Difference between revisions of "Biodiversity informatics"
Shawndouglas (talk | contribs) (Updated some content. Saving and updating more.) |
Shawndouglas (talk | contribs) (Updated content.) |
||
| Line 1: | Line 1: | ||
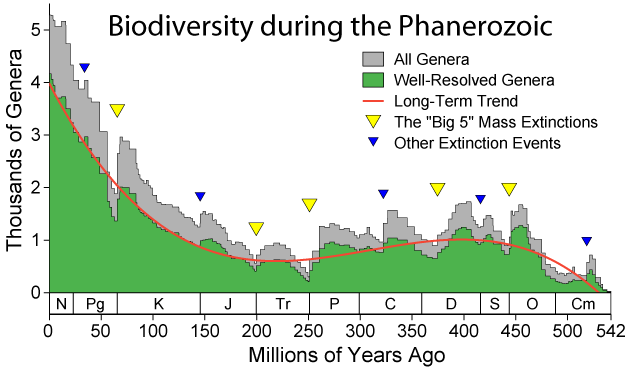
[[File:Phanerozoic Biodiversity.png|thumb|500px|right|Graphical representations of prehistoric biodiversity data like this are slowly becoming easier with the advancement of biodiversity informatics standards and tools.]] | |||
'''Biodiversity informatics''' is the application of informatics techniques to biodiversity [[information]] for improved management, presentation, discovery, exploration, and analysis. It typically builds on a foundation of taxonomic, biogeographic, and synecologic information stored in digital form, which, with the application of modern computer techniques, can yield new ways to view and analyze existing information, as well as predictive models for information that does not yet exist.<ref name="BerendsohnBio">{{cite journal |url=http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3234432/ |journal=ZooKeys |title=Biodiversity information platforms: From standards to interoperability |author=Berendsohn, W. G.; Güntsch, A.; Hoffmann, N.; Kohlbecker, A.; Luther, K.; Müller, A. |issue=150 |pages=71–87 |year=2011 |month=November |doi=10.3897/zookeys.150.2166 |pmc=3234432 |accessdate=18 June 2014}}</ref> | '''Biodiversity informatics''' is the application of informatics techniques to biodiversity [[information]] for improved management, presentation, discovery, exploration, and analysis. It typically builds on a foundation of taxonomic, biogeographic, and synecologic information stored in digital form, which, with the application of modern computer techniques, can yield new ways to view and analyze existing information, as well as predictive models for information that does not yet exist.<ref name="BerendsohnBio">{{cite journal |url=http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3234432/ |journal=ZooKeys |title=Biodiversity information platforms: From standards to interoperability |author=Berendsohn, W. G.; Güntsch, A.; Hoffmann, N.; Kohlbecker, A.; Luther, K.; Müller, A. |issue=150 |pages=71–87 |year=2011 |month=November |doi=10.3897/zookeys.150.2166 |pmc=3234432 |accessdate=18 June 2014}}</ref> | ||
| Line 4: | Line 5: | ||
==History== | ==History== | ||
According to correspondence reproduced by Walter Berendsohn<ref name="BITheTerm">{{cite web |url=http://www.bgbm.org/BioDivInf/TheTerm.htm |archiveurl=https://web.archive.org/web/20130511091435/http://www.bgbm.org/BioDivInf/TheTerm.htm |title="Biodiversity Informatics", The Term |author=Güntsch, Anton; Berendsohn, Walter |publisher=Botanic Garden and Botanical Museum Berlin-Dahlem |date=18 August 2010 |archivedate=11 May 2013 |accessdate=18 June 2014}}</ref>, the term "biodiversity informatics" was coined by John Whiting in 1992 to cover the activities of an entity known as the Canadian Biodiversity Informatics Consortium, a group involved with fusing basic biodiversity information with environmental economics and geospatial information. Subsequently it appears to have lost | According to correspondence reproduced by Walter Berendsohn<ref name="BITheTerm">{{cite web |url=http://www.bgbm.org/BioDivInf/TheTerm.htm |archiveurl=https://web.archive.org/web/20130511091435/http://www.bgbm.org/BioDivInf/TheTerm.htm |title="Biodiversity Informatics", The Term |author=Güntsch, Anton; Berendsohn, Walter |publisher=Botanic Garden and Botanical Museum Berlin-Dahlem |date=18 August 2010 |archivedate=11 May 2013 |accessdate=18 June 2014}}</ref>, the term "biodiversity informatics" was coined by John Whiting in 1992 to cover the activities of an entity known as the Canadian Biodiversity Informatics Consortium (CBIC), a group involved with fusing basic biodiversity information with environmental economics and geospatial information. Subsequently it appears to have lost at least some connection with the geospatial world, becoming more closely associated with the computerized management of biodiversity information.