Book:COVID-19 Testing, Reporting, and Information Management in the Laboratory/Diagnostic testing of COVID-19 and other coronaviruses/Testing terminology
2.1 Testing terminology
As you read through this chapter, you may discover terms like "polymerase chain reaction" (PCR) and "lateral flow assay" (LFA). If you're a laboratorian or have a clinical background, you're likely to be familiar with these terms. However, it seems prudent to at least briefly discuss a few of them before delving into coronavirus testing itself.
2.1.1. Introduction
Living organisms store information in their genetic material, using DNA or RNA as the information carrier. That information, or genetic code, essentially provides instructions for organism development, function, growth, and reproduction. In the late twentieth century, researchers were laying the groundwork for molecular diagnostics, the concept of examining an organism's genetic code and its associated biological markers to diagnose and treat disease on a more personalized basis. This requires an assay, an investigative procedure for assessing the presence of, or measuring the amount or functional activity of, a target analyte. In the case of molecular diagnostics, and more broadly molecular biology, the target is biological in nature, and thus biological assays are used. These biological assays are designed to accurately detect the presence of or enable counts of biological molecules, including DNA, RNA, proteins, cells, bacteria, and virus particles (e.g., viral plaque assays).[1]
2.1.2 Polymerase chain reaction (PCR)
Polymerase chain reaction or PCR is a molecular biology method that takes small amounts of DNA sequences and makes copies of (amplifies) them to the point of having enough material to sufficiently study or work with. The base technique can yield results in several hours and has a high level of sensitivity, with its ability to amplify the DNA to counts of millions or billions. PCR has been used in molecular diagnostics for testing prospective parents for being genetic carriers of particular diseases (i.e., expanded carrier screening)[2][3], tissue typing to ensure more effective organ transplants[4], and analyzing mutations in oncogenes to customize cancer treatments.[5] However, the method has also been applied to forensic science[6] and epidemiology.[7]
PCR and its variations have been used to characterize and detect infectious disease organisms such as human immunodeficiency virus (HIV), pathogenic tuberculosis bacteria, and Bordetella pertussis, which causes whooping cough.[8] Additionally, a selection of viruses can have their RNA detected using PCR, though the primers (short single-strand DNA fragments) used in the process must by sympathetic to the virus' genetic structure to ensure that only target virus material is amplified.[9] As it turns out, coronaviruses are RNA viruses, having some of the longest genomes of any RNA virus, and, detrimentally, the highest known frequency of recombination (the exchange of genetic material with another organism); this broadly means high rates of virus mutation, which interferes with maintaining consistent diagnostic detection and therapy.[10][11]
PCR comes in several variant methods. For example, while PCR monitors the amplification portion at the end of the overall process, real-time or "quantitative" PCR (qPCR) allows for the generation rate of the amplified product to be monitored at a particular point during each PCR cycle. Reverse transcription PCR (RT-PCR) is a combinatory process, applying reverse transcription (creating complementary double-stranded DNA [cDNA] from an RNA template) with PCR. If RT-PCR incorporates qPCR, you end up with "real-time RT-PCR" (rRT-PCR), sometimes referred to as "quantitative RT-PCR" (qRT-PCR). In the case of using PCR for detecting coronaviruses, more often than not we see some variation of RT-PCR, with or without real-time amplification monitoring. (It's important to not assume all RT-PCR processes incorporate real-time methods.)
