Journal:SUSHI: An exquisite recipe for fully documented, reproducible and reusable NGS data analysis
| Full article title | SUSHI: An exquisite recipe for fully documented, reproducible and reusable NGS data analysis |
|---|---|
| Journal | BMC Bioinformatics |
| Author(s) | Hatakeyama, Masaomi; Opitz, Lennart; Russo, Giancarlo; Qi, Weihong; Schlapbach, Ralph; Rehrauer, Hubert |
| Author affiliation(s) | ETH Zürich and University of Zürich |
| Primary contact | Email: hubert dot rehrauer at fgcz dot ethz dot ch |
| Year published | 2016 |
| Volume and issue | 17 |
| Page(s) | 228 |
| DOI | 10.1186/s12859-016-1104-8 |
| ISSN | 1471-2105 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | http://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-016-1104-8 |
| Download | http://bmcbioinformatics.biomedcentral.com/track/pdf/10.1186/s12859-016-1104-8 (PDF) |
Abstract
Background: Next generation sequencing (NGS) produces massive datasets consisting of billions of reads and up to thousands of samples. Subsequent bioinformatic analysis is typically done with the help of open-source tools, where each application performs a single step towards the final result. This situation leaves the bioinformaticians with the tasks of combining the tools, managing the data files and meta-information, documenting the analysis, and ensuring reproducibility.
Results: We present SUSHI, an agile data analysis framework that relieves bioinformaticians from the administrative challenges of their data analysis. SUSHI lets users build reproducible data analysis workflows from individual applications and manages the input data, the parameters, meta-information with user-driven semantics, and the job scripts. As distinguishing features, SUSHI provides an expert command line interface as well as a convenient web interface to run bioinformatics tools. SUSHI datasets are self-contained and self-documented on the file system. This makes them fully reproducible and ready to be shared. With the associated meta-information being formatted as plain text tables, the datasets can be readily further analyzed and interpreted outside SUSHI.
Conclusion: SUSHI provides an exquisite recipe for analysing NGS data. By following the SUSHI recipe, SUSHI makes data analysis straightforward and takes care of documentation and administration tasks. Thus, the user can fully dedicate his time to the analysis itself. SUSHI is suitable for use by bioinformaticians as well as life science researchers. It is targeted for, but by no means constrained to, NGS data analysis. Our SUSHI instance is in productive use and has served as data analysis interface for more than 1000 data analysis projects. SUSHI source code as well as a demo server are freely available.
Keywords: Data analysis framework, reproducible research, meta-level system design
Background
Today’s bioinformatics faces the practical challenge of analysing massive and diverse data in a well documented and reproducible fashion. The situation is particularly challenging in the area of NGS research where state-of-the-art algorithms are frequently available as standalone tools and where a complete data analysis consists of many individual data processing and analysis steps. The considerations associated with conducting such a data analysis in a research environment have been discussed by W. S. Noble[1], and guidelines as well as an example strategy for organizing computational data analysis have been given. According to Noble a key principle is to record every operation such that reproducibility is ensured.
In this paper, we present SUSHI, which stands for "Support Users for SHell-script Integration," a new approach to bioinformatics analysis that is centered on reusability, reproducibility and scalability. SUSHI produces analysis results as directories that are fully self-contained and hold all the information to be reproduced. Specifically, we document all parameters, input data, commands executed, as well as the versions of the tools and the reference data used. Additionally, we store meta-information on the experimental data together with the result files, so that those can be interpreted and further analyzed by other tools independently from SUSHI. This holds even if the analysis directory is transferred to collaborators with a different computing environment. SUSHI is extendable and we have put special emphasis on the simplicity of adding new software applications. A bioinformatician can define them within a single file and does not need special programming skills. SUSHI natively offers a command line interface as well as a web interface to run data analysis steps. Altogether, SUSHI lets bioinformaticians efficiently build analysis pipelines and ensures that analysis results are ready-to-be-shared and reproducible.