<ref name="BisbyRevo">{{cite journal |url=http://www.sciencemag.org/content/289/5488/2309.abstract |journal=Science |title=The Quiet Revolution: Biodiversity Informatics and the Internet |author=Bisby, Frank A. |volume=289 |issue=5488 |pages=2309–2312 |year=2000 |month=September |doi=10.1126/science.289.5488.2309 |pmid=11009408 |accessdate=18 June 2014}}</ref> However, modern efforts to document global biodiversity patterns and processes using georeferencing and other [[geoinformatics]] tools have re-emphasized some of the original spirit of the CBIC.<ref name="GuralnickBio">{{cite journal |url=http://bioinformatics.oxfordjournals.org/content/25/4/421.full |journal=Bioinformatics |title=Biodiversity Informatics: Automated Approaches for Documenting Global Biodiversity Patterns and Processes |author=Guralnick, R. P. |volume=25 |issue=4 |pages=421–428 |year=2009 |month=January |pmid=19129210 |doi=10.1093/bioinformatics/btn659}}</ref> | ||
Biodiversity informatics itself likely grew from the construction of the first computerized taxonomic databases in the early 1970s, progressing through the subsequent development of distributed search tools towards the late 1990s, including Species Analyst, the North American Biodiversity Information Network (NABIN), and CONABIO.<ref name="KrishtalkaCan">{{cite journal |url=http://bioscience.oxfordjournals.org/content/50/7/611.full |journal=BioScience |title=Can Natural History Museums Capture the Future? |author=Krishtalka, L.; Humphrey, P. S. |volume=50 |issue=7 |pages=611–617 |year=2000 |doi=10.1641/0006-3568(2000)050[0611:CNHMCT]2.0.CO;2 |accessdate=18 June 2014}}</ref> Other contributions came in the form of a variety of niche modeling tools and algorithms to process digitized biodiversity data from the mid-1980s onwards.<ref name="PetersonPredict">{{cite journal |url=http://www.cria.org.br/eventos/mfmpe/19_20jun2002_docs/BioScience%202001.pdf |journal=BioScience |title=Predicting Species Invasions Using Ecological Niche Modeling: New Approaches from Bioinformatics Attack a Pressing Problem |author=Peterson, A. T.; Vieglais, D. |volume=51 |issue=5 |pages=363–371 |year=2001 |month=May |doi=10.1641/0006-3568(2001)051[0363:PSIUEN]2.0.CO;2 |accessdate=18 June 2014}}</ref> | Biodiversity informatics itself likely grew from the construction of the first computerized taxonomic databases in the early 1970s, progressing through the subsequent development of distributed search tools towards the late 1990s, including Species Analyst, the North American Biodiversity Information Network (NABIN), and CONABIO.<ref name="KrishtalkaCan">{{cite journal |url=http://bioscience.oxfordjournals.org/content/50/7/611.full |journal=BioScience |title=Can Natural History Museums Capture the Future? |author=Krishtalka, L.; Humphrey, P. S. |volume=50 |issue=7 |pages=611–617 |year=2000 |doi=10.1641/0006-3568(2000)050[0611:CNHMCT]2.0.CO;2 |accessdate=18 June 2014}}</ref> Other contributions came in the form of a variety of niche modeling tools and algorithms to process digitized biodiversity data from the mid-1980s onwards.<ref name="PetersonPredict">{{cite journal |url=http://www.cria.org.br/eventos/mfmpe/19_20jun2002_docs/BioScience%202001.pdf |journal=BioScience |title=Predicting Species Invasions Using Ecological Niche Modeling: New Approaches from Bioinformatics Attack a Pressing Problem |author=Peterson, A. T.; Vieglais, D. |volume=51 |issue=5 |pages=363–371 |year=2001 |month=May |doi=10.1641/0006-3568(2001)051[0363:PSIUEN]2.0.CO;2 |accessdate=18 June 2014}}</ref> | ||
| Line 11: | Line 12: | ||
==Application== | ==Application== | ||
Biodiversity informatics can help tackle problems and tasks such as the following<ref name="eBiosphere" /><ref name="eBio09Reso">{{cite web |url=http://www.e-biosphere09.org/assets/files/workshop/Resolution.pdf |format=PDF |title=e-Biosphere 09 Planning Workshop - Resolution |publisher=Smithsonian Institution |date=05 June 2009 |accessdate=18 June 2014}}</ref><ref name="GordonHier">{{cite web |url=http://www.catalogueoflife.org/col/info/hierarchy |title=Towards a management hierarchy (classification) for the Catalogue of Life |author=Gordon, Dennis P. |publisher=Catalogue of Life |date=May 2009 |accessdate=18 June 2014}}</ref>: | |||