How does PCR work in practice? The simplified version (see this JAMA Patient Page for a useful graphical explanation, using COVID-19 as an example) has a clinician obtaining a biological specimen from the appropriate location or source material. Then, special techniques are used to isolate viral (or in some cases, bacterial) genetic material from the specimen. (If RT-PCR is performed, the next step of reverse transcription of the isolated viral RNA into cDNA is also performed.) Once the viral genetic material is isolated, suitable primers that are sympathetic to the structure of the isolated genetic material are introduced. Those primers bind to the virus' genetic material and begin making copies of it. Fluorescent or other biomarkers that were attached to the copies during the PCR process eventually release from the copies, and an attempt is made to detect the presence of those biomarkers. The presence or absence of these markers drives the determination of a positive or negative detection for the sought-after virus.[12]
2.1.3 Lateral flow assay (LFA)
The lateral flow assay or LFA is another molecular biology method for detecting the presence of a target analyte in specimen material. In comparison to PCR, LFA has the advantages of being more rapid, low-cost, easy-to-use, and applicable at the point of care, though the average LFA is at best semi-quantitative in its results and has slightly lower sensitivity.[13][14] The method involves a cellulose-based strip with an ordered collection of specific "pads" and reagents that are reactive to a target analyte in a liquid, which is placed on the strip and moves across the various reagents using capillary and electrostatic interactions.[13][15][16] LFA has been used in molecular diagnostics for testing urine, saliva, sweat, serum, plasma, and other biological fluids for the presence of specific antigens and antibodies, as well as signs of gene amplification. The method has also been effectively applied to other industries outside healthcare, including the food and beverage, chemical, and environmental industries.[13]
In the realm of infectious disease, the LFA has played an important role in disease diagnosis and control, particularly in resource-constrained settings where resources are limited and point-of-care testing is critical to filling the gap.[17] Testing for the presence of an infectious agent in body fluids using LFA can be performed in two ways: lateral flow immunoassay (LFI) or lateral flow nucleic acid (LFNA). The immunoassay variant looks for antibodies created as a result of the presence of an infectious agent, whereas the nucleic acid variant is built to detect an amplified nucleic acid sequence specific to a target infectious agent.[18] Speaking specifically to the SARS-CoV-2 coronavirus, the antibody-based immunoassay method of lateral flow is predominantly used, targeting one of either a monoclonal antibody directed at a viral antigen, or a viral antigen that is recognizable by a patient's developed antibodies.[19][20]
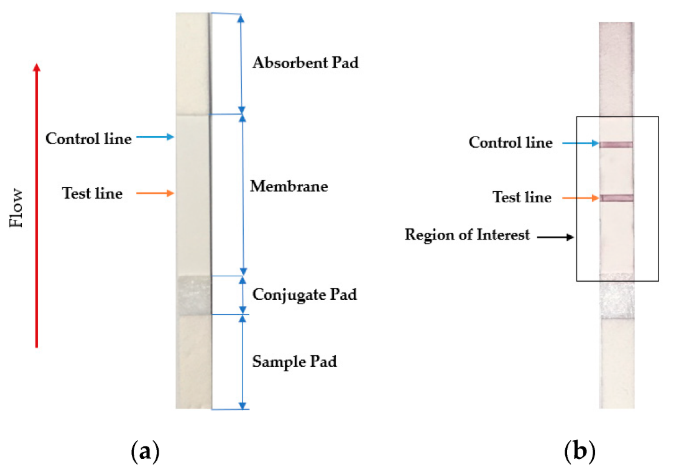
Testing using LFI is generally as follows. A specialized adsorbent sample pad at one end of the LFI strip receives the specimen material. That material then migrates to the next conjugate release pad, where the specimen material is exposed to "antibodies that are specific to the target analyte and are conjugated to coloured or fluorescent particles."[13] The material then progresses to a detection zone containing antibodies or antigens that are fixed in the zone and intended to react to a specific analyte. If the analyte is present, a test line produces a visual, qualitative response, and a control line ensures proper liquid flow across the strip. A wicking pad at the other end properly maintains the flow of liquids across the strip.[13]
References
- ↑ AdvaMedDx (2013). "Introduction to Molecular Diagnostics: The Essentials of Diagnostics Series". pp. 19. http://www.epemed.org/online/www/content2/108/469/3172/listdownloads/3175/507/ENG/dxinsights.pdf. Retrieved 06 September 2021.
- ↑ Gregg, A.R. (2018). "Expanded Carrier Screening". Obstetrics and Gynecology Clinics of North America 45 (1): 103–112. doi:10.1016/j.ogc.2017.10.005. PMID 29428278.
- ↑ Chokoshvili, D.; Vears, D.F.; Borry, P. (2018). "Reproductive autonomy in expanded carrier screening: More than meets the eye?". Expert Review of Molecular Diagnostics 18 (12): 993–94. doi:10.1080/14737159.2018.1544496. PMID 30394810.
- ↑ Edgerly, C.H.; Weimer, E.T. (2018). "The Past, Present, and Future of HLA Typing in Transplantation". Methods in Molecular Biology 1802: 1–10. doi:10.1007/978-1-4939-8546-3_1. PMID 29858798.
- ↑ Loda, M. (1994). "Polymerase chain reaction-based methods for the detection of mutations in oncogenes and tumor suppressor genes". Human Pathology 25 (6): 564–71. doi:10.1016/0046-8177(94)90220-8. PMID 7912220.
