Difference between revisions of "Severe acute respiratory syndrome coronavirus 2"
Shawndouglas (talk | contribs) (Created as needed.) |
(No difference)
|
Latest revision as of 20:05, 25 March 2020
| Severe acute respiratory syndrome coronavirus 2 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|

| |||||||||||
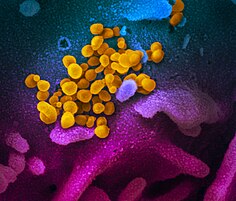
| Colourised transmission electron micrograph of SARS-CoV-2 virions with visible coronae | |||||||||||
| |||||||||||
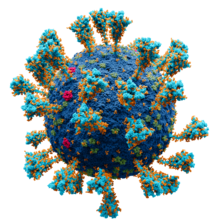
Model of the external structure of the SARS-CoV-2 virion[1]
| |||||||||||
| Virus classification | |||||||||||
| (unranked): | Virus | ||||||||||
| Realm: | Riboviria | ||||||||||
| Kingdom: | Orthornavirae | ||||||||||
| Phylum: | Pisuviricota | ||||||||||
| Class: | Pisoniviricetes | ||||||||||
| Order: | Nidovirales | ||||||||||
| Family: | Coronaviridae | ||||||||||
| Genus: | Betacoronavirus | ||||||||||
| Subgenus: | Sarbecovirus | ||||||||||
| Species: | |||||||||||
| Virus: | Severe acute respiratory syndrome coronavirus 2
| ||||||||||
| Notable variants | |||||||||||
| Synonyms | |||||||||||
| |||||||||||
| Part of a series on the |
| COVID-19 pandemic |
|---|
 |
|
|
|
Severe acute respiratory syndrome coronavirus 2 (SARS‑CoV‑2)[2] is a strain of coronavirus that causes COVID-19, the respiratory illness responsible for the COVID-19 pandemic.[3] The virus previously had the provisional name 2019 novel coronavirus (2019-nCoV),[4][5][6][7] and has also been called human coronavirus 2019 (HCoV-19 or hCoV-19).[8][9][10][11] First identified in the city of Wuhan, Hubei, China, the World Health Organization designated the outbreak a public health emergency of international concern from January 30, 2020, to May 5, 2023.[12][13][14] SARS‑CoV‑2 is a positive-sense single-stranded RNA virus[15] that is contagious in humans.[16]
SARS‑CoV‑2 is a strain of the species severe-acute-respiratory-syndrome-related coronavirus (SARSr-CoV), as is SARS-CoV-1, the virus that caused the 2002–2004 SARS outbreak.[2][17] There are animal-borne coronavirus strains more closely related to SARS-CoV-2, the most closely known relative being the BANAL-52 bat coronavirus. SARS-CoV-2 is of zoonotic origin; its close genetic similarity to bat coronaviruses suggests it emerged from such a bat-borne virus.[18] Research is ongoing as to whether SARS‑CoV‑2 came directly from bats or indirectly through any intermediate hosts.[19] The virus shows little genetic diversity, indicating that the spillover event introducing SARS‑CoV‑2 to humans is likely to have occurred in late 2019.[20]
Epidemiological studies estimate that in the period between December 2019 and September 2020 each infection resulted in an average of 2.4–3.4 new infections when no members of the community were immune and no preventive measures were taken.[21] However, some subsequent variants have become more infectious.[22] The virus is airborne and primarily spreads between people through close contact and via aerosols and respiratory droplets that are exhaled when talking, breathing, or otherwise exhaling, as well as those produced from coughs and sneezes.[23][24] It enters human cells by binding to angiotensin-converting enzyme 2 (ACE2), a membrane protein that regulates the renin–angiotensin system.[25][26]
Terminology

During the initial outbreak in Wuhan, China, various names were used for the virus; some names used by different sources included "the coronavirus" or "Wuhan coronavirus".[27][28] In January 2020, the World Health Organization (WHO) recommended "2019 novel coronavirus" (2019-nCoV)[5][29] as the provisional name for the virus. This was in accordance with WHO's 2015 guidance[30] against using geographical locations, animal species, or groups of people in disease and virus names.[31][32]
On 11 February 2020, the International Committee on Taxonomy of Viruses adopted the official name "severe acute respiratory syndrome coronavirus 2" (SARS‑CoV‑2).[33] To avoid confusion with the disease SARS, the WHO sometimes refers to SARS‑CoV‑2 as "the COVID-19 virus" in public health communications[34][35] and the name HCoV-19 was included in some research articles.[8][9][10] Referring to COVID-19 as the "Wuhan virus" has been described as dangerous by WHO officials, and as xenophobic by many journalists and academics.[36][37][38]
Infection and transmission
This section has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
|
Human-to-human transmission of SARS‑CoV‑2 was confirmed on 20 January 2020 during the COVID-19 pandemic.[16][39][40][41] Transmission was initially assumed to occur primarily via respiratory droplets from coughs and sneezes within a range of about 1.8 metres (6 ft).[42][43] Laser light scattering experiments suggest that speaking is an additional mode of transmission[44][45] and a far-reaching[46] one, indoors, with little air flow.[47][48] Other studies have suggested that the virus may be airborne as well, with aerosols potentially being able to transmit the virus.[49][50][51] During human-to-human transmission, between 200 and 800 infectious SARS‑CoV‑2 virions are thought to initiate a new infection.[52][53][54] If confirmed, aerosol transmission has biosafety implications because a major concern associated with the risk of working with emerging viruses in the laboratory is the generation of aerosols from various laboratory activities which are not immediately recognizable and may affect other scientific personnel.[55] Indirect contact via contaminated surfaces is another possible cause of infection.[56] Preliminary research indicates that the virus may remain viable on plastic (polypropylene) and stainless steel (AISI 304) for up to three days, but it does not survive on cardboard for more than one day or on copper for more than four hours.[10] The virus is inactivated by soap, which destabilizes its lipid bilayer.[57][58] Viral RNA has also been found in stool samples and semen from infected individuals.[59][60]
The degree to which the virus is infectious during the incubation period is uncertain, but research has indicated that the pharynx reaches peak viral load approximately four days after infection[61][62] or in the first week of symptoms and declines thereafter.[63] The duration of SARS-CoV-2 RNA shedding is generally between 3 and 46 days after symptom onset.[64]
A study by a team of researchers from the University of North Carolina found that the nasal cavity is seemingly the dominant initial site of infection, with subsequent aspiration-mediated virus-seeding into the lungs in SARS‑CoV‑2 pathogenesis.[65] They found that there was an infection gradient from high in proximal towards low in distal pulmonary epithelial cultures, with a focal infection in ciliated cells and type 2 pneumocytes in the airway and alveolar regions respectively.[65]
Studies have identified a range of animals—such as cats, ferrets, hamsters, non-human primates, minks, tree shrews, raccoon dogs, fruit bats, and rabbits—that are susceptible and permissive to SARS-CoV-2 infection.[66][67][68] Some institutions have advised that those infected with SARS‑CoV‑2 restrict their contact with animals.[69][70]
Asymptomatic and presymptomatic transmission
On 1 February 2020, the World Health Organization (WHO) indicated that "transmission from asymptomatic cases is likely not a major driver of transmission".[71] One meta-analysis found that 17% of infections are asymptomatic, and asymptomatic individuals were 42% less likely to transmit the virus.[72]
However, an epidemiological model of the beginning of the outbreak in China suggested that "pre-symptomatic shedding may be typical among documented infections" and that subclinical infections may have been the source of a majority of infections.[73] That may explain how out of 217 on board a cruise liner that docked at Montevideo, only 24 of 128 who tested positive for viral RNA showed symptoms.[74] Similarly, a study of ninety-four patients hospitalized in January and February 2020 estimated patients began shedding virus two to three days before symptoms appear and that "a substantial proportion of transmission probably occurred before first symptoms in the index case".[53] The authors later published a correction that showed that shedding began earlier than first estimated, four to five days before symptoms appear.[75]
Reinfection
There is uncertainty about reinfection and long-term immunity.[76] It is not known how common reinfection is, but reports have indicated that it is occurring with variable severity.[76]
The first reported case of reinfection was a 33-year-old man from Hong Kong who first tested positive on 26 March 2020, was discharged on 15 April 2020 after two negative tests, and tested positive again on 15 August 2020 (142 days later), which was confirmed by whole-genome sequencing showing that the viral genomes between the episodes belong to different clades.[77] The findings had the implications that herd immunity may not eliminate the virus if reinfection is not an uncommon occurrence and that vaccines may not be able to provide lifelong protection against the virus.[77]
Another case study described a 25-year-old man from Nevada who tested positive for SARS‑CoV‑2 on 18 April 2020 and on 5 June 2020 (separated by two negative tests). Since genomic analyses showed significant genetic differences between the SARS‑CoV‑2 variant sampled on those two dates, the case study authors determined this was a reinfection.[78] The man's second infection was symptomatically more severe than the first infection, but the mechanisms that could account for this are not known.[78]
Reservoir and origin

No natural reservoir for SARS-CoV-2 has been identified.[79] Prior to the emergence of SARS-CoV-2 as a pathogen infecting humans, there had been two previous zoonosis-based coronavirus epidemics, those caused by SARS-CoV-1 and MERS-CoV.[18]