* the tracking of invasive species | |||
* the creation of new biodiversity mapping, infrastructure, and species identification models | |||
* the development of new modeling and data integration tools | |||
* the creation of global registries for the resources that are basic to biodiversity informatics | |||
* the construction of a solid global taxonomic infrastructure | |||
* the creation of ontologies for biodiversity data | |||
* the creation of a single-consensus classification system | |||
* the development of algorithms to cope with variant representations of identifiers such as species names and authorities | |||
* the transition of content in taxonomic databases to a machine-readable and -queryable format | |||
==Informatics== | ==Informatics== | ||
Providing online, coherent, standardized digital access to the vast collection of disparate primary biodiversity data is a task at the heart of regional and global biodiversity data networks. Secondary sources of biodiversity data, including relevant scientific literature, can be potentially parsed by specialized information retrieval algorithms to extract the relevant primary biodiversity information that is reported therein, sometimes in summary form, but more frequently as primary observations in narrative or tabular form.<ref name="BerendsohnBio" /> The Biodiversity Heritage Library is an example of this, aiming to digitize substantial portions of the out-of-copyright taxonomic literature, which is then subjected to OCR (optical character recognition) so as to be amenable to further processing.<ref name="BHLAbout">{{cite web |url=http://biodivlib.wikispaces.com/About |title=Biodiversity Heritage Library - About |publisher=Tangient LLC |date=18 June 2014 |accessdate=18 June 2014}}</ref> | |||
Like other data-related disciplines, biodiversity informatics benefits from the adoption of appropriate standards and protocols in order to support machine-machine transmission and interoperability of information within its particular domain. Examples of relevant standards include<ref name="BerendsohnBio" />: | |||
* [http://rs.tdwg.org/dwc/terms/guides/xml/ Darwin Core XML], an XML schema for specimen- and observation-based biodiversity data | |||
* [http://www.tdwg.org/standards/117/ Taxonomic Concept Transfer Schema], a schema for taxonomic information providers to exchange information with other such providers | |||
* [http://www.tdwg.org/standards/116/ Structured Descriptive Data], a standard for the capture, transport, caching and archiving of descriptive data | |||
* [http://www.tdwg.org/standards/115/ Access to Biological Collection Data], a standard for access to and exchange of data about specimens and observations | |||
* [http://www.tdwg.org/standards/449/ TDWG Access Protocol for Information Retrieval] (TAPIR), a request and response protocol for accessing structured biodiversity data | |||
==Further reading== | ==Further reading== | ||
| Line 51: | Line 42: | ||
* {{cite journal |url=http://arjournals.annualreviews.org/doi/abs/10.1146/annurev.ento.52.110405.091259 |journal=Annual Review of Entomology |title=Biodiversity Informatics |author=Johnson, N. F. |volume=52 |pages=421–438 |year=2007 |doi=10.1146/annurev.ento.52.110405.091259 |pmid=16956323}} | * {{cite journal |url=http://arjournals.annualreviews.org/doi/abs/10.1146/annurev.ento.52.110405.091259 |journal=Annual Review of Entomology |title=Biodiversity Informatics |author=Johnson, N. F. |volume=52 |pages=421–438 |year=2007 |doi=10.1146/annurev.ento.52.110405.091259 |pmid=16956323}} | ||
