?
- Are metadata provided which enable access to subsets of data to address specific scientific questions?
- Is the recorded data described in sufficient detail, such that an external collaborator could understand them?
For the example experiment presented in Figure 1, we used these guiding questions to control if the content of the various metadata sources was sufficient, and enriched these where necessary. The process of planning the relevant metadata content and organizing the metadata into a comprehensive collection can be time-consuming, especially if the process happened after the experiment was performed.
In the following, we will outline use cases that illustrate why the time for creating a comprehensive metadata collection is well spent, nevertheless.
Advantage of a comprehensive metadata collection: Use cases
Based on the example experiment (Figure 1), we present five use cases that demonstrate the common scenarios in which access to metadata becomes important. For the implementation of each use case, we contrast the scenarios before and after having organized metadata in a comprehensive collection:
- Scenario 1: Metadata are organized in different files and formats which are stored in a metadata source directory.
- Scenario 2: Metadata sources of Scenario 1 are compiled into a comprehensive collection, stored in one file per recording using a standard file format.
We introduce the protagonists acting in the use cases and characterize their relationship:
- Alice: She is the experimenter who built up the setup, trained the monkeys, and carried out the experimental study. She has programming experience in MATLAB and performs the preprocessing step of spike sorting.
- Bob: He is a data analyst and a new member of Alice's laboratory. He has programming experience in MATLAB and Python. His task is to support Alice in implementing the preprocessing and first analysis of the data.
- Carol: She is a theoretical neuroscientist and an analyst of an external group who collaborates with Alice's laboratory. She is an experienced Python programmer. Information exchange with Alice's group is limited to phone, email, and video calls, and few in-person meetings. She is not an experimentalist and therefore not used to work with real neuronal data. She never participated in an experiment and thus never experienced the workflow of a typical recording session.
Each of the use cases below illustrates a different aspect of the analysis workflow: enrichment of metadata information, metadata accessibility, selection and screening of datasets, and formal queries to reference metadata.
Use Case 1: Enrichment of the metadata collection
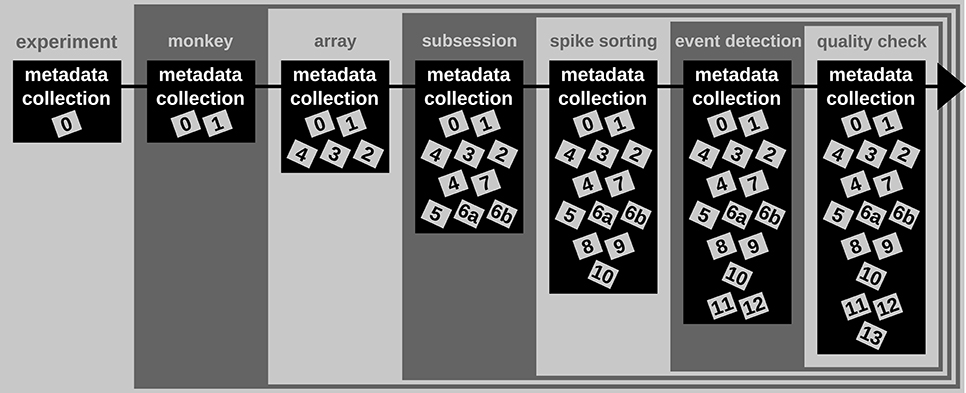
A difficult aspect of metadata management is that one cannot know all metadata in advance that are necessary to ensure the reproducibility of the experiment and the data analysis. Thus, the metadata collection needs to be enriched when new and updated information becomes available (see Figure 2). An example for such an enrichment of a metadata collection is the integration of results and parameters of preprocessing steps applied to the recorded data, since these are usually performed long after the primary data acquisition (cf. Table 2). Use Case 1 demonstrates the advantage of using a standard file format when a metadata collection is enriched with information from successive preprocessing steps.
|
In one of the first meetings with Alice, Bob learned that a number of preprocessing steps are performed by Alice. One of these is to identify and extract time points of the behavioral events describing the object displacement in each trial (OT, HS, OR, see Table 1) from the continuous force and displacement signals which were saved in the ns2-datafile (label 6b in Figure 1 and Table 2). For this, the signals were loaded by a program Alice developed in MATLAB to perform this preprocessing step. The extracted behavioral events are saved for each recording session along with the processing parameters in a respective .mat file (labels 11 in Figure 1 and Table 2). It is important that these preprocessing results are readily accessible to all collaboration partners, including Bob, because they are needed to correctly interpret the timeline and behavior in each trial.