The first known infections from SARS‑CoV‑2 were discovered in Wuhan, China.[80] The original source of viral transmission to humans remains unclear, as does whether the virus became pathogenic before or after the spillover event.[9][20][81] Because many of the early infectees were workers at the Huanan Seafood Market,[82][83] it has been suggested that the virus might have originated from the market.[9][84] However, other research indicates that visitors may have introduced the virus to the market, which then facilitated rapid expansion of the infections.[20][85] A March 2021 WHO-convened report stated that human spillover via an intermediate animal host was the most likely explanation, with direct spillover from bats next most likely. Introduction through the food supply chain and the Huanan Seafood Market was considered another possible, but less likely, explanation.[86] An analysis in November 2021, however, said that the earliest-known case had been misidentified and that the preponderance of early cases linked to the Huanan Market argued for it being the source.[87]
For a virus recently acquired through a cross-species transmission, rapid evolution is expected.[88] The mutation rate estimated from early cases of SARS-CoV-2 was of 6.54×10−4 per site per year.[86] Coronaviruses in general have high genetic plasticity,[89] but SARS-CoV-2's viral evolution is slowed by the RNA proofreading capability of its replication machinery.[90] For comparison, the viral mutation rate in vivo of SARS-CoV-2 has been found to be lower than that of influenza.[91]
Research into the natural reservoir of the virus that caused the 2002–2004 SARS outbreak has resulted in the discovery of many SARS-like bat coronaviruses, most originating in horseshoe bats. The closest match by far, published in Nature (journal) in February 2022, were viruses BANAL-52 (96.8% resemblance to SARS‑CoV‑2), BANAL-103 and BANAL-236, collected in three different species of bats in Feuang, Laos.[92][93][94] An earlier source published in February 2020 identified the virus RaTG13, collected in bats in Mojiang, Yunnan, China to be the closest to SARS‑CoV‑2, with 96.1% resemblance.[80][95] None of the above are its direct ancestor.[96]

Bats are considered the most likely natural reservoir of SARS‑CoV‑2.[86][97] Differences between the bat coronavirus and SARS‑CoV‑2 suggest that humans may have been infected via an intermediate host;[84] although the source of introduction into humans remains unknown.[98][79]
Although the role of pangolins as an intermediate host was initially posited (a study published in July 2020 suggested that pangolins are an intermediate host of SARS‑CoV‑2-like coronaviruses[99][100]), subsequent studies have not substantiated their contribution to the spillover.[86] Evidence against this hypothesis includes the fact that pangolin virus samples are too distant to SARS-CoV-2: isolates obtained from pangolins seized in Guangdong were only 92% identical in sequence to the SARS‑CoV‑2 genome (matches above 90 percent may sound high, but in genomic terms it is a wide evolutionary gap[101]). In addition, despite similarities in a few critical amino acids,[102] pangolin virus samples exhibit poor binding to the human ACE2 receptor.[103]
Phylogenetics and taxonomy
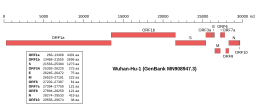 Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2 | |
| NCBI genome ID | 86693 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
| Genome browser (UCSC) | |
SARS‑CoV‑2 belongs to the broad family of viruses known as coronaviruses.[28] It is a positive-sense single-stranded RNA (+ssRNA) virus, with a single linear RNA segment. Coronaviruses infect humans, other mammals, including livestock and companion animals, and avian species.[104] Human coronaviruses are capable of causing illnesses ranging from the common cold to more severe diseases such as Middle East respiratory syndrome (MERS, fatality rate ~34%). SARS-CoV-2 is the seventh known coronavirus to infect people, after 229E, NL63, OC43, HKU1, MERS-CoV, and the original SARS-CoV.[105]
Like the SARS-related coronavirus implicated in the 2003 SARS outbreak, SARS‑CoV‑2 is a member of the subgenus Sarbecovirus (beta-CoV lineage B).[106][107] Coronaviruses undergo frequent recombination.[108] The mechanism of recombination in unsegmented RNA viruses such as SARS-CoV-2 is generally by copy-choice replication, in which gene material switches from one RNA template molecule to another during replication.[109] The SARS-CoV-2 RNA sequence is approximately 30,000 bases in length,[110] relatively long for a coronavirus—which in turn carry the largest genomes among all RNA families.[111] Its genome consists nearly entirely of protein-coding sequences, a trait shared with other coronaviruses.[108]

A distinguishing feature of SARS‑CoV‑2 is its incorporation of a polybasic site cleaved by furin,[102][112] which appears to be an important element enhancing its virulence.[113] It was suggested that the acquisition of the furin-cleavage site in the SARS-CoV-2 S protein was essential for zoonotic transfer to humans.[114] The furin protease recognizes the canonical peptide sequence RX[R/K] R↓X where the cleavage site is indicated by a down arrow and X is any amino acid.[115][116] In SARS-CoV-2 the recognition site is formed by the incorporated 12 codon nucleotide sequence CCT CGG CGG GCA which corresponds to the amino acid sequence P RR A.[117] This sequence is upstream of an arginine and serine which forms the S1/S2 cleavage site (P RR A R↓S) of the spike protein.[118] Although such sites are a common naturally-occurring feature of other viruses within the Subfamily Orthocoronavirinae,[117] it appears in few other viruses from the Beta-CoV genus,[119] and it is unique among members of its subgenus for such a site.[102] The furin cleavage site PRRAR↓ is highly similar to that of the feline coronavirus, an alphacoronavirus 1 strain.[120]
Viral genetic sequence data can provide critical information about whether viruses separated by time and space are likely to be epidemiologically linked.[121] With a sufficient number of sequenced genomes, it is possible to reconstruct a phylogenetic tree of the mutation history of a family of viruses. By 12 January 2020, five genomes of SARS‑CoV‑2 had been isolated from Wuhan and reported by the Chinese Center for Disease Control and Prevention (CCDC) and other institutions;[110][122] the number of genomes increased to 42 by 30 January 2020.[123] A phylogenetic analysis of those samples showed they were "highly related with at most seven mutations relative to a common ancestor", implying that the first human infection occurred in November or December 2019.[123] Examination of the topology of the phylogenetic tree at the start of the pandemic also found high similarities between human isolates.[124] As of 21 August 2021,[update] 3,422 SARS‑CoV‑2 genomes, belonging to 19 strains, sampled on all continents except Antarctica were publicly available.[125]
On 11 February 2020, the International Committee on Taxonomy of Viruses announced that according to existing rules that compute hierarchical relationships among coronaviruses based on five conserved sequences of nucleic acids, the differences between what was then called 2019-nCoV and the virus from the 2003 SARS outbreak were insufficient to make them separate viral species. Therefore, they identified 2019-nCoV as a virus of Severe acute respiratory syndrome–related coronavirus.[126]
In July 2020, scientists reported that a more infectious SARS‑CoV‑2 variant with spike protein variant G614 has replaced D614 as the dominant form in the pandemic.[127][128]
Coronavirus genomes and subgenomes encode six open reading frames (ORFs).[129] In October 2020, researchers discovered a possible overlapping gene named ORF3d, in the SARS‑CoV‑2 genome. It is unknown if the protein produced by ORF3d has any function, but it provokes a strong immune response. ORF3d has been identified before, in a variant of coronavirus that infects pangolins.[130][131]
Phylogenetic tree
A phylogenetic tree based on whole-genome sequences of SARS-CoV-2 and related coronaviruses is:[132][133]
| SARS‑CoV‑2 related coronavirus |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
SARS-CoV-1, 79% to SARS-CoV-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Variants
This section needs to be updated. (April 2023) |

There are many thousands of variants of SARS-CoV-2, which can be grouped into the much larger clades.[142] Several different clade nomenclatures have been proposed. Nextstrain divides the variants into five clades (19A, 19B, 20A, 20B, and 20C), while GISAID divides them into seven (L, O, V, S, G, GH, and GR).[143]
Several notable variants of SARS-CoV-2 emerged in late 2020. The World Health Organization has currently declared five variants of concern, which are as follows:[144]
- Alpha: Lineage B.1.1.7 emerged in the United Kingdom in September 2020, with evidence of increased transmissibility and virulence. Notable mutations include N501Y and P681H.
- An E484K mutation in some lineage B.1.1.7 virions has been noted and is also tracked by various public health agencies.
- Beta: Lineage B.1.351 emerged in South Africa in May 2020, with evidence of increased transmissibility and changes to antigenicity, with some public health officials raising alarms about its impact on the efficacy of some vaccines. Notable mutations include K417N, E484K and N501Y.
- Gamma: Lineage P.1 emerged in Brazil in November 2020, also with evidence of increased transmissibility and virulence, alongside changes to antigenicity. Similar concerns about vaccine efficacy have been raised. Notable mutations also include K417N, E484K and N501Y.
- Delta: Lineage B.1.617.2 emerged in India in October 2020. There is also evidence of increased transmissibility and changes to antigenicity.
- Omicron: Lineage B.1.1.529 emerged in Botswana in November 2021.