Various types of data analysis frameworks have already been implemented. They can be essentially divided into web-based frameworks and scripting frameworks. Examples for web-based frameworks are Galaxy[2], Chipster[3], GeneProf[4] or GenePattern.[5] They let users run individual steps or entire pipelines on a remote compute system with the framework keeping track of the executed analysis. Scripting frameworks like bpipe[6], Ruffus[7], nestly[8], NGSANE[9], Makeflow[10], and Snakemake[11] let users build bioinformatics pipelines in a command line fashion. Given the different types of user interactions, the former solutions are more targeted for the experienced biologists or the application-oriented bioinformaticians, while the latter address the needs of bioinformaticians who are more inclined to programming and high-throughput analysis of many datasets. However, there is no system as yet that natively offers both interfaces. Additionally, none of the existing frameworks puts an emphasis on having a human-readable and portable file-based representation of the meta-information and associated data.
Implementation
SUSHI data sets and applications
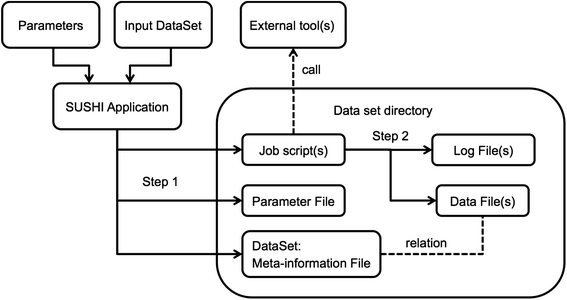
Within SUSHI, the original measured data as well as any derived analysis result is modeled as a data set that is represented as a fully self-sufficient directory on the file system. For the original data this means that it must be accompanied by meta-information on how the data has been measured. The meta-information must include information on the biological samples used as well as information on the measurement process. For analysis results this implies that the analysis result files must have accompanying meta-information that documents the input data, the analysis steps, and the analysis parameters. If these requirements are satisfied, we call this a "dataset." Figure 1 shows schematically how a dataset is generated as the result of running a SUSHI application. The meta-information associated with a dataset is represented in SUSHI as a DataSet object. On the file system a tabular file called "dataset.tsv" represents the dataset. Examples of meta-information are characteristics like sample name, species, tissue, but also e.g. the genome build that has been used for read alignment. Each characteristic is represented as a separate column in the tabular dataset.tsv.
|
A SUSHI application requires as input both a set of parameters and a DataSet object. This means that applications do not take bare data files as direct input. Instead, SUSHI applications take as input the DataSet meta-information object. The DataSet object holds, next to the data files, the meta-information necessary to process and interpret the data files. Based on its input, a SUSHI application first generates 1) the necessary job script(s), 2) a file representation of the parameters, and 3) the DataSet for the output data (Fig. 1 Step 1). The actual result data file(s) are generated by the job script(s) (Fig. 1 Step 2). The columns of the output DataSet hold again the meta-information, which now include additionally the parameters of the executed analysis if relevant for the further analysis or interpretation. The set of characteristics that is added to the annotation columns is defined and generated by the SUSHI application. The SUSHI framework itself does not require any specific annotation columns. Thus, the semantics of the DataSet columns are determined by the SUSHI applications (described in detail in the next section).
Every column of meta-information has a unique header that identifies the content and optional tags that characterize the information type in the column. Tags are represented as comma-separated strings within square brackets in the column headers. Currently supported tags are "File," "Link," "Factor," and "Characteristics." Depending on the tags, the SUSHI framework provides appropriate actions for the corresponding columns. The File tag is reserved for actual file paths, and SUSHI checks if the file actually exists. If a column has a Link tag, SUSHI will automatically add a hyperlink to the data. Finally, the Factor column data will be used to group samples according to experimental factors, which is typically required in a differential gene expression analysis.