In Scenario 1, Alice does not create a comprehensive metadata collection, and Bob has to deal with the fact that the behavioral events are not only not stored along with the digitally available trial events (e.g., TS, see Table 1) in the recorded nev-file (label 5 in Figure 1 and Table 2) but also saved in the custom-made mat-file, which requires him to write a corresponding loading routine. In Scenario 2, Alice saved the digital, directly available trial events into one comprehensive metadata collection per recording. With the preprocessing of each recording, she enriched then the corresponding primary collection with all results and parameters of the behavioral trial event extraction. As a result, the access to all trial events becomes easier for Bob, so that he is able to quickly scan the behavior in each trial.
In summary, a comprehensive metadata collection has the advantage that it bundles metadata that conceptually belongs together, and it simplifies access by combining metadata into one single, standard file format. Moreover, the flexible enrichment of a comprehensive metadata collection simplifies the organization of metadata that originate from multiple versions of performing a preprocessing step, e.g., as in the case of offline spike sorting (see Supplementary Material). With this, a comprehensive metadata collection not only simplifies the reproducibility of Alice's work (e.g., repetition of preprocessing steps), but also guarantees better reproducibility for the collaboration with Bob (e.g., standardized access to parameters and results of preprocessing steps).
Use Case 2: Metadata accessibility
Complex experimental studies usually include organizational structures on the level of the file system (e.g., directories) and file format to store all data and metadata. Even in cases where all data are very well organized, metadata are usually distributed over several files and formats (see Table 2). Use Case 2 demonstrates how a standard file format of a comprehensive metadata collection improves the accessibility and readability of metadata.
When Bob first arrived in the lab, Alice explained to him the structure of the data and metadata of the experimental study. Unfortunately, Bob could not start working on the data directly after the meeting with Alice. Of course he made several notes during this meeting, but due to the complexity of the information, he forgot where to find the individual pieces of metadata in sufficient detail.
In Scenario 1, Bob starts to scan all different metadata sources to regain an overview of what can be found where. This means that he has not only to go through several files, but also that he needs to deal with several formats, and that he needs to understand the different design of each file. In Scenario 2, Bob knows that all metadata are collected and stored in one file per recording, which is readable using standard software, e.g., a text editor. To access again an overview of the available metadata, he can open a file of any recording, screen its content or search for specific metadata.
Thus, the metadata organization in Scenario 1 has the following disadvantages: (i) it is difficult to keep track of which metadata source contains which piece of information; (ii) not every metadata source is readable with a simple, easily accessible software tool (e.g., the binary data files, .nev and .nsX; labels 5, 6a, and 6b in Figure 1 and Table 2, respectively); (iii) not every software tool offers the option to search for specific metadata content. In contrast, in Scenario 2 a comprehensive, standardized, and searchable organization of the metadata guarantees complete and easy access to all metadata related to an experiment, even over a long period of time. This simplifies the long-term comprehensibility of Alice's experiment not only for herself, but also for all further group members and collaboration partners.
Use Case 3: Selection of datasets
Data of an experiment can usually be used to address multiple scientific questions. In this context, it is necessary to select datasets according to certain criteria, which are defined by the requirements and constraints of the scientific question. These criteria are usually represented by metadata related to one recording session and generated during or after the recording period (see Table 2). Use Case 3 demonstrates how a comprehensive metadata collection with a standard file format supports the automatic selection of datasets according to defined selection criteria.