Other notable variants include 6 other WHO-designated variants under investigation and Cluster 5, which emerged among mink in Denmark and resulted in a mink euthanasia campaign rendering it virtually extinct.[145]
Virology
Virus structure

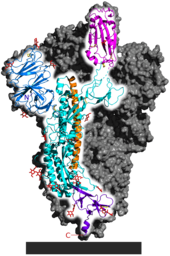
Each SARS-CoV-2 virion is 60–140 nanometres (2.4×10−6–5.5×10−6 in) in diameter;[105][83] its mass within the global human populace has been estimated as being between 0.1 and 10 kilograms.[146] Like other coronaviruses, SARS-CoV-2 has four structural proteins, known as the S (spike), E (envelope), M (membrane), and N (nucleocapsid) proteins; the N protein holds the RNA genome, and the S, E, and M proteins together create the viral envelope.[147] Coronavirus S proteins are glycoproteins and also type I membrane proteins (membranes containing a single transmembrane domain oriented on the extracellular side).[114] They are divided into two functional parts (S1 and S2).[104] In SARS-CoV-2, the spike protein, which has been imaged at the atomic level using cryogenic electron microscopy,[148][149] is the protein responsible for allowing the virus to attach to and fuse with the membrane of a host cell;[147] specifically, its S1 subunit catalyzes attachment, the S2 subunit fusion.[150]
Genome
As of early 2022, about 7 million SARS-CoV-2 genomes had been sequenced and deposited into public databases and another 800,000 or so were added each month.[151] By September 2023, the GISAID EpiCoV database contained more than 16 million genome sequences.[152]
SARS-CoV-2 has a linear, positive-sense, single-stranded RNA genome about 30,000 bases long.[104] Its genome has a bias against cytosine (C) and guanine (G) nucleotides, like other coronaviruses.[153] The genome has the highest composition of U (32.2%), followed by A (29.9%), and a similar composition of G (19.6%) and C (18.3%).[154] The nucleotide bias arises from the mutation of guanines and cytosines to adenosines and uracils, respectively.[155] The mutation of CG dinucleotides is thought to arise to avoid the zinc finger antiviral protein related defense mechanism of cells,[156] and to lower the energy to unbind the genome during replication and translation (adenosine and uracil base pair via two hydrogen bonds, cytosine and guanine via three).[155] The depletion of CG dinucleotides in its genome has led the virus to have a noticeable codon usage bias. For instance, arginine's six different codons have a relative synonymous codon usage of AGA (2.67), CGU (1.46), AGG (.81), CGC (.58), CGA (.29), and CGG (.19).[154] A similar codon usage bias trend is seen in other SARS–related coronaviruses.[157]
Replication cycle
Virus infections start when viral particles bind to host surface cellular receptors.[158] Protein modeling experiments on the spike protein of the virus soon suggested that SARS‑CoV‑2 has sufficient affinity to the receptor angiotensin converting enzyme 2 (ACE2) on human cells to use them as a mechanism of cell entry.[159] By 22 January 2020, a group in China working with the full virus genome and a group in the United States using reverse genetics methods independently and experimentally demonstrated that ACE2 could act as the receptor for SARS‑CoV‑2.[80][160][161][162] Studies have shown that SARS‑CoV‑2 has a higher affinity to human ACE2 than the original SARS virus.[148][163] SARS‑CoV‑2 may also use basigin to assist in cell entry.[164]
Initial spike protein priming by transmembrane protease, serine 2 (TMPRSS2) is essential for entry of SARS‑CoV‑2.[25] The host protein neuropilin 1 (NRP1) may aid the virus in host cell entry using ACE2.[165] After a SARS‑CoV‑2 virion attaches to a target cell, the cell's TMPRSS2 cuts open the spike protein of the virus, exposing a fusion peptide in the S2 subunit, and the host receptor ACE2.[150] After fusion, an endosome forms around the virion, separating it from the rest of the host cell. The virion escapes when the pH of the endosome drops or when cathepsin, a host cysteine protease, cleaves it.[150] The virion then releases RNA into the cell and forces the cell to produce and disseminate copies of the virus, which infect more cells.[166]
SARS‑CoV‑2 produces at least three virulence factors that promote shedding of new virions from host cells and inhibit immune response.[147] Whether they include downregulation of ACE2, as seen in similar coronaviruses, remains under investigation (as of May 2020).[167]
Treatment and drug development
Very few drugs are known to effectively inhibit SARS‑CoV‑2. Masitinib is a clinically safe drug and was recently found to inhibit its main protease, 3CLpro and showed >200-fold reduction in viral titers in the lungs and nose in mice. However, it is not approved for the treatment of COVID-19 in humans as of August 2021.[168][needs update] In December 2021, the United States granted emergency use authorization to Nirmatrelvir/ritonavir for the treatment of the virus;[169] the European Union, United Kingdom, and Canada followed suit with full authorization soon after.[170][171][172] One study found that Nirmatrelvir/ritonavir reduced the risk of hospitalization and death by 88%.[173]
COVID Moonshot is an international collaborative open-science project started in March 2020 with the goal of developing an un-patented oral antiviral drug for treatment of SARS-CoV-2.[174]
Epidemiology
Retrospective tests collected within the Chinese surveillance system revealed no clear indication of substantial unrecognized circulation of SARS‑CoV‑2 in Wuhan during the latter part of 2019.[86]
A meta-analysis from November 2020 estimated the basic reproduction number () of the virus to be between 2.39 and 3.44.[21] This means each infection from the virus is expected to result in 2.39 to 3.44 new infections when no members of the community are immune and no preventive measures are taken. The reproduction number may be higher in densely populated conditions such as those found on cruise ships.[175] Human behavior affects the R0 value and hence estimates of R0 differ between different countries, cultures, and social norms. For instance, one study found relatively low R0 (~3.5) in Sweden, Belgium and the Netherlands, while Spain and the US had significantly higher R0 values (5.9 to 6.4, respectively).[176]
| Variant | R0 | Source |
|---|---|---|
| Reference/ancestral strain | ~2.8 | [177] |
| Alpha (B.1.1.7) | (40-90% higher than previous variants) | [178] |
| Delta (B.1.617.2) | ~5 (3-8) | [179] |
There have been about 96,000 confirmed cases of infection in mainland China.[180] While the proportion of infections that result in confirmed cases or progress to diagnosable disease remains unclear,[181] one mathematical model estimated that 75,815 people were infected on 25 January 2020 in Wuhan alone, at a time when the number of confirmed cases worldwide was only 2,015.[182] Before 24 February 2020, over 95% of all deaths from COVID-19 worldwide had occurred in Hubei province, where Wuhan is located.[183][184] As of 10 March 2023, the percentage had decreased to 0.047%.[180]
As of 10 March 2023, there were 676,609,955 total confirmed cases of SARS‑CoV‑2 infection.[180] The total number of deaths attributed to the virus was 6,881,955.[180]
See also
- 3C-like protease (NS5)
References
- ^ Solodovnikov A, Arkhipova V (29 July 2021). "Достоверно красиво: как мы сделали 3D-модель SARS-CoV-2" [Truly beautiful: how we made the SARS-CoV-2 3D model] (in Russian). N+1. Archived from the original on 30 July 2021. Retrieved 30 July 2021.
- ^ Zimmer C (26 February 2021). "The Secret Life of a Coronavirus – An oily, 100-nanometer-wide bubble of genes has killed more than two million people and reshaped the world. Scientists don't quite know what to make of it". The New York Times. Archived from the original on 27 February 2021. Retrieved 28 February 2021.
- ^ Surveillance case definitions for human infection with novel coronavirus (nCoV): interim guidance v1, January 2020 (Report). World Health Organization. January 2020. hdl:10665/330376. WHO/2019-nCoV/Surveillance/v2020.1.
- ^ a b "Healthcare Professionals: Frequently Asked Questions and Answers". United States Centers for Disease Control and Prevention (CDC). 11 February 2020. Archived from the original on 14 February 2020. Retrieved 15 February 2020.
- ^ "About Novel Coronavirus (2019-nCoV)". United States Centers for Disease Control and Prevention (CDC). 11 February 2020. Archived from the original on 11 February 2020. Retrieved 25 February 2020.
- ^ Harmon A (4 March 2020). "We Spoke to Six Americans with Coronavirus". The New York Times. Archived from the original on 13 March 2020. Retrieved 16 March 2020.
- ^ a b Wong G, Bi YH, Wang QH, Chen XW, Zhang ZG, Yao YG (May 2020). "Zoonotic origins of human coronavirus 2019 (HCoV-19 / SARS-CoV-2): why is this work important?". Zoological Research. 41 (3): 213–219. doi:10.24272/j.issn.2095-8137.2020.031. PMC 7231470. PMID 32314559.
- ^ a b c d Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF (April 2020). "The proximal origin of SARS-CoV-2". Nature Medicine. 26 (4): 450–452. doi:10.1038/s41591-020-0820-9. PMC 7095063. PMID 32284615.
- ^ a b c van Doremalen N, Bushmaker T, Morris DH, Holbrook MG, Gamble A, Williamson BN, Tamin A, Harcourt JL, Thornburg NJ, Gerber SI, Lloyd-Smith JO, de Wit E, Munster VJ (April 2020). "Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1". The New England Journal of Medicine. 382 (16): 1564–1567. doi:10.1056/NEJMc2004973. PMC 7121658. PMID 32182409.
- ^ "hCoV-19 Database". China National GeneBank. Archived from the original on 17 June 2020. Retrieved 2 June 2020.
- ^ Statement on the second meeting of the International Health Regulations (2005) Emergency Committee regarding the outbreak of novel coronavirus (2019-nCoV). World Health Organization (WHO). 30 January 2020. Archived from the original on 31 January 2020. Retrieved 30 January 2020.
- ^ WHO Director-General's opening remarks at the media briefing on COVID-19 – 11 March 2020. World Health Organization (WHO). 11 March 2020. Archived from the original on 11 March 2020. Retrieved 12 March 2020.
- ^ Rigby J, Satija B (5 May 2023). "WHO declares end to COVID global health emergency". Reuters. Retrieved 6 May 2023.
- ^ Machhi J, Herskovitz J, Senan AM, Dutta D, Nath B, Oleynikov MD, Blomberg WR, Meigs DD, Hasan M, Patel M, Kline P, Chang RC, Chang L, Gendelman HE, Kevadiya BD (September 2020). "The Natural History, Pathobiology, and Clinical Manifestations of SARS-CoV-2 Infections". Journal of Neuroimmune Pharmacology. 15 (3): 359–386. doi:10.1007/s11481-020-09944-5. PMC 7373339. PMID 32696264.
- ^ a b Chan JF, Yuan S, Kok KH, To KK, Chu H, Yang J, Xing F, Liu J, Yip CC, Poon RW, Tsoi HW, Lo SK, Chan KH, Poon VK, Chan WM, Ip JD, Cai JP, Cheng VC, Chen H, Hui CK, Yuen KY (February 2020). "A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster". Lancet. 395 (10223): 514–523. doi:10.1016/S0140-6736(20)30154-9. PMC 7159286. PMID 31986261.
- ^ "New coronavirus stable for hours on surfaces". National Institutes of Health (NIH). NIH.gov. 17 March 2020. Archived from the original on 23 March 2020. Retrieved 4 May 2020.
- ^ a b V'kovski P, Kratzel A, Steiner S, Stalder H, Thiel V (March 2021). "Coronavirus biology and replication: implications for SARS-CoV-2". Nat Rev Microbiol (Review). 19 (3): 155–170. doi:10.1038/s41579-020-00468-6. PMC 7592455. PMID 33116300.
- ^ Novel Coronavirus (2019-nCoV): situation report, 22 (Report). World Health Organization. 11 February 2020. hdl:10665/330991.
- ^ a b c Cohen J (January 2020). "Wuhan seafood market may not be source of novel virus spreading globally". Science. doi:10.1126/science.abb0611. S2CID 214574620.
- ^ a b Billah MA, Miah MM, Khan MN (11 November 2020). "Reproductive number of coronavirus: A systematic review and meta-analysis based on global level evidence". PLOS ONE. 15 (11): e0242128. Bibcode:2020PLoSO..1542128B. doi:10.1371/journal.pone.0242128. PMC 7657547. PMID 33175914.
- ^ "COVID-19 variants: What's the concern?". Mayo Clinic. 27 August 2022. Retrieved 10 October 2022.
- ^ "How Coronavirus Spreads Archived 3 April 2020 at the Wayback Machine", Centers for Disease Control and Prevention, Retrieved 14 May 2021.
- ^ "Coronavirus disease (COVID-19): How is it transmitted? Archived 15 October 2020 at the Wayback Machine", World Health Organization
- ^ a b Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S (April 2020). "SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor". Cell. 181 (2): 271–280.e8. doi:10.1016/j.cell.2020.02.052. PMC 7102627. PMID 32142651.