Example DataSet holding RNA-seq reads
We follow the convention that sequencing read files are represented as a DataSet with the following columns:
- Name: the name of the sample measured
- Read1 [File]: path to the file holding the reads; if reads are paired-end, this must be the first read
- Adapter1: potentially contaminating adapter sequence at the 3′-end of read 1
- Read2 [File]: path to the second read for paired-end data (only for paired-end data)
- Adapter2: potentially contaminating adapter sequence at the 3′-end of read 2 (only for paired-end data)
- Species: the species of the sample
- StrandMode: specifies whether the library preparation protocol preserved strand information
- Enrichment Kit: the kit employed to enrich the input material (e.g. poly-A selection kit)
- Read Count: the number of reads in the file
Additionally, there can be columns that specify experimental factors. Table 1 shows an example (due to space constraints only a subset of the columns is shown).
| ||||||||||||||||||||||||
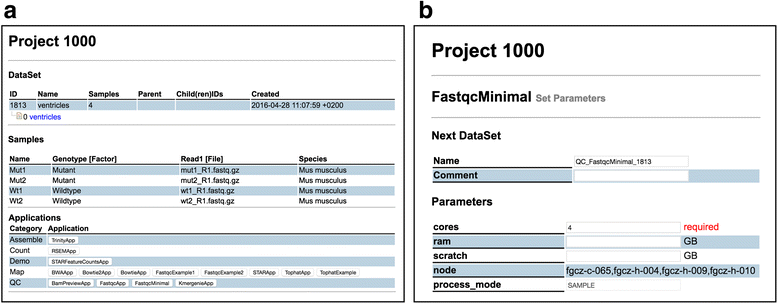
It is important to mention here that the SUSHI framework does not impose any constraints or semantics on the columns of the DataSet table. It is up to the user to identify which meta-information is relevant for his data. In particular, we do not require specific ontologies or controlled vocabularies. Users are free to define their own meta-information. The content definition and interpretation is entirely delegated to the user and the SUSHI applications. Figure 2a shows how the above DataSet is visualized in the SUSHI DataSet view.
|
Example SUSHI application performing a FastQC report
A common task is to generate a FastQC report[12] for each sample in a read data set. If the FastQC package is installed, one would run e.g. fastqc -o . -t 4 p1001/ventricles/mut1_R1.fastq.gz which creates the FastQC report mut1_R1_fastqc.zip for the first sample in the above data set. With the SUSHI framework this can be turned into a SUSHI application defined by the following Ruby code:
The example code shows the essential features of a SUSHI application (See also Additional file 1). The "@required_columns" tells the SUSHI framework which columns a DataSet must have so that "FastqcMinimal" is applicable. In Fig. 2a, all applications that are compatible with the example reads data set are shown at the bottom, including the "FastqcMinimal" application. The "@params['cores']" defines the number of cores to be used for multi-threading as a parameter with default value 4. This parameter is automatically turned into an input field in the web interface (see Fig. 2b. The code also defines with the method "next_dataset" the columns and content for the resulting DataSet. Finally, the method "commands" defines the command to be executed.
The SUSHI framework automatically performs administrative tasks such as putting the resulting file in the correct directory and managing the log files The full result directory is available as Additional file 2. A second example is the example for TopHat mapping[13] in Additional files 3 and 4. Both examples are for the illustrative purpose kept minimal. In real world application one would define additional parameters, support paired-end reads, and so on.
A list of all SUSHI applications that is in use at the Functional Genomics Center Zurich is available as Additional file 5 or on the SUSHI demo server: http://fgcz-sushi-demo.uzh.ch/sushi_application.
SUSHI applications are meta-process objects
Conceptually, a SUSHI application is a meta-process that generates an actual application (shell script). The entire data analysis functionality is contained within the shell script. For example, for a read alignment application, the job script contains the call to the aligner. If a SUSHI application is suitable only for a certain data type, it defines requirements on the input data. The requirements are specified as mandatory DataSet columns that an input dataset must have. The SUSHI framework guarantees that applications are only available for compatible DataSets. At run time the SUSHI application defines the columns of the resulting DataSet it will produce. With this meta-process modeling approach, the applications define the semantics of the meta-information without referring to other applications but only to DataSet content. Finally, from the input data and the parameters provided, the core method of a SUSHI application builds the command lines as a shell script that will eventually produce the output. The shell script can call other scripts or tools on the execution server. The only constraints for tools are:
- must be runnable from command line;
- must not require interactive input at run time; and
- result must be representable as a file or a set of files in a directory.