Alice reported that she had the feeling that the weekend break influences the monkey's performance on Mondays and asked Bob to investigate her suspicion. To solve this new task, Bob first wants to identify the recording sessions for each weekday (Criterion 1) which were recorded with standard task settings (Criterion 2) and under the standard experimental paradigm (instructed delayed reach-to-grasp task with two cues; Criterion 3). To automatize the data selection, Bob writes a Python program where he loops through all available recording sessions, uses Criterion 1 to identify the weekday, and adds each session name to a corresponding list if in the session Criteria 2 and 3 were fulfilled.
To write this program, Bob uses in Scenario 1 the knowledge he gained from the manual inspection in Use Case 2 to identify for the chosen selection criteria the different metadata source files. Criterion 1 was stored in the .nev data file (Label 5 in Figure 1 and Table 2) which Bob can access via the data loading routine. Criteria 2 and 3 are instead stored in the recording-specific spreadsheet (Label 7 in Figure 1 and Table 2). To automatize their extraction, Bob additionally needs to write a spreadsheet loading routine, because there is no standard loading routine available for the homemade structure of the spreadsheet. In contrast, in Scenario 2, Bob extracts all three criteria for each recording from one comprehensive metadata file using an available loading routine of the chosen standard file format.
In summary, compared to Scenario 1, Scenario 2 improved the workflow of selecting datasets according to certain criteria in two aspects: (i) to check for the selection criteria, only one metadata file per recording needs to be loaded, and (ii) to extract metadata stored in a standardized format, a loading routine is already available. Thus, in Scenario 2 the scripts for automatized data selection are less complicated, which improved the reproducibility of the operation.
Use Case 4: Metadata screening
Sometimes it can be helpful to gain an overview of the metadata of an entire experiment. Such a screening process is often negatively influenced by the following aspects: (i) Some metadata are stored along with the actual electrophysiological data, (ii) metadata are often distributed over several files and formats, and (iii) some metadata need to be computed from other metadata. All three aspects slow down the screening procedure and complicate the corresponding code. Use Case 4 demonstrates how a comprehensive metadata collection improves the speed and reproducibility of a metadata screening procedure.
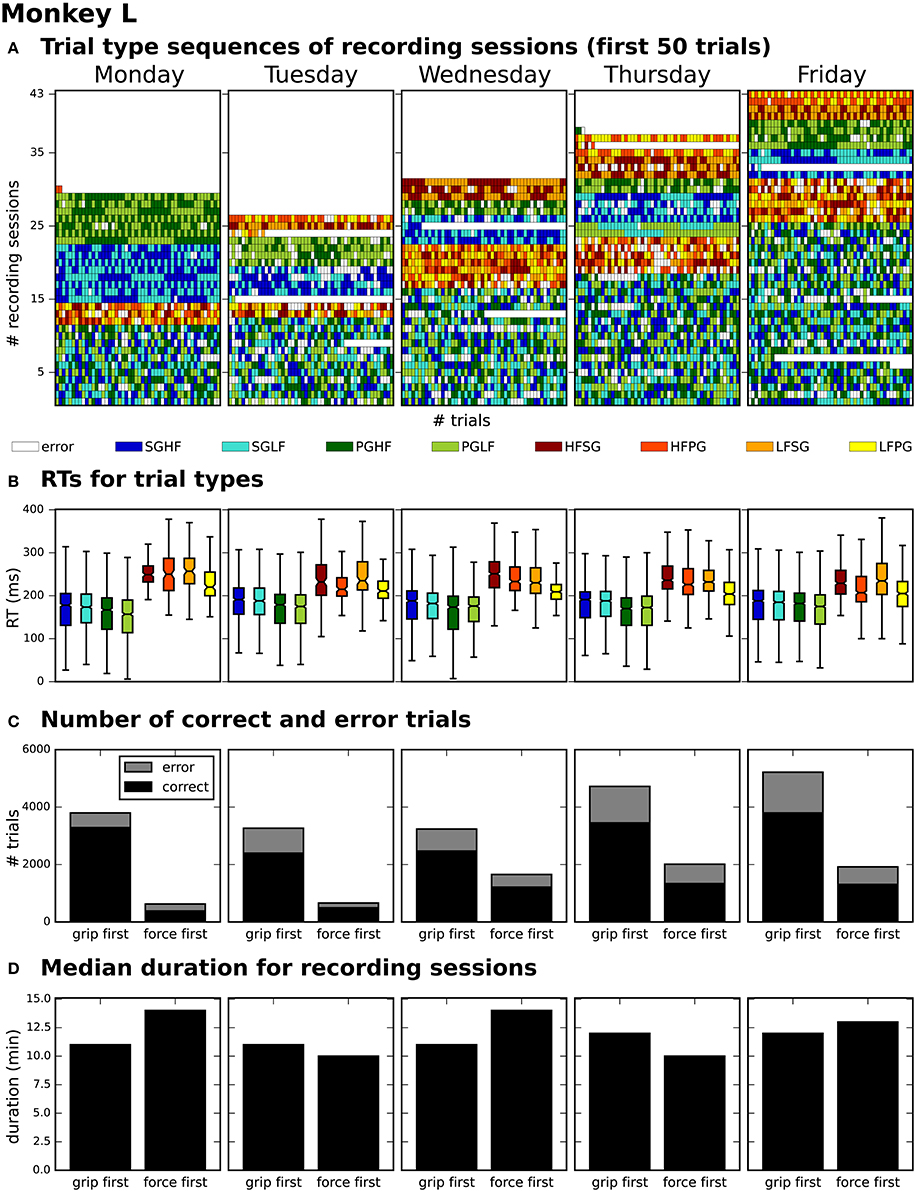
After generating for each weekday a corresponding list of recording file names (see Use Case 3, Section 3.3), Bob would like to generate an overview figure summarizing the set of metadata that best reflect the performance of the monkey during the selected recordings (Figure 3). This set includes the following metadata: the RTs of the trials for each recording (Figure 3B), the number of correct vs. error trials (Figure 3C), and the total duration of the recording (Figure 3D). To exclude a bias due to a variable distribution of the different task conditions, Bob also wants to include the trial type combinations and their sequential order (Figure 3A).