- ^ Zhao P, Praissman JL, Grant OC, Cai Y, Xiao T, Rosenbalm KE, Aoki K, Kellman BP, Bridger R, Barouch DH, Brindley MA, Lewis NE, Tiemeyer M, Chen B, Woods RJ, Wells L (October 2020). "Virus-Receptor Interactions of Glycosylated SARS-CoV-2 Spike and Human ACE2 Receptor". Cell Host & Microbe. 28 (4): 586–601.e6. doi:10.1016/j.chom.2020.08.004. PMC 7443692. PMID 32841605.
- ^ Huang P (22 January 2020). "How Does Wuhan Coronavirus Compare with MERS, SARS and the Common Cold?". NPR. Archived from the original on 2 February 2020. Retrieved 3 February 2020.
- ^ a b Fox D (January 2020). "What you need to know about the novel coronavirus". Nature. doi:10.1038/d41586-020-00209-y. PMID 33483684. S2CID 213064026.
- ^ World Health Organization (30 January 2020). Novel Coronavirus (2019-nCoV): situation report, 10 (Report). World Health Organization. hdl:10665/330775.
- ^ "World Health Organization Best Practices for the Naming of New Human Infectious Diseases" (PDF). WHO. May 2015. Archived (PDF) from the original on 12 February 2020.
- ^ "Novel coronavirus named 'Covid-19': WHO". TODAYonline. Archived from the original on 21 March 2020. Retrieved 11 February 2020.
- ^ "The coronavirus spreads racism against—and among—ethnic Chinese". The Economist. 17 February 2020. Archived from the original on 17 February 2020. Retrieved 17 February 2020.
- ^ "Naming the coronavirus disease (COVID-2019) and the virus that causes it". World Health Organization. Archived from the original on 28 February 2020. Retrieved 14 December 2020.
ICTV announced "severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)" as the name of the new virus on 11 February 2020. This name was chosen because the virus is genetically related to the coronavirus responsible for the SARS outbreak of 2003. While related, the two viruses are different.
- ^ Hui M (18 March 2020). "Why won't the WHO call the coronavirus by its name, SARS-CoV-2?". Quartz. Archived from the original on 25 March 2020. Retrieved 26 March 2020.
- ^ "Naming the coronavirus disease (COVID-2019) and the virus that causes it". World Health Organization. Archived from the original on 28 February 2020. Retrieved 14 December 2020.
From a risk communications perspective, using the name SARS can have unintended consequences in terms of creating unnecessary fear for some populations. ... For that reason and others, WHO has begun referring to the virus as "the virus responsible for COVID-19" or "the COVID-19 virus" when communicating with the public. Neither of these designations is [sic] intended as replacements for the official name of the virus as agreed by the ICTV.
- ^ "Standing Up to Hate and Bias Related to COVID-19". National Education Association. 5 June 2020.
It's racist and it creates xenophobia," University of California at Berkeley Asian American studies lecturer Harvey Dong told The Washington Post. "It's a very dangerous situation."
- ^ Gstalter M (19 March 2020). "WHO official warns against calling it 'Chinese virus,' says 'there is no blame in this'". The Hill. Retrieved 15 September 2022.
Ryan is not the first WHO official to push back against the phrase. Director-General Tedros Adhanom Ghebreyesus said earlier this month that the term is "painful to see" and "more dangerous than the virus itself."
- ^ Gover AR, Harper SB, Langton L (July 2020). "Anti-Asian Hate Crime During the COVID-19 Pandemic: Exploring the Reproduction of Inequality". American Journal of Criminal Justice. 45 (4): 647–667. doi:10.1007/s12103-020-09545-1. PMC 7364747. PMID 32837171.
- ^ Li JY, You Z, Wang Q, Zhou ZJ, Qiu Y, Luo R, Ge XY (March 2020). "The epidemic of 2019-novel-coronavirus (2019-nCoV) pneumonia and insights for emerging infectious diseases in the future". Microbes and Infection. 22 (2): 80–85. doi:10.1016/j.micinf.2020.02.002. PMC 7079563. PMID 32087334.
- ^ Kessler G (17 April 2020). "Trump's false claim that the WHO said the coronavirus was 'not communicable'". The Washington Post. Archived from the original on 17 April 2020. Retrieved 17 April 2020.
- ^ Kuo L (21 January 2020). "China confirms human-to-human transmission of coronavirus". The Guardian. Archived from the original on 22 March 2020. Retrieved 18 April 2020.
- ^ "How COVID-19 Spreads". U.S. Centers for Disease Control and Prevention (CDC). 27 January 2020. Archived from the original on 28 January 2020. Retrieved 29 January 2020.
- ^ Edwards E (25 January 2020). "How does coronavirus spread?". NBC News. Archived from the original on 28 January 2020. Retrieved 13 March 2020.
- ^ Anfinrud P, Stadnytskyi V, Bax CE, Bax A (May 2020). "Visualizing Speech-Generated Oral Fluid Droplets with Laser Light Scattering". The New England Journal of Medicine. 382 (21): 2061–2063. doi:10.1056/NEJMc2007800. PMC 7179962. PMID 32294341.
- ^ Stadnytskyi V, Bax CE, Bax A, Anfinrud P (June 2020). "The airborne lifetime of small speech droplets and their potential importance in SARS-CoV-2 transmission". Proceedings of the National Academy of Sciences of the United States of America. 117 (22): 11875–11877. Bibcode:2020PNAS..11711875S. doi:10.1073/pnas.2006874117. PMC 7275719. PMID 32404416.
- ^ Klompas M, Baker MA, Rhee C (August 2020). "Airborne Transmission of SARS-CoV-2: Theoretical Considerations and Available Evidence". JAMA. 324 (5): 441–442. doi:10.1001/jama.2020.12458. PMID 32749495. S2CID 220500293.
Investigators have demonstrated that speaking and coughing produce a mixture of both droplets and aerosols in a range of sizes, that these secretions can travel together for up to 27 feet, that it is feasible for SARS-CoV-2 to remain suspended in the air and viable for hours, that SARS-CoV-2 RNA can be recovered from air samples in hospitals, and that poor ventilation prolongs the amount of time that aerosols remain airborne.
- ^ Rettner R (21 January 2021). "Talking is worse than coughing for spreading COVID-19 indoors". Live Science. Retrieved 10 October 2022.
In one modeled scenario, the researchers found that after a short cough, the number of infectious particles in the air would quickly fall after 1 to 7 minutes; in contrast, after speaking for 30 seconds, only after 30 minutes would the number of infectious particles fall to similar levels; and a high number of particles were still suspended after one hour. In other words, a dose of virus particles capable of causing an infection would linger in the air much longer after speech than a cough. (In this modeled scenario, the same number of droplets were admitted during a 0.5-second cough as during the course of 30 seconds of speech.)
- ^ de Oliveira PM, Mesquita LC, Gkantonas S, Giusti A, Mastorakos E (January 2021). "Evolution of spray and aerosol from respiratory releases: theoretical estimates for insight on viral transmission". Proceedings of the Royal Society A: Mathematical, Physical and Engineering Sciences. 477 (2245): 20200584. Bibcode:2021RSPSA.47700584D. doi:10.1098/rspa.2020.0584. PMC 7897643. PMID 33633490. S2CID 231643585.
- ^ Mandavilli A (4 July 2020). "239 Experts With One Big Claim: The Coronavirus Is Airborne – The W.H.O. has resisted mounting evidence that viral particles floating indoors are infectious, some scientists say. The agency maintains the research is still inconclusive". The New York Times. Archived from the original on 17 November 2020. Retrieved 5 July 2020.
- ^ Tufekci Z (30 July 2020). "We Need to Talk About Ventilation". The Atlantic. Archived from the original on 17 November 2020. Retrieved 8 September 2020.
- ^ Lewis D (July 2020). "Mounting evidence suggests coronavirus is airborne - but health advice has not caught up". Nature. 583 (7817): 510–513. Bibcode:2020Natur.583..510L. doi:10.1038/d41586-020-02058-1. PMID 32647382. S2CID 220470431.
- ^ Popa A, Genger JW, Nicholson MD, Penz T, Schmid D, Aberle SW, Agerer B, Lercher A, Endler L, Colaço H, Smyth M, Schuster M, Grau ML, Martínez-Jiménez F, Pich O, Borena W, Pawelka E, Keszei Z, Senekowitsch M, Laine J, Aberle JH, Redlberger-Fritz M, Karolyi M, Zoufaly A, Maritschnik S, Borkovec M, Hufnagl P, Nairz M, Weiss G, Wolfinger MT, von Laer D, Superti-Furga G, Lopez-Bigas N, Puchhammer-Stöckl E, Allerberger F, Michor F, Bock C, Bergthaler A (December 2020). "Genomic epidemiology of superspreading events in Austria reveals mutational dynamics and transmission properties of SARS-CoV-2". Science Translational Medicine. 12 (573): eabe2555. doi:10.1126/scitranslmed.abe2555. PMC 7857414. PMID 33229462.
- ^ a b He X, Lau EH, Wu P, Deng X, Wang J, Hao X, Lau YC, Wong JY, Guan Y, Tan X, Mo X, Chen Y, Liao B, Chen W, Hu F, Zhang Q, Zhong M, Wu Y, Zhao L, Zhang F, Cowling BJ, Li F, Leung GM (May 2020). "Temporal dynamics in viral shedding and transmissibility of COVID-19". Nature Medicine. 26 (5): 672–675. doi:10.1038/s41591-020-0869-5. PMID 32296168.
- ^ Watanabe T, Bartrand TA, Weir MH, Omura T, Haas CN (July 2010). "Development of a dose-response model for SARS coronavirus". Risk Analysis. 30 (7): 1129–38. Bibcode:2010RiskA..30.1129W. doi:10.1111/j.1539-6924.2010.01427.x. PMC 7169223. PMID 20497390.
- ^ Artika IM, Ma'roef CN (May 2017). "Laboratory biosafety for handling emerging viruses". Asian Pacific Journal of Tropical Biomedicine. 7 (5): 483–491. doi:10.1016/j.apjtb.2017.01.020. PMC 7103938. PMID 32289025.
- ^ "Getting your workplace ready for COVID-19" (PDF). World Health Organization. 27 February 2020. Archived (PDF) from the original on 2 March 2020. Retrieved 3 March 2020.