* {{cite journal |url=http://bib.oxfordjournals.org/content/8/5/347.full |journal=Briefings in Bioinformatics |title=Biodiversity Informatics: Organizing and Linking Information Across the Spectrum of Life |author=Sarkar, I. N. |volume=8 |issue=5 |pages=347–357 |year=2007 |month=August |pmid=17704120 |doi=10.1093/bib/bbm037}} | * {{cite journal |url=http://bib.oxfordjournals.org/content/8/5/347.full |journal=Briefings in Bioinformatics |title=Biodiversity Informatics: Organizing and Linking Information Across the Spectrum of Life |author=Sarkar, I. N. |volume=8 |issue=5 |pages=347–357 |year=2007 |month=August |pmid=17704120 |doi=10.1093/bib/bbm037}} | ||
==External links== | ==External links== | ||
* [http://www.biodiversitylibrary.org/ Biodiversity Heritage Library] | |||
* [http://journals.ku.edu/index.php/jbi Biodiversity Informatics] open-access journal | * [http://journals.ku.edu/index.php/jbi Biodiversity Informatics] open-access journal | ||
* [http://www.biomedcentral.com/bmcbioinformatics/supplements/10/S14 Biodiversity Informatics at BMC Bioinformatics] | * [http://www.biomedcentral.com/bmcbioinformatics/supplements/10/S14 Biodiversity Informatics at BMC Bioinformatics] | ||
* [http://www.tdwg.org/biodiv-projects Biodiversity Information Projects of the World] | * [http://www.tdwg.org/biodiv-projects Biodiversity Information Projects of the World] | ||
* [http://www.tdwg.org/ Biodiversity Information Standards] | * [http://www.tdwg.org/ Biodiversity Information Standards] | ||
* [http://www.catalogueoflife.org/ Catalogue of Life] | |||
* [http://eol.org/ Encyclopedia of Life] | |||
* [http://www.pensoftonline.net/zookeys ZooKeys] open-access journal | * [http://www.pensoftonline.net/zookeys ZooKeys] open-access journal | ||
==Notes== | ==Notes== | ||
This article | This article reuses some content from [http://en.wikipedia.org/wiki/Biodiversity_informatics the Wikipedia article]. | ||
==References== | ==References== | ||
Revision as of 22:29, 18 June 2014
Biodiversity informatics is the application of informatics techniques to biodiversity information for improved management, presentation, discovery, exploration, and analysis. It typically builds on a foundation of taxonomic, biogeographic, and synecologic information stored in digital form, which, with the application of modern computer techniques, can yield new ways to view and analyze existing information, as well as predictive models for information that does not yet exist.[1]
Biodiversity informatics has also been described by others as "the creation, integration, analysis, and understanding of information regarding biological diversity"[2] and a field of science "that brings information science and technologies to bear on the data and information generated by the study of organisms, their genes, and their interactions."[3]
History
According to correspondence reproduced by Walter Berendsohn[4], the term "biodiversity informatics" was coined by John Whiting in 1992 to cover the activities of an entity known as the Canadian Biodiversity Informatics Consortium (CBIC), a group involved with fusing basic biodiversity information with environmental economics and geospatial information. Subsequently it appears to have lost at least some connection with the geospatial world, becoming more closely associated with the computerized management of biodiversity information.[5] However, modern efforts to document global biodiversity patterns and processes using georeferencing and other geoinformatics tools have re-emphasized some of the original spirit of the CBIC.[6]
Biodiversity informatics itself likely grew from the construction of the first computerized taxonomic databases in the early 1970s, progressing through the subsequent development of distributed search tools towards the late 1990s, including Species Analyst, the North American Biodiversity Information Network (NABIN), and CONABIO.[7] Other contributions came in the form of a variety of niche modeling tools and algorithms to process digitized biodiversity data from the mid-1980s onwards.[8]
The U.S. journal Science devoted a special issue to "Bioinformatics for Biodiversity" in September 2000[9], the Global Biodiversity Information Facility (GBIF) was officially formed in 2001[10], the journal Biodiversity Informatics commenced publication in 2004, and several international conferences brought together biodiversity researchers during the twenty-first century.[3][11]