|
To create the overview figure in Scenario 1, Bob has to load the .nev data file (Label 5 in Figure 1 and Table 2) in which most selection criteria are stored, and additionally the .ns2 data file (Label 6b in Figure 1 and Table 2) to extract the duration of the recording. For both files, Bob is able to use available loading routines, but for accessing the desired metadata, he always has to load the complete data files. Depending on the data size this processing can be very time consuming. In Scenario 2, Bob is able to directly and efficiently extract all metadata from one comprehensive metadata file without having to load the neuronal data in parallel.
In summary, compared to Scenario 1, the workflow of creating an overview of certain metadata of an entire experiment is improved in Scenario 2 by reducing the number of metadata files which need to be screened to one per recording, and by drastically lowering the run time to collect the criteria used in the figure. In addition, Bob benefits from better reproducibility as in Use Case 3.
Use Case 5: Metadata queries for data selection
It is common that datasets are analyzed not only by members of the experimenter's lab but also collaborators. Two difficulties may arise in this context. First, the partners will often base their work on different workflow strategies and software technologies, making it difficult to share their code, in particular code that is used to access data objects. Second, the geographical separation represents a communication barrier resulting from infrequent and impersonal communication by telephone, chat, or email, requiring extra care in conveying relevant information to the partner in a precise way. Use Case 5 demonstrates how a standard format used to save the comprehensive metadata collection improves cross-lab collaborations by formalizing the communication process through the use of queries on the metadata.
Alice has detected a systematic noise artifact in the LFP signals of some channels in recording sessions performed in July (possibly due to additional air conditioning). As a consequence, Alice decided to exclude recordings performed in July from her LFP spectral analyses. Alice collaborates with Carol to perform complementary spike correlation analyses on an identical subset of recordings in order to find out if the network correlation structure is affected by task performance. To ensure that they analyze exactly the same datasets, the best data selection solution is to rely on metadata information that is located in the data files, rather than error-prone measures such as interpreting the file name or file creation date.
In Scenario 1, since Alice and Carol use different programming languages (MATLAB and Python), they need to ensure that their routines extract the same recording date. This procedure will require providing corresponding cross-validations between their MATLAB and Python routines to extract identical dates.