- ^ Yong E (20 March 2020). "Why the Coronavirus Has Been So Successful". The Atlantic. Archived from the original on 20 March 2020. Retrieved 20 March 2020.
- ^ Gibbens S (18 March 2020). "Why soap is preferable to bleach in the fight against coronavirus". National Geographic. Archived from the original on 2 April 2020. Retrieved 2 April 2020.
- ^ Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, Spitters C, Ericson K, Wilkerson S, Tural A, Diaz G, Cohn A, Fox L, Patel A, Gerber SI, Kim L, Tong S, Lu X, Lindstrom S, Pallansch MA, Weldon WC, Biggs HM, Uyeki TM, Pillai SK (March 2020). "First Case of 2019 Novel Coronavirus in the United States". The New England Journal of Medicine. 382 (10): 929–936. doi:10.1056/NEJMoa2001191. PMC 7092802. PMID 32004427.
- ^ Li D, Jin M, Bao P, Zhao W, Zhang S (May 2020). "Clinical Characteristics and Results of Semen Tests Among Men With Coronavirus Disease 2019". JAMA Network Open. 3 (5): e208292. doi:10.1001/jamanetworkopen.2020.8292. PMC 7206502. PMID 32379329.
- ^ Wölfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Müller MA, Niemeyer D, Jones TC, Vollmar P, Rothe C, Hoelscher M, Bleicker T, Brünink S, Schneider J, Ehmann R, Zwirglmaier K, Drosten C, Wendtner C (May 2020). "Virological assessment of hospitalized patients with COVID-2019". Nature. 581 (7809): 465–469. Bibcode:2020Natur.581..465W. doi:10.1038/s41586-020-2196-x. PMID 32235945.
- ^ Kupferschmidt K (February 2020). "Study claiming new coronavirus can be transmitted by people without symptoms was flawed". Science. doi:10.1126/science.abb1524. S2CID 214094598.
- ^ To KK, Tsang OT, Leung WS, Tam AR, Wu TC, Lung DC, Yip CC, Cai JP, Chan JM, Chik TS, Lau DP, Choi CY, Chen LL, Chan WM, Chan KH, Ip JD, Ng AC, Poon RW, Luo CT, Cheng VC, Chan JF, Hung IF, Chen Z, Chen H, Yuen KY (May 2020). "Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study". The Lancet. Infectious Diseases. 20 (5): 565–574. doi:10.1016/S1473-3099(20)30196-1. PMC 7158907. PMID 32213337.
- ^ Avanzato VA, Matson MJ, Seifert SN, Pryce R, Williamson BN, Anzick SL, Barbian K, Judson SD, Fischer ER, Martens C, Bowden TA, de Wit E, Riedo FX, Munster VJ (December 2020). "Case Study: Prolonged Infectious SARS-CoV-2 Shedding from an Asymptomatic Immunocompromised Individual with Cancer". Cell. 183 (7): 1901–1912.e9. doi:10.1016/j.cell.2020.10.049. PMC 7640888. PMID 33248470.
- ^ a b Hou YJ, Okuda K, Edwards CE, Martinez DR, Asakura T, Dinnon KH, Kato T, Lee RE, Yount BL, Mascenik TM, Chen G, Olivier KN, Ghio A, Tse LV, Leist SR, Gralinski LE, Schäfer A, Dang H, Gilmore R, Nakano S, Sun L, Fulcher ML, Livraghi-Butrico A, Nicely NI, Cameron M, Cameron C, Kelvin DJ, de Silva A, Margolis DM, Markmann A, Bartelt L, Zumwalt R, Martinez FJ, Salvatore SP, Borczuk A, Tata PR, Sontake V, Kimple A, Jaspers I, O'Neal WK, Randell SH, Boucher RC, Baric RS (July 2020). "SARS-CoV-2 Reverse Genetics Reveals a Variable Infection Gradient in the Respiratory Tract". Cell. 182 (2): 429–446.e14. doi:10.1016/j.cell.2020.05.042. PMC 7250779. PMID 32526206.
- ^ Banerjee A, Mossman K, Baker ML (February 2021). "Zooanthroponotic potential of SARS-CoV-2 and implications of reintroduction into human populations". Cell Host & Microbe. 29 (2): 160–164. doi:10.1016/j.chom.2021.01.004. PMC 7837285. PMID 33539765.
- ^ "Questions and Answers on the COVID-19: OIE – World Organisation for Animal Health". www.oie.int. Archived from the original on 31 March 2020. Retrieved 16 April 2020.
- ^ Goldstein J (6 April 2020). "Bronx Zoo Tiger Is Sick with the Coronavirus". The New York Times. Archived from the original on 9 April 2020. Retrieved 10 April 2020.
- ^ "USDA Statement on the Confirmation of COVID-19 in a Tiger in New York". United States Department of Agriculture. 5 April 2020. Archived from the original on 15 April 2020. Retrieved 16 April 2020.
- ^ "If You Have Animals—Coronavirus Disease 2019 (COVID-19)". Centers for Disease Control and Prevention (CDC). 13 April 2020. Archived from the original on 1 April 2020. Retrieved 16 April 2020.
- ^ World Health Organization (1 February 2020). Novel Coronavirus (2019-nCoV): situation report, 12 (Report). World Health Organization. hdl:10665/330777.
- ^ Nogrady B (November 2020). "What the data say about asymptomatic COVID infections". Nature. 587 (7835): 534–535. Bibcode:2020Natur.587..534N. doi:10.1038/d41586-020-03141-3. PMID 33214725.
- ^ Li R, Pei S, Chen B, Song Y, Zhang T, Yang W, Shaman J (May 2020). "Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2)". Science. 368 (6490): 489–493. Bibcode:2020Sci...368..489L. doi:10.1126/science.abb3221. PMC 7164387. PMID 32179701.
- ^ Daily Telegraph, Thursday 28 May 2020, page 2 column 1, which refers to the medical journal Thorax; Thorax May 2020 article COVID-19: in the footsteps of Ernest Shackleton Archived 30 May 2020 at the Wayback Machine
- ^ He X, Lau EH, Wu P, Deng X, Wang J, Hao X, Lau YC, Wong JY, Guan Y, Tan X, Mo X, Chen Y, Liao B, Chen W, Hu F, Zhang Q, Zhong M, Wu Y, Zhao L, Zhang F, Cowling BJ, Li F, Leung GM (September 2020). "Author Correction: Temporal dynamics in viral shedding and transmissibility of COVID-19". Nature Medicine. 26 (9): 1491–1493. doi:10.1038/s41591-020-1016-z. PMC 7413015. PMID 32770170.
- ^ a b Ledford H (September 2020). "Coronavirus reinfections: three questions scientists are asking". Nature. 585 (7824): 168–169. doi:10.1038/d41586-020-02506-y. PMID 32887957. S2CID 221501940.
- ^ a b To KK, Hung IF, Ip JD, Chu AW, Chan WM, Tam AR, Fong CH, Yuan S, Tsoi HW, Ng AC, Lee LL, Wan P, Tso E, To WK, Tsang D, Chan KH, Huang JD, Kok KH, Cheng VC, Yuen KY (August 2020). "COVID-19 re-infection by a phylogenetically distinct SARS-coronavirus-2 strain confirmed by whole genome sequencing". Clinical Infectious Diseases. 73 (9): e2946–e2951. doi:10.1093/cid/ciaa1275. PMC 7499500. PMID 32840608. S2CID 221308584.
- ^ a b Tillett RL, Sevinsky JR, Hartley PD, Kerwin H, Crawford N, Gorzalski A, Laverdure C, Verma SC, Rossetto CC, Jackson D, Farrell MJ, Van Hooser S, Pandori M (January 2021). "Genomic evidence for reinfection with SARS-CoV-2: a case study". The Lancet. Infectious Diseases. 21 (1): 52–58. doi:10.1016/S1473-3099(20)30764-7. PMC 7550103. PMID 33058797.
- ^ a b Holmes EC, Goldstein SA, Rasmussen AL, Robertson DL, Crits-Christoph A, Wertheim JO, Anthony SJ, Barclay WS, Boni MF, Doherty PC, Farrar J (August 2021). "The Origins of SARS-CoV-2: A Critical Review". Cell. 184 (19): 4848–4856. doi:10.1016/j.cell.2021.08.017. PMC 8373617. PMID 34480864.
- ^ a b c Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL (March 2020). "A pneumonia outbreak associated with a new coronavirus of probable bat origin". Nature. 579 (7798): 270–273. Bibcode:2020Natur.579..270Z. doi:10.1038/s41586-020-2012-7. PMC 7095418. PMID 32015507.
- ^ Eschner K (28 January 2020). "We're still not sure where the Wuhan coronavirus really came from". Popular Science. Archived from the original on 30 January 2020. Retrieved 30 January 2020.
- ^ Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B (February 2020). "Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China". Lancet. 395 (10223): 497–506. doi:10.1016/S0140-6736(20)30183-5. PMC 7159299. PMID 31986264.
- ^ a b Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L (February 2020). "Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study". Lancet. 395 (10223): 507–513. doi:10.1016/S0140-6736(20)30211-7. PMC 7135076. PMID 32007143.
- ^ a b Cyranoski D (March 2020). "Mystery deepens over animal source of coronavirus". Nature. 579 (7797): 18–19. Bibcode:2020Natur.579...18C. doi:10.1038/d41586-020-00548-w. PMID 32127703.
- ^ Yu WB, Tang GD, Zhang L, Corlett RT (May 2020). "Decoding the evolution and transmissions of the novel pneumonia coronavirus (SARS-CoV-2 / HCoV-19) using whole genomic data". Zoological Research. 41 (3): 247–257. doi:10.24272/j.issn.2095-8137.2020.022. PMC 7231477. PMID 32351056.
- ^ a b c d e Joint WHO-China Study Team (30 March 2021). WHO-convened global study of origins of SARS-CoV-2: China Part. Geneva, Switzerland: World Health Organization. Retrieved 31 May 2023.
- ^ Worobey M (December 2021). "Dissecting the early COVID-19 cases in Wuhan". Science. 374 (6572): 1202–1204. Bibcode:2021Sci...374.1202W. doi:10.1126/science.abm4454. PMID 34793199. S2CID 244403410.
