Journal:Efficient sample tracking with OpenLabFramework
| Full article title | Efficient sample tracking with OpenLabFramework |
|---|---|
| Journal | Scientific Reports |
| Author(s) | List, Markus; Schmidt, Steffen; Trojnar, Jakub; Thomas, Jochen; Thomassen, Mads; Kruse, Torben A.; Tan, Qihua; Baumbach, Jan; Mollenhauer, Jan |
| Author affiliation(s) | University of Southern Denmark, io-consultants GmbH & Co. KG |
| Primary contact | Email: http://www.nature.com/articles/srep04278/email/correspondent/c1/new (Requires login) |
| Year published | 2014 |
| Volume and issue | 4 |
| Page(s) | 4278 |
| DOI | 10.1038/srep04278 |
| ISSN | 2045-2322 |
| Distribution license | Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported |
| Website | http://www.nature.com/articles/srep04278 |
| Download | http://www.nature.com/articles/srep04278.pdf (PDF) |
Abstract
The advance of new technologies in biomedical research has led to a dramatic growth in experimental throughput. Projects therefore steadily grow in size and involve a larger number of researchers. Spreadsheets traditionally used are thus no longer suitable for keeping track of the vast amounts of samples created and need to be replaced with state-of-the-art laboratory information management systems. Such systems have been developed in large numbers, but they are often limited to specific research domains and types of data. One domain so far neglected is the management of libraries of vector clones and genetically engineered cell lines. OpenLabFramework is a newly developed web-application for sample tracking, particularly laid out to fill this gap, but with an open architecture allowing it to be extended for other biological materials and functional data. Its sample tracking mechanism is fully customizable and aids productivity further through support for mobile devices and barcoded labels.
Introduction
With the development of high-throughput technologies, laboratory work has seen a paradigm shift from small projects involving single or few researchers towards large-scale projects involving several laboratories and often hundreds or thousands of samples. Sample management is therefore a growing issue, especially since most laboratories still attempt to keep track of their samples using spreadsheet tools. A high turn-over of academic staff coupled with maintenance of individual files that are often locked or outdated, as well as inconsistent nomenclature and labeling, can lead to tedious repetition of previously existing work. The significant amount of time that is often spent on locating samples would be better used for performing experiments. Moreover, expensive storage space is wasted, since samples are often not labeled properly and cannot be identified. Even if a label is given, it usually does not include a standardized minimal amount of information that allows unambiguous identification of the materials or the experiments they were derived from. Numerous commercial and open-source solutions have been developed in an attempt to overcome these problems.
Although solutions are offered by commercial companies like LabVantage, most academic laboratories find it difficult to afford the license costs, which usually rise with additional users and technical features. The focus of this paper is thus open-source systems.
As Table 1 shows, open-source laboratory information management systems (LIMS) are often customized towards specific types of biomaterials or research data, as for instance genotyping[1][2][3], protein production[4][5], protein-protein-interaction[6], 2D gel electrophoresis[7], or protein crystallography[8] data. Some generic LIMS target specific laboratory tasks, such as sample management[3][9][10], laboratory work-flows and protocols[2][11][12][13], documentation, management of lab stocks, or clinical studies.[14] Further solutions exist for molecular genetics and the creation of vector libraries.[15] There is, however, no dedicated LIMS for the management of large vector construct and cell line libraries. At our Lundbeck Foundation Center of Excellence in Nanomedicine (NanoCAN) at the University of Southern Denmark in Odense such large-scale libraries need to be handled efficiently (see Mollenhauer et al.[16] for a short overview about our work). This motivated us to develop a novel open-source LIMS platform: OpenLabFramework (OLF).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Results
Any LIMS that involves sample management on a large scale should fulfill a number of requirements listed in the following as R1-15. Existing open-source LIMS fulfill these requirements to varying degrees (Table 2).
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Implementation
A LIMS for an academic environment needs to be open-source (R1), in order to save costs and to allow for adaptation to the specific requirements of a given scientific field and laboratory. Since adaptation can be a difficult and time-consuming task, a LIMS that is modular and extensible by design (R2) would be most appropriate. Although difficult to assess for existing projects, a LIMS should be reliable and its implementation simple. Existing frameworks and software packages that are maintained and tested by a large community are often more reliable than individual solutions and should thus be incorporated.
Data handling
Dealing with a large number of samples in a library or biobank requires efficient mechanisms for sample management (R3) and physical sample tracking over several hierarchical levels (R4). Since related information and experimental results are usually stored in additional documents, a management system, where files can be linked to an arbitrary number of samples (R5), would be most useful. Another requirement is that raw data previously entered into the system can be exported to various file formats. This requirement is usually met through an integrated reporting mechanism (R6).
Flexibility in deployment
Academic laboratories are often part of an existing IT infrastructure, but support is in many cases limited, e.g. to a single database management system (DBMS), such as MySQL. LIMS deployment should thus be as flexible as possible not be bound to a specific operating system or DBMS. While the first requirement is fulfilled by all LIMS considered in this paper, multiple database support remains an issue (R7). Furthermore, if a suitable server is not available, deployment locally (R8) or to a cloud service (R9) is advantageous.
User acceptance and excess value
Triplet et al. have identified approachability as a major hurdle in the acceptance of a LIMS.[9] Modern web-technologies like Ajax allow for a more responsive and intuitive user interface, which in turn improves the user experience and reduces the learning period. Another crucial requirement for a successful adaptation of a LIMS is good documentation (R10). User acceptance can also be improved by offering an excess value over traditional spreadsheet tools, for instance by incorporating the use of barcodes (R11), label printing (R12), and mobile devices, such as smartphones (R13). A further advantage would be the incorporation of data analysis tools directly within the LIMS (R14).