The framework cares about job execution, data file placement and cleanup of temporary data. Technically, a SUSHI application is implemented in a single Ruby file as a single Ruby class that takes over the SUSHI application super class using the template method design pattern (see Additional file 6).
SUSHI architecture follows meta-level design
SUSHI consists of three modules that are implemented in Ruby: 1) The SUSHI application module, as discussed above; 2) a Workflow Manager that performs job execution either on a local host or on a grid or cloud environment; and 3) the SUSHI server that relies on Ruby on Rails and provides the web front-end. The SUSHI server delegates the core function of generating a job script to the SUSHI application, and the SUSHI application communicates with the Workflow Manager for job submission. SUSHI relies heavily on Ruby meta-programming in that the executable code is dynamically generated at run time. During the code execution, all graphical components are dynamically generated based on the SUSHI application. This process follows the basic principles of Ruby on Rails: DRY (Don’t Repeat Yourself) and CvC (Convention over Configuration). Namely, the SUSHI application serves as the data source for the generation of the graphical components of the GUI. As a consequence, every data analysis that is implemented as a SUSHI application is directly available on the web front-end of the SUSHI server. The implementation of a SUSHI application does not need any web development knowledge, so that the bioinformatician can focus on data analysis aspects.
Installation
The SUSHI server is implemented in Ruby on Rails and can be installed in one step including all dependencies. The default installation uses the WEBrick web server application and SQLite3 database management system. Alternatively, SUSHI can be configured to run using an Apache web-server and a MySQL backend. For more details, please refer to README.rdoc in the source repository, https://github.com/uzh/sushi.
Results and discussion
Use case RNA-seq data analysis
A commonly used minimal workflow in RNA-seq data analysis consists of the following steps:
1. Map Reads with the STAR aligner.[14]
2. Compute expression values with the featureCounts function (Rsubread package).[15]
3. Detect differential expressed genes with edgeR.[16]
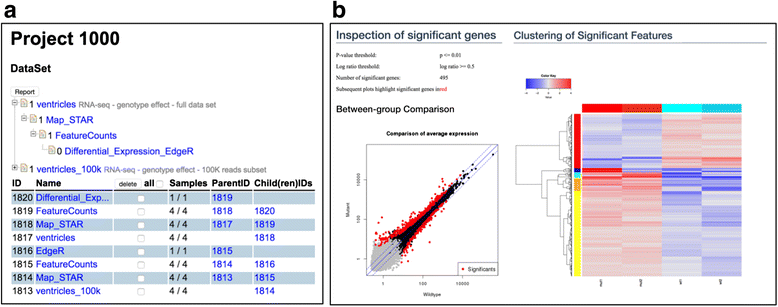
Figure 3a shows a representation of the generated datasets in SUSHI. The tree at the top indicates the hierarchical relationship and the bottom list shows the timeline of the generated data sets. Every data set is derived from its parent by running an application. Figure 3b shows visualizations that are generated by the last step, the assessment of differential expression between the wild-type and mutant samples in the data set. The scatter plot and the heat map indicate the significant genes and visualize the expression profiles. As with every SUSHI dataset, this result including all visualizations is downloadable and ready for offline use.
|
Comparison with existing systems
We compare SUSHI to similar bioinformatics frameworks (Table 2 and Additional file 7). A main distinction of the various frameworks is whether they have a graphical user interface (GUI) or a command line interface (CLI). The majority of CLI systems is implemented as an extension of a programming language or implemented in a domain specific language (DSL), and targeted for bioinformaticians with programming skills. The GUI systems are implemented as web applications with a variety of components and functions. Their installation and configuration is more complex relative to CLI systems but their usage does not require programming skills, so that they are targeted towards biomedical researchers.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Chipster[17] is designed as a GUI application and does not have a CLI or batch process mode, meaning a user must run applications manually one by one. GenePattern[5] and Taverna[18] provide both GUI and CLI. These systems are designed for GUI usage but provide also command-line access through special clients with an application programming interface (API). On the other hand, the SUSHI application model is designed for command line usage, and the web front-end is auto-generated using the Ruby on Rails meta-process.