Application
Biodiversity informatics can help tackle problems and tasks such as the following[3][12][13]:
- the tracking of invasive species
- the creation of new biodiversity mapping, infrastructure, and species identification models
- the development of new modeling and data integration tools
- the creation of global registries for the resources that are basic to biodiversity informatics
- the construction of a solid global taxonomic infrastructure
- the creation of ontologies for biodiversity data
- the creation of a single-consensus classification system
- the development of algorithms to cope with variant representations of identifiers such as species names and authorities
- the transition of content in taxonomic databases to a machine-readable and -queryable format
Informatics
Providing online, coherent, standardized digital access to the vast collection of disparate primary biodiversity data is a task at the heart of regional and global biodiversity data networks. Secondary sources of biodiversity data, including relevant scientific literature, can be potentially parsed by specialized information retrieval algorithms to extract the relevant primary biodiversity information that is reported therein, sometimes in summary form, but more frequently as primary observations in narrative or tabular form.[1] The Biodiversity Heritage Library is an example of this, aiming to digitize substantial portions of the out-of-copyright taxonomic literature, which is then subjected to OCR (optical character recognition) so as to be amenable to further processing.[14]
Like other data-related disciplines, biodiversity informatics benefits from the adoption of appropriate standards and protocols in order to support machine-machine transmission and interoperability of information within its particular domain. Examples of relevant standards include[1]:
- Darwin Core XML, an XML schema for specimen- and observation-based biodiversity data
- Taxonomic Concept Transfer Schema, a schema for taxonomic information providers to exchange information with other such providers
- Structured Descriptive Data, a standard for the capture, transport, caching and archiving of descriptive data
- Access to Biological Collection Data, a standard for access to and exchange of data about specimens and observations
- TDWG Access Protocol for Information Retrieval (TAPIR), a request and response protocol for accessing structured biodiversity data
Further reading
- OECD Megascience Forum Working Group on Biological Informatics (January 1999). "Final Report of the OECD Megascience Forum Working Group on Biological Informatics, January 1999" (PDF). OECD. pp. 1–74. http://www.oecd.org/science/sci-tech/2105199.pdf.
- Canhos, V. P.; Souza, S.; Giovanni, R.; Canhos, D. A. L. (2004). "Global Biodiversity Informatics: Setting the Scene for a "New World" of Ecological Modeling" (PDF). Biodiversity Informatics 1: 1–13. https://journals.ku.edu/index.php/jbi/article/viewFile/3/1.
- Soberón, J.; Peterson, A. T. (March 2004). "Biodiversity Informatics: Managing and Applying Primary Biodiversity Data" (PDF). Philosophical Transactions of the Royal Society of London B359: 689–698. doi:10.1098/rstb.2003.1439. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1693343/pdf/15253354.pdf.
- Chapman, A. D. (2005). Uses of Primary Species-Occurrence Data. Copenhagen: Global Biodiversity Information Facility. pp. 1–106. http://www.gbif.org/resources/2834.
- Johnson, N. F. (2007). "Biodiversity Informatics". Annual Review of Entomology 52: 421–438. doi:10.1146/annurev.ento.52.110405.091259. PMID 16956323. http://arjournals.annualreviews.org/doi/abs/10.1146/annurev.ento.52.110405.091259.
- Sarkar, I. N. (August 2007). "Biodiversity Informatics: Organizing and Linking Information Across the Spectrum of Life". Briefings in Bioinformatics 8 (5): 347–357. doi:10.1093/bib/bbm037. PMID 17704120. http://bib.oxfordjournals.org/content/8/5/347.full.