- ^ Kang L, He G, Sharp AK, Wang X, Brown AM, Michalak P, Weger-Lucarelli J (August 2021). "A selective sweep in the Spike gene has driven SARS-CoV-2 human adaptation". Cell. 184 (17): 4392–4400.e4. doi:10.1016/j.cell.2021.07.007. PMC 8260498. PMID 34289344.
- ^ Decaro N, Lorusso A (May 2020). "Novel human coronavirus (SARS-CoV-2): A lesson from animal coronaviruses". Veterinary Microbiology. 244: 108693. doi:10.1016/j.vetmic.2020.108693. PMC 7195271. PMID 32402329.
- ^ Robson F, Khan KS, Le TK, Paris C, Demirbag S, Barfuss P, Rocchi P, Ng WL (August 2020). "Coronavirus RNA Proofreading: Molecular Basis and Therapeutic Targeting [published correction appears in Mol Cell. 2020 Dec 17;80(6):1136–1138]". Molecular Cell. 79 (5): 710–727. doi:10.1016/j.molcel.2020.07.027. PMC 7402271. PMID 32853546.
- ^ Tao K, Tzou PL, Nouhin J, Gupta RK, de Oliveira T, Kosakovsky Pond SL, Fera D, Shafer RW (December 2021). "The biological and clinical significance of emerging SARS-CoV-2 variants". Nature Reviews Genetics. 22 (12): 757–773. doi:10.1038/s41576-021-00408-x. PMC 8447121. PMID 34535792.
- ^ Temmam S, Vongphayloth K, Salazar EB, Munier S, Bonomi M, Régnault B, Douangboubpha B, Karami Y, Chretien D, Sanamxay D, Xayaphet V (February 2022). "Bat coronaviruses related to SARS-CoV-2 and infectious for human cells". Nature. 604 (7905): 330–336. Bibcode:2022Natur.604..330T. doi:10.1038/s41586-022-04532-4. PMID 35172323. S2CID 246902858.
- ^ Mallapaty S (24 September 2021). "Closest known relatives of virus behind COVID-19 found in Laos". Nature. 597 (7878): 603. Bibcode:2021Natur.597..603M. doi:10.1038/d41586-021-02596-2. PMID 34561634. S2CID 237626322.
- ^ "Newly Discovered Bat Viruses Give Hints to Covid's Origins". The New York Times. 14 October 2021.
- ^ "Bat coronavirus isolate RaTG13, complete genome". National Center for Biotechnology Information (NCBI). 10 February 2020. Archived from the original on 15 May 2020. Retrieved 5 March 2020.
- ^ "The 'Occam's Razor Argument' Has Not Shifted in Favor of a Lab Leak". Snopes.com. Snopes. 16 July 2021. Retrieved 18 July 2021.
- ^ Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W (February 2020). "Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding". Lancet. 395 (10224): 565–574. doi:10.1016/S0140-6736(20)30251-8. PMC 7159086. PMID 32007145.
- ^ O'Keeffe J, Freeman S, Nicol A (21 March 2021). The Basics of SARS-CoV-2 Transmission. Vancouver, BC: National Collaborating Centre for Environmental Health (NCCEH). ISBN 978-1-988234-54-0. Archived from the original on 12 May 2021. Retrieved 12 May 2021.
- ^ Xiao K, Zhai J, Feng Y, Zhou N, Zhang X, Zou JJ, Li N, Guo Y, Li X, Shen X, Zhang Z, Shu F, Huang W, Li Y, Zhang Z, Chen RA, Wu YJ, Peng SM, Huang M, Xie WJ, Cai QH, Hou FH, Chen W, Xiao L, Shen Y (July 2020). "Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins". Nature. 583 (7815): 286–289. Bibcode:2020Natur.583..286X. doi:10.1038/s41586-020-2313-x. PMID 32380510. S2CID 218557880.
- ^ Zhao J, Cui W, Tian BP (2020). "The Potential Intermediate Hosts for SARS-CoV-2". Frontiers in Microbiology. 11: 580137. doi:10.3389/fmicb.2020.580137. PMC 7554366. PMID 33101254.
- ^ "Why it's so tricky to trace the origin of COVID-19". Science. National Geographic. 10 September 2021.
- ^ a b c Hu B, Guo H, Zhou P, Shi ZL (March 2021). "Characteristics of SARS-CoV-2 and COVID-19". Nature Reviews. Microbiology. 19 (3): 141–154. doi:10.1038/s41579-020-00459-7. PMC 7537588. PMID 33024307.
- ^ Giovanetti M, Benedetti F, Campisi G, Ciccozzi A, Fabris S, Ceccarelli G, Tambone V, Caruso A, Angeletti S, Zella D, Ciccozzi M (January 2021). "Evolution patterns of SARS-CoV-2: Snapshot on its genome variants". Biochemical and Biophysical Research Communications. 538: 88–91. doi:10.1016/j.bbrc.2020.10.102. PMC 7836704. PMID 33199021. S2CID 226988090.
- ^ a b c V'kovski P, Kratzel A, Steiner S, Stalder H, Thiel V (March 2021). "Coronavirus biology and replication: implications for SARS-CoV-2". Nature Reviews. Microbiology. 19 (3): 155–170. doi:10.1038/s41579-020-00468-6. PMC 7592455. PMID 33116300.
- ^ a b Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W (February 2020). "A Novel Coronavirus from Patients with Pneumonia in China, 2019". The New England Journal of Medicine. 382 (8): 727–733. doi:10.1056/NEJMoa2001017. PMC 7092803. PMID 31978945.
- ^ "Phylogeny of SARS-like betacoronaviruses". nextstrain. Archived from the original on 20 January 2020. Retrieved 18 January 2020.
- ^ Wong AC, Li X, Lau SK, Woo PC (February 2019). "Global Epidemiology of Bat Coronaviruses". Viruses. 11 (2): 174. doi:10.3390/v11020174. PMC 6409556. PMID 30791586.
- ^ a b Singh D, Yi SV (April 2021). "On the origin and evolution of SARS-CoV-2". Experimental & Molecular Medicine. 53 (4): 537–547. doi:10.1038/s12276-021-00604-z. PMC 8050477. PMID 33864026.
- ^ Jackson B, Boni MF, Bull MJ, Colleran A, Colquhoun RM, Darby AC, Haldenby S, Hill V, Lucaci A, McCrone JT, Nicholls SM, O'Toole Á, Pacchiarini N, Poplawski R, Scher E, Todd F, Webster HJ, Whitehead M, Wierzbicki C, Loman NJ, Connor TR, Robertson DL, Pybus OG, Rambaut A (September 2021). "Generation and transmission of interlineage recombinants in the SARS-CoV-2 pandemic". Cell. 184 (20): 5179–5188.e8. doi:10.1016/j.cell.2021.08.014. PMC 8367733. PMID 34499854. S2CID 237099659.
- ^ a b "CoV2020". GISAID EpifluDB. Archived from the original on 12 January 2020. Retrieved 12 January 2020.
- ^ Kim D, Lee JY, Yang JS, Kim JW, Kim VN, Chang H (May 2020). "The Architecture of SARS-CoV-2 Transcriptome". Cell. 181 (4): 914–921.e10. doi:10.1016/j.cell.2020.04.011. PMC 7179501. PMID 32330414.
- ^ Hossain MG, Tang Yd, Akter S, Zheng C (May 2022). "Roles of the polybasic furin cleavage site of spike protein in SARS-CoV-2 replication, pathogenesis, and host immune responses and vaccination". Journal of Medical Virology. 94 (5): 1815–1820. doi:10.1002/jmv.27539. PMID 34936124. S2CID 245430230.
- ^ To KK, Sridhar S, Chiu KH, Hung DL, Li X, Hung IF, Tam AR, Chung TW, Chan JF, Zhang AJ, Cheng VC, Yuen KY (December 2021). "Lessons learned 1 year after SARS-CoV-2 emergence leading to COVID-19 pandemic". Emerging Microbes & Infections. 10 (1): 507–535. doi:10.1080/22221751.2021.1898291. PMC 8006950. PMID 33666147.
- ^ a b Jackson CB, Farzan M, Chen B, Choe H (January 2022). "Mechanisms of SARS-CoV-2 entry into cells". Nature Reviews Molecular Cell Biology. 23 (1): 3–20. doi:10.1038/s41580-021-00418-x. PMC 8491763. PMID 34611326.
- ^ Braun E, Sauter D (2019). "Furin-mediated protein processing in infectious diseases and cancer". Clinical & Translational Immunology. 8 (8): e1073. doi:10.1002/cti2.1073. PMC 6682551. PMID 31406574.
- ^ Vankadari N (August 2020). "Structure of Furin Protease Binding to SARS-CoV-2 Spike Glycoprotein and Implications for Potential Targets and Virulence". The Journal of Physical Chemistry Letters. 11 (16): 6655–6663. doi:10.1021/acs.jpclett.0c01698. PMC 7409919. PMID 32787225.
- ^ a b Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E (April 2020). "The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade". Antiviral Research. 176 (7): 104742. Bibcode:2020CBio...30E1346Z. doi:10.1016/j.cub.2020.03.022. PMC 7114094. PMID 32057769.
- ^ Zhang T, Wu Q, Zhang Z (April 2020). "Probable Pangolin Origin of SARS-CoV-2 Associated with the COVID-19 Outbreak". Current Biology. 30 (7): 1346–1351.e2. Bibcode:2020CBio...30E1346Z. doi:10.1016/j.cub.2020.03.022. PMC 7156161. PMID 32197085.
- ^ Wu Y, Zhao S (December 2020). "Furin cleavage sites naturally occur in coronaviruses". Stem Cell Research. 50: 102115. doi:10.1016/j.scr.2020.102115. PMC 7836551. PMID 33340798.
- ^ Worobey M, Pekar J, Larsen BB, Nelson MI, Hill V, Joy JB, Rambaut A, Suchard MA, Wertheim JO, Lemey P (October 2020). "The emergence of SARS-CoV-2 in Europe and North America". Science. 370 (6516): 564–570. doi:10.1126/science.abc8169. PMC 7810038. PMID 32912998.
- ^ "Initial genome release of novel coronavirus". Virological. 11 January 2020. Archived from the original on 12 January 2020. Retrieved 12 January 2020.
- ^ a b Bedford T, Neher R, Hadfield N, Hodcroft E, Ilcisin M, Müller N. "Genomic analysis of nCoV spread: Situation report 2020-01-30". nextstrain.org. Archived from the original on 15 March 2020. Retrieved 18 March 2020.