In terms of reproducibility and documentation, CLI systems tend to have analysis information in a single text file, while GUI systems come with a relational database management system (RDBMS) that provides this and additional functionality. Generally, the framework itself is required to run an application or workflow again to reproduce results. Different from that, SUSHI provides the inherent advantages of GUI systems but still generates a set of human-readable shell script files that contain all the processing information and can be run independent of SUSHI (see the scripts in Additional files 1 and 3). In essence, SUSHI conveniently ensures full documentation and high reproducibility but is not needed to reproduce the analysis results.
Representation of meta-information
Accurate and high quality meta-information is necessary for the interoperability and integration of different data sources. However, even 10 years after "minimum information about a microarray experiment" (MIAME) guidelines[19] has been established, there is no consensus on how this should be implemented. This can be seen from the fact that different repositories and consortia use different implementations. Examples include GEO SOFT[20], Sequence Read Archive (SRA) XML[21], ENCODE[22], modENCODE[23], and ISA-TAB.[24] Given this situation, we decided that SUSHI should not implement its own constraints on meta-information. It is up to the data providers to decide, for example, which meta-information fields are provided and whether they are filled with free text, controlled vocabularies or terms from an ontology. SUSHI simply makes sure that the meta-information is preserved and that the analysis results are linked to the source data and accompanying meta-information. It goes without saying that we always encourage the use of controlled vocabularies and ontologies wherever possible.
Productive use and user acceptance
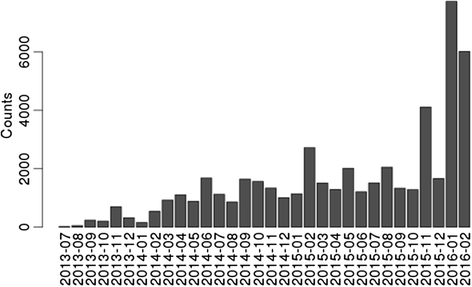
SUSHI is in productive use at the Functional Genomics Center Zurich and has been used to analyze NGS data in more than 1000 projects. A publicly available instance is placed at http://fgcz-sushi-demo.uzh.ch. The productive instance is integrated with the project and data management system B-Fabric[25][26] that handles projects, users, generated data, and access control. Currently we have implemented applications that support NGS analysis workflows for RNA-seq, small RNA-seq, ChIP-seq, de novo assembly, genotyping and variant analysis. The number of submitted jobs using SUSHI has been constantly increasing. Figure 4 shows the monthly submitted jobs at the Functional Genomics Center Zurich since 2013 which has reached now up to 5000 submitted jobs per month.
|
We have introduced SUSHI step-wise at the Functional Genomics Center Zurich. Initially, we supported only the most frequently used analyses such as RNA-seq and SNP calling, and subsequently we supported other applications like small RNA-seq, ChIP-seq and de novo assembly. SUSHI was readily adopted by the bioinformaticians because they had an immediate direct net benefit without a compromise on the flexibility in terms parameter choices. SUSHI development was driven and shaped by direct user feedback, which also broadened the acceptance.
Advanced separation of SUSHI concerns
The separation of concerns (SoC) is a software design principle in computer science.[27] It is now widely accepted and adopted in a variety of computer systems and software design such as object oriented programming and modularity of software design. The Model-View-Controller (MVC) design pattern is a typical example of separation of concerns for better software maintainability. The meta-level design is one type of advanced separation of concerns (ASoC) beyond object-oriented which can be seen in recent software paradigms such as generic programming, generative programming, and meta-programming. For example, the reflection architecture in Pattern-Oriented Software Architecture (POSA)[28] separates a system into meta-level and base-level and by controlling a meta-level it triggers a change at the base-level that actually provides a service to a user.
The current situation of the common NGS data analysis, such that several independent software applications are combined and chained to produce a final result, presents the two following main aspects: 1) which applications are used, and 2) how they are actually used. SUSHI separates these aspects (concerns) into a meta-process with meta-information (SUSHI application and DataSet object) and a base-process (shell scripts that do the data analysis). This separation of concerns results in the loose coupling of SUSHI into its system level and user application level. This improves the independence of SUSHI applications and shell scripts.