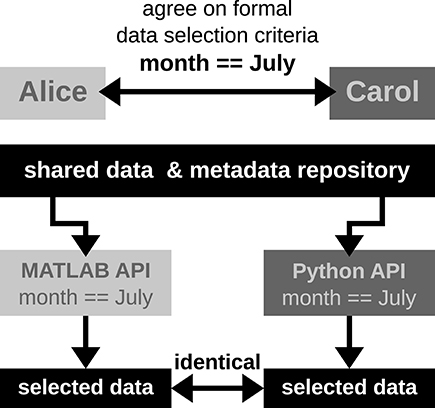
In Scenario 2, Alice and Carol agree on a concrete formal specification of the dataset selection (Figure 4) via the comprehensive metadata collection. In such a specification, metadata are stored in a defined format that reflects the structure of key-value pairs: in our example, Alice would specify the data selection by telling Carol to allow for only those recording sessions where the key month has the exact value "July." For the chosen standardized format of the collection, the metadata query can be handled by an application program interface (API) available both at the MATLAB and Python levels for Alice and Carol, respectively. Therefore, the query is guaranteed to produce the same result for both scientists. The formalization of such metadata queries will result in a more coherent and less error-prone synchronization between the work in the two laboratories.
|
References
- ↑ Nicolelis, M.A.; Ribeiro, S. (2002). "Multielectrode recordings: The next steps". Current Opinion in Neurobiology 12 (5): 602–6. doi:10.1016/S0959-4388(02)00374-4. PMID 12367642.
- ↑ Verkhratsky, A.; Krishtal, O.A.; Petersen, O.H. (2006). "From Galvani to patch clamp: The development of electrophysiology". Pflügers Archiv 453 (3): 233–47. doi:10.1007/s00424-006-0169-z. PMID 17072639.
- ↑ Obien, M.E.; Deligkaris, K.; Bullmann, T. et al. (2015). "Revealing neuronal function through microelectrode array recordings". Frontiers in Neuroscience 8: 423. doi:10.3389/fnins.2014.00423. PMC PMC4285113. PMID 25610364. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4285113.
- ↑ Deisseroth, K.; Schnitzer, M.J. (2013). "Engineering approaches to illuminating brain structure and dynamics". Neuron 80 (3): 568–77. doi:10.1016/j.neuron.2013.10.032. PMC PMC5731466. PMID 24183010. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5731466.
- ↑ Miyamoto, D.; Murayama, M. (2016). "The fiber-optic imaging and manipulation of neural activity during animal behavior". Neuroscience Research 103: 1–9. doi:10.1016/j.neures.2015.09.004. PMID 26427958.
- ↑ Maldonado, P.; Babul, C.; Singer, W. et al. (2008). "Synchronization of neuronal responses in primary visual cortex of monkeys viewing natural images". Journal of Neurophysiology 100 (3): 1523-32. doi:10.1152/jn.00076.2008. PMID 18562559.
- ↑ Vargas-Irwin, C.E.; Shakhnarovich, G.; Yadollahpour, P. et al. (2010). "Decoding complete reach and grasp actions from local primary motor cortex populations". Journal of Neuroscience 30 (29): 9659-69. doi:10.1523/JNEUROSCI.5443-09.2010. PMC PMC2921895. PMID 20660249. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2921895.
- ↑ Berényi, A.; Somogyvári, Z.; Nagy, A.J. et al. (2014). "Large-scale, high-density (up to 512 channels) recording of local circuits in behaving animals". Journal of Neurophysiology 111 (5): 1132-49. doi:10.1152/jn.00785.2013. PMC PMC3949233. PMID 24353300. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3949233.
- ↑ Lewis, C.M.; Bosman, C.A.; Fries, P. et al. (2015). "Recording of brain activity across spatial scales". Current Opinion in Neurobiology 32: 68–77. doi:10.1016/j.conb.2014.12.007. PMID 25544724.
- ↑ Lisman, J. (2015). "The challenge of understanding the brain: Where we stand in 2015". Neuron 86 (4): 864-882. doi:10.1016/j.neuron.2015.03.032. PMC PMC4441769. PMID 25996132. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4441769.