- ^ Sun J, He WT, Wang L, Lai A, Ji X, Zhai X, Li G, Suchard MA, Tian J, Zhou J, Veit M, Su S (May 2020). "COVID-19: Epidemiology, Evolution, and Cross-Disciplinary Perspectives". Trends in Molecular Medicine. 26 (5): 483–495. doi:10.1016/j.molmed.2020.02.008. PMC 7118693. PMID 32359479.
- ^ "Genomic epidemiology of novel coronavirus – Global subsampling". Nextstrain. 25 October 2021. Archived from the original on 20 April 2020. Retrieved 26 October 2021.
- ^ Coronaviridae Study Group of the International Committee on Taxonomy of Viruses (April 2020). "The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2". Nature Microbiology. 5 (4): 536–544. doi:10.1038/s41564-020-0695-z. PMC 7095448. PMID 32123347.
- ^ "New, more infectious strain of COVID-19 now dominates global cases of virus: study". medicalxpress.com. Archived from the original on 17 November 2020. Retrieved 16 August 2020.
- ^ Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, Hengartner N, Giorgi EE, Bhattacharya T, Foley B, Hastie KM, Parker MD, Partridge DG, Evans CM, Freeman TM, de Silva TI, McDanal C, Perez LG, Tang H, Moon-Walker A, Whelan SP, LaBranche CC, Saphire EO, Montefiori DC (August 2020). "Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus". Cell. 182 (4): 812–827.e19. doi:10.1016/j.cell.2020.06.043. PMC 7332439. PMID 32697968.
- ^ Dhama K, Khan S, Tiwari R, Sircar S, Bhat S, Malik YS, Singh KP, Chaicumpa W, Bonilla-Aldana DK, Rodriguez-Morales AJ (September 2020). "Coronavirus Disease 2019-COVID-19". Clinical Microbiology Reviews. 33 (4). doi:10.1128/CMR.00028-20. PMC 7405836. PMID 32580969.
- ^ Dockrill P (11 November 2020). "Scientists Just Found a Mysteriously Hidden 'Gene Within a Gene' in SARS-CoV-2". ScienceAlert. Archived from the original on 17 November 2020. Retrieved 11 November 2020.
- ^ Nelson CW, Ardern Z, Goldberg TL, Meng C, Kuo CH, Ludwig C, Kolokotronis SO, Wei X (October 2020). "Dynamically evolving novel overlapping gene as a factor in the SARS-CoV-2 pandemic". eLife. 9. doi:10.7554/eLife.59633. PMC 7655111. PMID 33001029.
- ^ a b Zhou H, Ji J, Chen X, Bi Y, Li J, Wang Q, et al. (August 2021). "Identification of novel bat coronaviruses sheds light on the evolutionary origins of SARS-CoV-2 and related viruses". Cell. 184 (17): 4380–4391.e14. doi:10.1016/j.cell.2021.06.008. PMC 8188299. PMID 34147139.
- ^ a b Wacharapluesadee S, Tan CW, Maneeorn P, Duengkae P, Zhu F, Joyjinda Y, et al. (February 2021). "Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia". Nature Communications. 12 (1): 972. Bibcode:2021NatCo..12..972W. doi:10.1038/s41467-021-21240-1. PMC 7873279. PMID 33563978.
- ^ Murakami S, Kitamura T, Suzuki J, Sato R, Aoi T, Fujii M, et al. (December 2020). "Detection and Characterization of Bat Sarbecovirus Phylogenetically Related to SARS-CoV-2, Japan". Emerging Infectious Diseases. 26 (12): 3025–3029. doi:10.3201/eid2612.203386. PMC 7706965. PMID 33219796.
- ^ a b Zhou H, Chen X, Hu T, Li J, Song H, Liu Y, et al. (June 2020). "A Novel Bat Coronavirus Closely Related to SARS-CoV-2 Contains Natural Insertions at the S1/S2 Cleavage Site of the Spike Protein". Current Biology. 30 (11): 2196–2203.e3. doi:10.1016/j.cub.2020.05.023. PMC 7211627. PMID 32416074.
- ^ Lam TT, Jia N, Zhang YW, Shum MH, Jiang JF, Zhu HC, et al. (July 2020). "Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins". Nature. 583 (7815): 282–285. Bibcode:2020Natur.583..282L. doi:10.1038/s41586-020-2169-0. PMID 32218527. S2CID 214683303.
- ^ Xiao K, Zhai J, Feng Y, Zhou N, Zhang X, Zou JJ, et al. (July 2020). "Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins". Nature. 583 (7815): 286–289. Bibcode:2020Natur.583..286X. doi:10.1038/s41586-020-2313-x. PMID 32380510. S2CID 256822274.
- ^ a b Delaune D, Hul V, Karlsson EA, Hassanin A, Ou TP, Baidaliuk A, et al. (November 2021). "A novel SARS-CoV-2 related coronavirus in bats from Cambodia". Nature Communications. 12 (1): 6563. Bibcode:2021NatCo..12.6563D. doi:10.1038/s41467-021-26809-4. PMC 8578604. PMID 34753934.
- ^ Zhou H, Chen X, Hu T, Li J, Song H, Liu Y, et al. (June 2020). "A Novel Bat Coronavirus Closely Related to SARS-CoV-2 Contains Natural Insertions at the S1/S2 Cleavage Site of the Spike Protein". Current Biology. 30 (11): 2196–2203.e3. doi:10.1016/j.cub.2020.05.023. PMC 7211627. PMID 32416074.
- ^ Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, et al. (March 2020). "A pneumonia outbreak associated with a new coronavirus of probable bat origin". Nature. 579 (7798): 270–273. Bibcode:2020Natur.579..270Z. doi:10.1038/s41586-020-2012-7. PMC 7095418. PMID 32015507.
- ^ Temmam S, Vongphayloth K, Baquero E, Munier S, Bonomi M, Regnault B, et al. (April 2022). "Bat coronaviruses related to SARS-CoV-2 and infectious for human cells". Nature. 604 (7905): 330–336. Bibcode:2022Natur.604..330T. doi:10.1038/s41586-022-04532-4. PMID 35172323. S2CID 246902858.
- ^ Koyama T, Platt D, Parida L (July 2020). "Variant analysis of SARS-CoV-2 genomes". Bulletin of the World Health Organization. 98 (7): 495–504. doi:10.2471/BLT.20.253591. PMC 7375210. PMID 32742035.
We detected in total 65776 variants with 5775 distinct variants.
- ^ Alm E, Broberg EK, Connor T, Hodcroft EB, Komissarov AB, Maurer-Stroh S, Melidou A, Neher RA, O'Toole Á, Pereyaslov D (August 2020). "Geographical and temporal distribution of SARS-CoV-2 clades in the WHO European Region, January to June 2020". Euro Surveillance. 25 (32). doi:10.2807/1560-7917.ES.2020.25.32.2001410. PMC 7427299. PMID 32794443.
- ^ World Health Organization (27 November 2021). "Tracking SARS-CoV-2 variants". World Health Organization. Archived from the original on 6 June 2021. Retrieved 28 November 2021.
- ^ "SARS-CoV-2 mink-associated variant strain – Denmark". WHO. 3 December 2020. Archived from the original on 31 December 2020. Retrieved 30 December 2020.
- ^ Sender R, Bar-On YM, Gleizer S, Bernsthein B, Flamholz A, Phillips R, Milo R (April 2021). "The total number and mass of SARS-CoV-2 virions". MedRxiv: The Preprint Server for Health Sciences. doi:10.1101/2020.11.16.20232009. PMC 7685332. PMID 33236021.
- ^ a b c Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y, Wang Q, Xu Y, Li M, Li X, Zheng M, Chen L, Li H (May 2020). "Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods". Acta Pharmaceutica Sinica B. 10 (5): 766–788. doi:10.1016/j.apsb.2020.02.008. PMC 7102550. PMID 32292689.
- ^ a b Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS (March 2020). "Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation". Science. 367 (6483): 1260–1263. Bibcode:2020Sci...367.1260W. doi:10.1126/science.abb2507. PMC 7164637. PMID 32075877.
- ^ Mandelbaum RF (19 February 2020). "Scientists Create Atomic-Level Image of the New Coronavirus's Potential Achilles Heel". Gizmodo. Archived from the original on 8 March 2020. Retrieved 13 March 2020.
- ^ Sokhansanj BA, Rosen GL (26 April 2022). Gaglia MM (ed.). "Mapping Data to Deep Understanding: Making the Most of the Deluge of SARS-CoV-2 Genome Sequences". mSystems. 7 (2): e00035–22. doi:10.1128/msystems.00035-22. ISSN 2379-5077. PMC 9040592. PMID 35311562.
- ^ "GISAID - gisaid.org". gisaid.org. Retrieved 16 September 2023.
- ^ Kandeel M, Ibrahim A, Fayez M, Al-Nazawi M (June 2020). "From SARS and MERS CoVs to SARS-CoV-2: Moving toward more biased codon usage in viral structural and nonstructural genes". Journal of Medical Virology. 92 (6): 660–666. doi:10.1002/jmv.25754. PMC 7228358. PMID 32159237.
- ^ a b Hou W (September 2020). "Characterization of codon usage pattern in SARS-CoV-2". Virology Journal. 17 (1): 138. doi:10.1186/s12985-020-01395-x. PMC 7487440. PMID 32928234.
- ^ a b Wang Y, Mao JM, Wang GD, Luo ZP, Yang L, Yao Q, Chen KP (July 2020). "Human SARS-CoV-2 has evolved to reduce CG dinucleotide in its open reading frames". Scientific Reports. 10 (1): 12331. Bibcode:2020NatSR..1012331W. doi:10.1038/s41598-020-69342-y. PMC 7378049. PMID 32704018.
- ^ Rice AM, Castillo Morales A, Ho AT, Mordstein C, Mühlhausen S, Watson S, Cano L, Young B, Kudla G, Hurst LD (January 2021). "Evidence for Strong Mutation Bias toward, and Selection against, U Content in SARS-CoV-2: Implications for Vaccine Design". Molecular Biology and Evolution. 38 (1): 67–83. doi:10.1093/molbev/msaa188. PMC 7454790. PMID 32687176.
- ^ Gu H, Chu DK, Peiris M, Poon LL (January 2020). "Multivariate analyses of codon usage of SARS-CoV-2 and other betacoronaviruses". Virus Evolution. 6 (1): veaa032. doi:10.1093/ve/veaa032. PMC 7223271. PMID 32431949.