Conclusions
SUSHI targets both biologists and bioinformaticians as users. Analyzing data with SUSHI does not require programming skills, while adding new SUSHI applications requires only basic experience in writing scripts using the syntax of shell, R, Python, Ruby, or similar. SUSHI is particularly attractive to data analysis experts and bioinformaticians. SUSHI relieves users from administrative burdens and aids documentation and data organization. The full flexibility of the underlying tools stays untouched and can be directly accessed. Additionally, the fact that datasets are fully defined on the file system lets users prototype new workflows without the need to integrate those in the SUSHI instance. Finally, experts can automate data analysis tasks with the command line interface and are not limited to the web interface.
The design of the SUSHI system is driven by the idea of having analysis results fully defined and self-contained on the file system. In fact, if SUSHI is shut down, all the results can still be used and interpreted. All the meta-information is available in human readable tabular formats and all job scripts are contained with no back reference to the SUSHI framework. SUSHI provides no laboratory information management system (LIMS) or computing functionality. Instead, through its open architecture, it readily integrates with existing LIMS systems and computing resources.
In one solution, SUSHI provides at the same time fully documented, high level NGS analysis tools to biologists and an easy to administer, reproducible approach for large and complicated NGS data to bioinformaticians. This is mostly obtained by using the meta-level system design. Bioinformaticians will be freed from the boring tasks of managing software application and documentation and they can focus more on method development and on data analysis itself. The separation of the Workflow Manager from the SUSHI server makes the adaptation to any kind of computing facility easy and leaves the possibility to scale up.
The meta-level system design gives the simple but powerful framework of no data representation: SUSHI fully delegates the definition of dataset semantics to the user. The SUSHI system itself only defines how meta-information is used in a SUSHI application. It yields portability of datasets and lowers the barrier to reuse data and augment human readability of data set meta-information. The meta-level system design produces the decoupling of the meta-process from the base application process, which increases the degrees of freedom on the user side and contributes to the flexibility and scalability of the system.
Availability and requirements
Project name: SUSHI
Project homepage: https://github.com/uzh/sushi
Demo installation: http://fgcz-sushi-demo.uzh.ch.
Operating system(s): Platform-independent, but we recommend Unix-like system such as Ubuntu Linux and MacOS X
Programming language: Ruby (> = 1.9.3)
Other requirements: Ruby on Rails (> = 3.2.9, < 4.0)
License: MIT
For the RubyGems library dependency, please refer to the Additional file 8, Gemfile.lock, which is also included in the Git repository.
Abbreviations
API: application programming interface
ASoC: advanced separation of concerns
ChIP-seq: chromatin immunoprecipitation and next-generation DNA sequencing
CLI: command line interface
CvC: convention over configuration
DRY: don’t repeat yourself
DSL: domain-specific language
GUI: graphical user interface
LIMS: laboratory information management system
MIAME: minimum information about a microarray experiment
MVC: model-view-controller
NGS: next generation sequencing
RDBMS: relational database management system
RNA-seq: RNA sequencing
SoC: separation of concerns
SRA: sequence read archive
Declarations
Acknowledgements
We would like to kindly thank the members of the IT and Genomics/Transcriptomics groups at the Functional Genomic Center Zurich for useful discussions and suggestions. Additionally we would like to thank all users for their valuable feedback.
Funding
No funding was obtained for this study.