- ↑ Geisler, W.S. (2008). "Visual perception and the statistical properties of natural scenes". Annual Review of Psychology 59: 167–92. doi:10.1146/annurev.psych.58.110405.085632. PMID 17705683.
- ↑ Schwarz, D.A.; Lebedev, M.A.; Hanson, T.L. et al. (2014). "Chronic, wireless recordings of large-scale brain activity in freely moving rhesus monkeys". Nature Methods 11 (6): 670-6. doi:10.1038/nmeth.2936. PMC PMC4161037. PMID 24776634. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4161037.
- ↑ Laine, C.; Goodman, S.N.; Griswold, M.E.; Sox, H.C. (2007). "Reproducible research: Moving toward research the public can really trust". Annals of Internal Medicine 146 (6): 450–3. doi:10.7326/0003-4819-146-6-200703200-00154. PMID 17339612.
- ↑ Peng, R.D. (2011). "Reproducible research in computational science". Science 334 (6060): 1226-7. doi:10.1126/science.1213847. PMC PMC3383002. PMID 22144613. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3383002.
- ↑ Tomasello, M.; Call, J. (2011). "Methodological challenges in the study of primate cognition". Science 334 (6060): 1227-8. doi:10.1126/science.1213443. PMID 22144614.
- ↑ Hines, W.C.; Su, Y.; Kuhn, I. et al. (2014). "Sorting out the FACS: A devil in the details". Cell Reports 6 (5): 779-81. doi:10.1016/j.celrep.2014.02.021. PMID 24630040.
- ↑ Open Science Collaboration (2015). "Estimating the reproducibility of psychological science". Science 349 (6251): aac4716. doi:10.1126/science.aac4716. PMID 26315443.
- ↑ 18.0 18.1 Grewe, J.; Wachtler, T.; Benda, J. (2011). "A Bottom-up Approach to Data Annotation in Neurophysiology". Frontiers in Neuroinformatics 5: 16. doi:10.3389/fninf.2011.00016. PMC PMC3171061. PMID 21941477. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3171061.
- ↑ Hucka, M.; Finney, A.; Sauro, H.M. et al. (2003). "The systems biology markup language (SBML): A medium for representation and exchange of biochemical network models". Bioinformatics 19 (4): 524–31. doi:10.1093/bioinformatics/btg015. PMID 12611808.
- ↑ Gleeson, P.; Crook, S.; Cannon, R.C. et al. (2010). "NeuroML: a language for describing data driven models of neurons and networks with a high degree of biological detail". PLoS Computational Biology 6 (6): e1000815. doi:10.1371/journal.pcbi.1000815. PMC PMC2887454. PMID 20585541. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2887454.
- ↑ Crook, S.M.; Bednar, J.A.; Berger, S. et al. (2012). "Creating, documenting and sharing network models". Network 23 (4): 131-49. doi:10.3109/0954898X.2012.722743. PMID 22994683.
- ↑ 22.0 22.1 Riehle, A.; Wirtssohn, S.; Grün, S.; Brochier, T. (2013). "Mapping the spatio-temporal structure of motor cortical LFP and spiking activities during reach-to-grasp movements". Frontiers in Neural Circuits 7: 48. doi:10.3389/fncir.2013.00048. PMC PMC3608913. PMID 23543888. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3608913.
- ↑ Baca, M. publisher=Getty Publications, ed. (2016). Introduction to Metadata (3rd ed.). http://www.getty.edu/publications/intrometadata/.
- ↑ "Metadata". Merriam-Webster.com. Merriam-Webster. https://www.merriam-webster.com/dictionary/metadata. Retrieved 06 July 2016.
- ↑ Milekovic, T.; Truccolo, W.; Grün, S. et al. (2015). "Local field potentials in primate motor cortex encode grasp kinetic parameters". Neuroimage 114: 338–55. doi:10.1016/j.neuroimage.2015.04.008. PMC PMC4562281. PMID 25869861. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4562281.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added. The original article lists references alphabetically, but this version — by design — lists them in order of appearance. What were originally footnotes have been turned into inline external links.