- ↑ Ninfa, A.J.; Ballou, D.P.; Benore, M. (2009). Fundamental Laboratory Approaches for Biochemistry and Biotechnology. Wiley. pp. 408–410. ISBN 9780470087664. https://books.google.com/books?id=k6_XQwAACAAJ&pg=PA408. Retrieved 08 April 2020.
- ↑ Hamborsky, J.; Kroger, A.; Wolfe, C., ed. (2015). Epidemiology and Prevention of Vaccine-Preventable Diseases (13th ed.). Centers for Disease Control and Prevention. https://www.cdc.gov/vaccines/pubs/pinkbook/index.html. Retrieved 08 April 2020.
- ↑ Buckingham, L. (2019). "Chapter 11: Detection and Identification of Microorganisms". Molecular Diagnostics: Fundamentals, Methods and Clinical Applications (3rd ed.). F.A. Davis Company. pp. 301–343. ISBN 9780803699540. https://books.google.com/books?hl=en&lr=&id=dJWNDwAAQBAJ&pg=301.
- ↑ Kim, H.; Kang, N.; An, K. et al. (2017). "MRPrimerV: A database of PCR primers for RNA virus detection". Nucleic Acids Research 45 (D1): D475–81. doi:10.1093/nar/gkw1095. PMC PMC5210568. PMID 27899620. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5210568.
- ↑ Makin, S. (5 February 2020). "How Coronaviruses Cause Infection—from Colds to Deadly Pneumonia". Scientific American. https://www.scientificamerican.com/article/how-coronaviruses-cause-infection-from-colds-to-deadly-pneumonia1/. Retrieved 08 April 2020.
- ↑ Rohde, R. (31 January 2020). "2019 Novel Coronavirus (2019-nCoV) Update: Uncoating the Virus". American Society for Microbiology. https://asm.org/Articles/2020/January/2019-Novel-Coronavirus-2019-nCoV-Update-Uncoating. Retrieved 08 April 2020.
- ↑ Hadaya, J.; Schumm, M.; Livingston, E.H. (2020). "Testing Individuals for Coronavirus Disease 2019 (COVID-19)". JAMA. doi:10.1001/jama.2020.5388. PMID 32236503.
- ↑ 13.0 13.1 13.2 13.3 13.4 Koczula, K.M.; Gallotta, A. (2016). "Lateral flow assays". Essays in Biochemistry 60 (1): 111–20. doi:10.1042/EBC20150012. PMC PMC4986465. PMID 27365041. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4986465.
- ↑ Liu, Z.; Hu, J.; Qu, Z.; Xu, F. (2018). "Chapter 8: Paper-Based Immunoassays". In Vashist, S.K.; Luong, J.H.T.. Handbook of Immunoassay Technologies: Approaches, Performances, and Applications. Academic Press. pp. 183–202. ISBN 9780128117620. https://books.google.com/books?id=jSk0DwAAQBAJ&pg=PA183. Retrieved 08 April 2020.
- ↑ Jauset-Rubio, M.; Svobodová, M.; Mairal, T. et al. (2016). "Ultrasensitive, rapid and inexpensive detection of DNA using paper based lateral flow assay". Scientific Reports 6: 37732. doi:10.1038/srep37732. PMC PMC5123575. PMID 27886248. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5123575.
- ↑ Foysal, K.H.; Seo, S.E.; Kim, M.J. et al. (2019). "Analyte Quantity Detection from Lateral Flow Assay Using a Smartphone". Sensors 19 (21): 4812. doi:10.3390/s19214812. PMC PMC6864604. PMID 31694281. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6864604.
- ↑ Hanafiah, K.M.; Arifin, N.; Bustami, T. et al. (2017). "Development of Multiplexed Infectious Disease Lateral Flow Assays: Challenges and Opportunities". Diagnostics 7 (3): 51. doi:10.3390/diagnostics7030051. PMC PMC5617951. PMID 28880218. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5617951.
- ↑ Ngom, B.; Guo, Y.; Wang, X.; Bi, D. (2010). "Development and application of lateral flow test strip technology for detection of infectious agents and chemical contaminants: A review". Analytical and Bioanalytical Chemistry 397: 1113–1135. doi:10.1007/s00216-010-3661-4. PMID 20422164.
- ↑ Sheridan, C. (23 March 2020). "Fast, portable tests come online to curb coronavirus pandemic". Nature Biotechnology - News. doi:10.1038/d41587-020-00010-2. https://www.nature.com/articles/d41587-020-00010-2. Retrieved 08 April 2020.