Availability of data and material
Source code: https://github.com/uzh/sushi
Public demo installation: http://fgcz-sushi-demo.uzh.ch
Competing interests
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable
Additional files
- Additional file 1: An example of minimal SUSHI application to generate FastQC job script. (RB 658 bytes)
- Additional file 2: The result dataset produced by the FastQC SUSHI application written as Additional file 1. (TGZ 706 kb)
- Additional file 3: An example of SUSHI application to generate TopHat job script. (RB 2 kb)
- Additional file 4: The result dataset produced by the TopHat SUSHI application written as Additional file 3. (TGZ 2223 kb)
- Additional file 5: List of SUSHI application. (XLS 5 kb)
- Additional file 6: Class diagram of SUSHI application. (PNG 61 kb)
- Additional file 7: Comparison of workflow management systems. (XLS 35 kb)
- Additional file 8: List of required RubyGems library. It is bundled in the SUSHI source code as Gemfile.lock. (TXT 4 kb)
References
- ↑ Noble, W.S. (2009). "A quick guide to organizing computational biology projects". PLoS Computational Biology 5 (7): e1000424. doi:10.1371/journal.pcbi.1000424. PMC PMC2709440. PMID 19649301. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2709440.
- ↑ Goecks, Jeremey; Nekrutenko, Anton; Taylor, James; The Galaxy Team (2010). "Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences". Genome Biology 11 (8): R86. doi:10.1186/gb-2010-11-8-r86. PMC PMC2945788. PMID 20738864. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2945788.
- ↑ Fisch, K.M.; Meißner, T.; Gioia, L. et al. (2015). "Omics Pipe: A community-based framework for reproducible multi-omics data analysis". Bioinformatics 31 (11): 1724-1728. doi:10.1093/bioinformatics/btv061. PMC PMC4443682. PMID 25637560. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4443682.
- ↑ Halbritter, F.; Vaidya, H.J.; Tomlinson, S.R. (2011). "GeneProf: Analysis of high-throughput sequencing experiments". Nature Methods 9 (1): 7–8. doi:10.1038/nmeth.1809. PMID 22205509.
- ↑ 5.0 5.1 Reich, M.; Liefeld, T.; Gould, J. et al. (2006). "GenePattern 2.0". Nature Genetics 38 (5): 500–501. doi:10.1038/ng0506-500. PMID 16642009.
- ↑ Sadedin, S.P.; Pope, B.; Oshlack, A. (2012). "Bpipe: a tool for running and managing bioinformatics pipelines". Bioinformatics 28 (11): 1525-6. doi:10.1093/bioinformatics/bts167. PMID 22500002.
- ↑ Goodstadt, L. (2010). "Ruffus: A lightweight Python library for computational pipelines". Bioinformatics 26 (21): 2778-9. doi:10.1093/bioinformatics/btq524. PMID 20847218.
- ↑ McCoy, C.O.; Gallagher, A.; Hoffman, N.G.; Matsen, F.A. (2013). "nestly: A framework for running software with nested parameter choices and aggregating results". Bioinformatics 29 (3): 387–8. doi:10.1093/bioinformatics/bts696. PMC PMC3562064. PMID 23220574. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3562064.
- ↑ Buske, F.A.; French, H.J.; Smith, M.A. et al. (2014). "NGSANE: A lightweight production informatics framework for high-throuput data analysis". Bioinformatics 30 (10): 1471-1472. doi:10.1093/bioinformatics/btu036. PMC PMC4016703. PMID 24470576. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4016703.
- ↑ Yu, L.; Moretti, C.; Thrasher, A. et al. (2010). "Harnessing parallelism in multicore clusters with the All-Pairs, Wavefront, and Makeflow abstractions". Cluster Computing 13 (3): 243-256. doi:10.1007/s10586-010-0134-7.
- ↑ Köster, J.; Rahmann, S. (2012). "Snakemake: A scalable bioinformatics workflow engine". Bioinformatics 28 (19): 2520-2. doi:10.1093/bioinformatics/bts480. PMID 22908215.
- ↑ Andrews, S. (2010). "FastQC: A quality control tool for high throughput sequence date". Babraham Bioinformatics. Babraham Institute. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- ↑ Kim, D.; Pertea, G.; Trapnell, C. et al. (2013). "TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions". Genome Biology 14: R36. doi:10.1186/gb-2013-14-4-r36. PMC PMC4053844. PMID 23618408. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4053844.
- ↑ Dobin, A.; Davis, C.A.; Schlesinger, F. et al. (2013). "STAR: Ultrafast universal RNA-seq aligner". Bioinformatics 29 (1): 15–21. doi:10.1093/bioinformatics/bts635. PMC PMC3530905. PMID 23104886. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3530905.