External links
- Biodiversity Heritage Library
- Biodiversity Informatics open-access journal
- Biodiversity Informatics at BMC Bioinformatics
- Biodiversity Information Projects of the World
- Biodiversity Information Standards
- Catalogue of Life
- Encyclopedia of Life
- ZooKeys open-access journal
Notes
This article reuses some content from the Wikipedia article.
References
- ↑ 1.0 1.1 1.2 Berendsohn, W. G.; Güntsch, A.; Hoffmann, N.; Kohlbecker, A.; Luther, K.; Müller, A. (November 2011). "Biodiversity information platforms: From standards to interoperability". ZooKeys (150): 71–87. doi:10.3897/zookeys.150.2166. PMC 3234432. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3234432/. Retrieved 18 June 2014.
- ↑ "Biodiversity Informatics". University of Kansas Libraries. https://journals.ku.edu/index.php/jbi. Retrieved 18 June 2014.
- ↑ 3.0 3.1 3.2 "e-Biosphere '09: International Conference on Biodiversity Informatics". Smithsonian Institution. 2009. http://www.e-biosphere09.org/. Retrieved 18 June 2014.
- ↑ Güntsch, Anton; Berendsohn, Walter (18 August 2010). ""Biodiversity Informatics", The Term". Botanic Garden and Botanical Museum Berlin-Dahlem. Archived from the original on 11 May 2013. https://web.archive.org/web/20130511091435/http://www.bgbm.org/BioDivInf/TheTerm.htm. Retrieved 18 June 2014.
- ↑ Bisby, Frank A. (September 2000). "The Quiet Revolution: Biodiversity Informatics and the Internet". Science 289 (5488): 2309–2312. doi:10.1126/science.289.5488.2309. PMID 11009408. http://www.sciencemag.org/content/289/5488/2309.abstract. Retrieved 18 June 2014.
- ↑ Guralnick, R. P. (January 2009). "Biodiversity Informatics: Automated Approaches for Documenting Global Biodiversity Patterns and Processes". Bioinformatics 25 (4): 421–428. doi:10.1093/bioinformatics/btn659. PMID 19129210. http://bioinformatics.oxfordjournals.org/content/25/4/421.full.
- ↑ Krishtalka, L.; Humphrey, P. S. (2000). "Can Natural History Museums Capture the Future?". BioScience 50 (7): 611–617. doi:10.1641/0006-3568(2000)050[0611:CNHMCT]2.0.CO;2. http://bioscience.oxfordjournals.org/content/50/7/611.full. Retrieved 18 June 2014.
- ↑ Peterson, A. T.; Vieglais, D. (May 2001). "Predicting Species Invasions Using Ecological Niche Modeling: New Approaches from Bioinformatics Attack a Pressing Problem". BioScience 51 (5): 363–371. doi:10.1641/0006-3568(2001)051[0363:PSIUEN]2.0.CO;2. http://www.cria.org.br/eventos/mfmpe/19_20jun2002_docs/BioScience%202001.pdf. Retrieved 18 June 2014.
- ↑ "Bioinformatics for Biodiversity". Science 289 (5488): 2229–2440. September 2000. http://www.sciencemag.org/content/289/5488.toc. Retrieved 18 June 2014.
- ↑ "What is GBIF?". GBIF. http://www.gbif.org/whatisgbif. Retrieved 18 June 2014.
- ↑ "Biodiversity Informatics Horizons 2013". LifeWatch. 2013. http://conference.lifewatch.unisalento.it/index.php/EBIC/index/index. Retrieved 18 June 2014.
- ↑ "e-Biosphere 09 Planning Workshop - Resolution" (PDF). Smithsonian Institution. 5 June 2009. http://www.e-biosphere09.org/assets/files/workshop/Resolution.pdf. Retrieved 18 June 2014.
- ↑ Gordon, Dennis P. (May 2009). "Towards a management hierarchy (classification) for the Catalogue of Life". Catalogue of Life. http://www.catalogueoflife.org/col/info/hierarchy. Retrieved 18 June 2014.
- ↑ "Biodiversity Heritage Library - About". Tangient LLC. 18 June 2014. http://biodivlib.wikispaces.com/About. Retrieved 18 June 2014.