- ^ Wang Q, Zhang Y, Wu L, Niu S, Song C, Zhang Z, Lu G, Qiao C, Hu Y, Yuen KY, Wang Q, Zhou H, Yan J, Qi J (May 2020). "Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2". Cell. 181 (4): 894–904.e9. doi:10.1016/j.cell.2020.03.045. PMC 7144619. PMID 32275855.
- ^ Xu X, Chen P, Wang J, Feng J, Zhou H, Li X, Zhong W, Hao P (March 2020). "Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission". Science China Life Sciences. 63 (3): 457–460. doi:10.1007/s11427-020-1637-5. PMC 7089049. PMID 32009228.
- ^ Letko M, Marzi A, Munster V (April 2020). "Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses". Nature Microbiology. 5 (4): 562–569. doi:10.1038/s41564-020-0688-y. PMC 7095430. PMID 32094589.
- ^ Letko M, Marzi A, Munster V (April 2020). "Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses". Nature Microbiology. 5 (4): 562–569. doi:10.1038/s41564-020-0688-y. PMC 7095430. PMID 32094589.
- ^ El Sahly HM. "Genomic Characterization of the 2019 Novel Coronavirus". The New England Journal of Medicine. Archived from the original on 17 February 2020. Retrieved 9 February 2020.
- ^ "Novel coronavirus structure reveals targets for vaccines and treatments". National Institutes of Health (NIH). 2 March 2020. Archived from the original on 1 April 2020. Retrieved 3 April 2020.
- ^ Wang K, Chen W, Zhang Z, Deng Y, Lian JQ, Du P, Wei D, Zhang Y, Sun XX, Gong L, Yang X, He L, Zhang L, Yang Z, Geng JJ, Chen R, Zhang H, Wang B, Zhu YM, Nan G, Jiang JL, Li L, Wu J, Lin P, Huang W, Xie L, Zheng ZH, Zhang K, Miao JL, Cui HY, Huang M, Zhang J, Fu L, Yang XM, Zhao Z, Sun S, Gu H, Wang Z, Wang CF, Lu Y, Liu YY, Wang QY, Bian H, Zhu P, Chen ZN (December 2020). "CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells". Signal Transduction and Targeted Therapy. 5 (1): 283. bioRxiv 10.1101/2020.03.14.988345. doi:10.1038/s41392-020-00426-x. PMC 7714896. PMID 33277466. S2CID 214725955.
- ^ Zamorano Cuervo N, Grandvaux N (November 2020). "ACE2: Evidence of role as entry receptor for SARS-CoV-2 and implications in comorbidities". eLife. 9. doi:10.7554/eLife.61390. PMC 7652413. PMID 33164751.
- ^ "Anatomy of a Killer: Understanding SARS-CoV-2 and the drugs that might lessen its power". The Economist. 12 March 2020. Archived from the original on 14 March 2020. Retrieved 14 March 2020.
- ^ Beeching NJ, Fletcher TE, Fowler R (22 May 2020). "BMJ Best Practice: Coronavirus Disease 2019 (COVID-19)" (PDF). BMJ. Archived (PDF) from the original on 13 June 2020. Retrieved 25 May 2020.
- ^ Drayman N, DeMarco JK, Jones KA, Azizi SA, Froggatt HM, Tan K, Maltseva NI, Chen S, Nicolaescu V, Dvorkin S, Furlong K, Kathayat RS, Firpo MR, Mastrodomenico V, Bruce EA, Schmidt MM, Jedrzejczak R, Muñoz-Alía MÁ, Schuster B, Nair V, Han KY, O'Brien A, Tomatsidou A, Meyer B, Vignuzzi M, Missiakas D, Botten JW, Brooke CB, Lee H, Baker SC, Mounce BC, Heaton NS, Severson WE, Palmer KE, Dickinson BC, Joachimiak A, Randall G, Tay S (August 2021). "Masitinib is a broad coronavirus 3CL inhibitor that blocks replication of SARS-CoV-2". Science. 373 (6557): 931–936. Bibcode:2021Sci...373..931D. doi:10.1126/science.abg5827. PMC 8809056. PMID 34285133.
- ^ Fact sheet for healthcare providers: Emergency Use Authorization for Paxlovid (PDF). Pfizer. 22 December 2021. Archived from the original on 23 December 2021.
- ^ "Paxlovid EPAR". European Medicines Agency (EMA). 24 January 2022. Retrieved 3 February 2022. Text was copied from this source which is copyright European Medicines Agency. Reproduction is authorized provided the source is acknowledged.
- ^ Oral COVID-19 antiviral, Paxlovid, approved by UK regulator. Medicines and Healthcare products Regulatory Agency. 31 December 2021.
- ^ Health Canada authorizes Paxlovid for patients with mild to moderate COVID-19 at high risk of developing serious disease. Health Canada. 17 January 2022. Retrieved 24 April 2022.
- ^ FDA Authorizes First Oral Antiviral for Treatment of COVID-19. U.S. Food and Drug Administration (FDA). 22 December 2021. Retrieved 22 December 2021.
 This article incorporates text from this source, which is in the public domain.
This article incorporates text from this source, which is in the public domain.
- ^ Whipple T (23 October 2021). "Moonshot is the spanner in the Covid-19 works the country needs". The Times. Retrieved 5 November 2021.
- ^ Rocklöv J, Sjödin H, Wilder-Smith A (May 2020). "COVID-19 outbreak on the Diamond Princess cruise ship: estimating the epidemic potential and effectiveness of public health countermeasures". Journal of Travel Medicine. 27 (3). doi:10.1093/jtm/taaa030. PMC 7107563. PMID 32109273.
- ^ Ke R, Romero-Severson E, Sanche S, Hengartner N (May 2021). "Estimating the reproductive number R0 of SARS-CoV-2 in the United States and eight European countries and implications for vaccination". Journal of Theoretical Biology. 517: 110621. Bibcode:2021JThBi.51710621K. doi:10.1016/j.jtbi.2021.110621. PMC 7880839. PMID 33587929.
- ^ Liu Y, Gayle AA, Wilder-Smith A, Rocklöv J (March 2020). "The reproductive number of COVID-19 is higher compared to SARS coronavirus". Journal of Travel Medicine. 27 (2): taaa021. doi:10.1093/jtm/taaa021. PMC 7074654. PMID 32052846.
- ^ Davies NG, Abbott S, Barnard RC, Jarvis CI, Kucharski AJ, Munday JD, Pearson CA, Russell TW, Tully DC, Washburne AD, Wenseleers T, Gimma A, Waites W, Wong KL, van Zandvoort K, Silverman JD, Diaz-Ordaz K, Keogh R, Eggo RM, Funk S, Jit M, Atkins KE, Edmunds WJ (April 2021). "Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England". Science. 372 (6538): eabg3055. doi:10.1126/science.abg3055. PMC 8128288. PMID 33658326.
- ^ Liu Y, Rocklöv J (October 2021). "The reproductive number of the Delta variant of SARS-CoV-2 is far higher compared to the ancestral SARS-CoV-2 virus". Journal of Travel Medicine. 28 (7): taab124. doi:10.1093/jtm/taab124. PMC 8436367. PMID 34369565.
- ^ a b c d "COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU)". ArcGIS. Johns Hopkins University. Retrieved 10 March 2023.
- ^ Branswell H (30 January 2020). "Limited data on coronavirus may be skewing assumptions about severity". STAT. Archived from the original on 1 February 2020. Retrieved 13 March 2020.
- ^ Wu JT, Leung K, Leung GM (February 2020). "Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study". Lancet. 395 (10225): 689–697. doi:10.1016/S0140-6736(20)30260-9. PMC 7159271. PMID 32014114.
- ^ Boseley S, McCurry J (30 January 2020). "Coronavirus deaths leap in China as countries struggle to evacuate citizens". The Guardian. Archived from the original on 6 February 2020. Retrieved 10 March 2020.
- ^ Paulinus A (25 February 2020). "Coronavirus: China to repay Africa in safeguarding public health". The Sun. Archived from the original on 9 March 2020. Retrieved 10 March 2020.
Further reading
- Bar-On YM, Flamholz A, Phillips R, Milo R (April 2020). "SARS-CoV-2 (COVID-19) by the numbers". eLife. 9. arXiv:2003.12886. Bibcode:2020arXiv200312886B. doi:10.7554/eLife.57309. PMC 7224694. PMID 32228860.
- Brüssow H (May 2020). "The Novel Coronavirus - A Snapshot of Current Knowledge". Microbial Biotechnology. 13 (3): 607–612. doi:10.1111/1751-7915.13557. PMC 7111068. PMID 32144890.
- Cascella M, Rajnik M, Aleem A, Dulebohn S, Di Napoli R (January 2020). "Features, Evaluation and Treatment Coronavirus (COVID-19)". StatPearls. PMID 32150360. Archived from the original on 6 April 2020. Retrieved 4 April 2020.
- Laboratory testing for coronavirus disease 2019 (COVID-19) in suspected human cases (Report). World Health Organization. 2 March 2020. hdl:10665/331329.
- Zoumpourlis V, Goulielmaki M, Rizos E, Baliou S, Spandidos DA (October 2020). "[Comment] The COVID‑19 pandemic as a scientific and social challenge in the 21st century". Molecular Medicine Reports (Review). 22 (4): 3035–3048. doi:10.3892/mmr.2020.11393. PMC 7453598. PMID 32945405.
External links
- "Coronavirus Disease 2019 (COVID-19)". Centers for Disease Control and Prevention (CDC). 11 February 2020.
- "Coronavirus disease (COVID-19) Pandemic". World Health Organization (WHO).
- "SARS-CoV-2 (Severe acute respiratory syndrome coronavirus 2) Sequences". National Center for Biotechnology Information (NCBI).
- "COVID-19 Resource Centre". The Lancet.
- "Coronavirus (Covid-19)". The New England Journal of Medicine.
- "Covid-19: Novel Coronavirus Outbreak". Wiley. Archived from the original on 24 September 2020. Retrieved 13 February 2020.
- "SARS-CoV-2". Virus Pathogen Database and Analysis Resource.
- "SARS-CoV-2 related protein structures". Protein Data Bank.
Notes
This article is a direct transclusion of the Wikipedia article and therefore may not meet the same editing standards as LIMSwiki.