- ↑ Liao, Y.; Smyth, G.K.; Shi, W. (2013). "The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote". Nucleic Acids Research 41 (10): e108. doi:10.1093/nar/gkt214. PMC PMC3664803. PMID 23558742. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3664803.
- ↑ Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. (2010). "edgeR: A Bioconductor package for differential expression analysis of digital gene expression data". Bioinformatics 26 (1): 139–140. doi:10.1093/bioinformatics/btp616. PMC PMC2796818. PMID 19910308. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2796818.
- ↑ Kallio, M.A.; Tuimala, J.T.; Hupponen, T. et al. (2011). "Chipster: User-friendly analysis software for microarray and other high-throughput data". BMC Genomics 12: 507. doi:10.1186/1471-2164-12-507. PMC PMC3215701. PMID 21999641. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3215701.
- ↑ Wolstencroft, K.; Haines, R.; Fellows, D. et al. (2013). "The Taverna workflow suite: Designing and executing workflows of Web Services on the desktop, web or in the cloud". Nucleic Acids Research 41 (W1): W557-W561. doi:10.1093/nar/gkt328. PMC PMC3692062. PMID 23640334. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3692062.
- ↑ Edgar, R.; Barrett, T. (2006). "NCBI GEO standards and services for microarray data". Nature Biotechnology 24 (12): 1471–2. doi:10.1038/nbt1206-1471. PMC PMC2270403. PMID 17160034. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2270403.
- ↑ Barrett, T.; Wilhite, S.E.; Ledoux, P. et al. (2013). "NCBI GEO: Archive for functional genomics data sets - update". Nucleic Acids Research 41 (D1): D991-5. doi:10.1093/nar/gks1193. PMC PMC3531084. PMID 23193258. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3531084.
- ↑ NCBI Resource Coordinators (2015). "Database resources of the National Center for Biotechnology Information". Nucleic Acids Research 44 (D1): D7-D19. doi:10.1093/nar/gku1130. PMC PMC4383943. PMID 25398906. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4383943.
- ↑ Hong, E.L.; Sloan, C.A.; Chan, E.T. et al. (2016). "Principles of metadata organization at the ENCODE data coordination center". Database 2016: baw001. doi:10.1093/database/baw001. PMC PMC4792520. PMID 26980513. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4792520.
- ↑ Washington, N.L.; Stinson, E.O.; Perry, M.D. et al. (2011). "The modENCODE Data Coordination Center: Lessons in harvesting comprehensive experimental details". Database 2011: bar023. doi:10.1093/database/bar023. PMC PMC3170170. PMID 21856757. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3170170.
- ↑ Sansone, S.A.; Rocca-Serra, P.; Field, D. et al. (2012). "Toward interoperable bioscience data". Nature Genetics 44 (2): 121–126. doi:10.1038/ng.1054. PMC PMC3428019. PMID 22281772. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3428019.
- ↑ Türker, C.; Stolte, E.; Joho, D.; Schlapbach, R. (2007). "B-Fabric: A Data and Application Integration Framework for Life Sciences Research". In Cohen-Boulakia, S.; Tannen, V.. Data Integration in the Life Sciences. Springer Berlin Heidelberg. pp. 37–47. doi:10.1007/978-3-540-73255-6_6. ISBN 9783540732556.
- ↑ Türker, C.; Akal, F.; Joho, D.; Schlapbach, R. (2009). "B-Fabric: An Open Source Life Sciences Data Management System". In Winslett, M.. Scientific and Statistical Database Management. Springer Berlin Heidelberg. pp. 185–190. doi:10.1007/978-3-642-02279-1_13. ISBN 9783642022791.
- ↑ Dijkstra, E.W. (1982). Selected Writings on Computing: A Personal Perspective. Springer New York. pp. 362. ISBN 9781461256953.
- ↑ Buschmann, F.; Meunier, R.; Rohnert, H. et al. (1996). Pattern-Oriented Software Architecture: A System of Patterns. John Wiley & Sons, Inc. pp. 476. ISBN 9780471958697.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added.