Security
LIMS typically address security concerns by restricting access through secure user logins and different user roles. Security would also be enhanced by audit logging features (R15), where a version number is added to each database entry. Any change will then result in a copy of the entry with a new version number, so that accidentally overwritten entries can be restored.
OpenLabFramework
We present OpenLabFramework (OLF), a laboratory information management system (LIMS) primarily targeted at advanced sample and storage management in mid-sized laboratories with less than 50 users. It facilitates a seamless integration of virtual and real world storage handling by making use of mobile devices, which are carried by lab personal anyways, in combination with cheap and fully integrated barcode labeling technology. In the following we shed a light on how OLF fulfills the LIMS requirements that we have identified before (R1–R15). A brief comparison with existing open-source LIMS is given in Table 2.
Modularity and extendibility
OLF is published as open-source (R1) and, due to its modular structure, it can be adapted to different types of laboratory data and sample types. New functionality can also be added and integrated (R2). Various features are covered by the following modules.
GeneTracker: GeneTracker is intended to fulfill requirements specific to the hierarchical organization of genes, gene variants, vector constructs, and genetically engineered cell lines, thus helping to keep track of extensive sample libraries in the field of targeted genomics. The organization of these samples is further supported through OLF's built-in user and project management features.
Sample storage: The Storage module adds options for tracking and organizing samples in a customizable storage infrastructure (R3–4). This infrastructure is hierarchical, starting from buildings and rooms and ending in individual freezers and storage boxes. Interactive grids help the user to assess the content of a storage box at a glance. Together with GeneTracker, samples can be added or removed from storage in an intuitive manner, while providing an overview of remaining copies and related samples.
File uploads: The FileAttachments module allows users to up- and download arbitrary files, allowing for a better organization of their results and documents. Files are stored with a combination of timestamp and original file name to avoid conflicts arising from identical file names. Files are uploaded to a configurable folder on the server and not to the database itself. They can be linked to an arbitrary number of samples, so that other users can quickly obtain an overview of files relevant to a sample (R5).
Barcode and label support: The functionality of the Storage module is complemented by the Barcode module, with which a user can create and print barcode labels (R11–12). These can later be used to locate a sample in OLF by scanning the barcode using a USB-connected scanner or a mobile device (R13). The Barcode module currently requires a connected DYMO label printer but can be extended in the future to support other devices.
Reporting
Apache POI is utilized to export lists of samples to various file formats, including Excel (XLSX), Open Document Spreadsheets (ODS), PDF, and comma separated values (CSV). This feature is currently available for lists of genes, vector constructs, and cell lines. The storage hierarchy and individual boxes can also be exported to Excel spreadsheets (R6).
Flexibility
Grails applications are not bound to a specific database management system and will even work with non-SQL solutions, such as MongoDB (R7). OLF is compiled either as WAR file, which is suitable for deployment on a large number of Java-based web containers, or as locally executable JAR file, which comes packed with its own web container and file-based SQL solution (R8). It should be noted that OLF has only been tested thoroughly on Tomcat versions 6 and 7.
Cloud deployment
Grails also offers a plug-in for cloud deployment using the VMware Cloud-Foundry service (http://www.cloudfoundry.com/) (R9). Apart from CloudFoundry credentials and memory settings, no further configuration is needed. Upon deployment CloudFoundry automatically configures a suitable database to work with the application.
Mobile support
OLF utilizes the Spring Mobile Grails plug-in to distinguish mobile clients from desktop clients. If a mobile device is detected, a different view is shown that is tailored for the small-sized screen and touch-screen interaction (R13).
User approachability and excess value
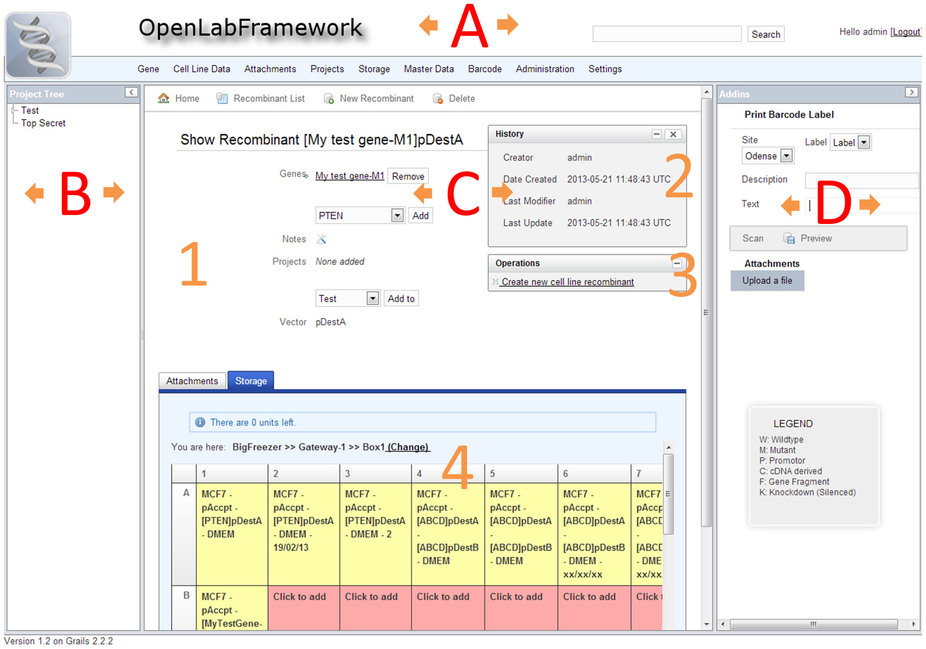
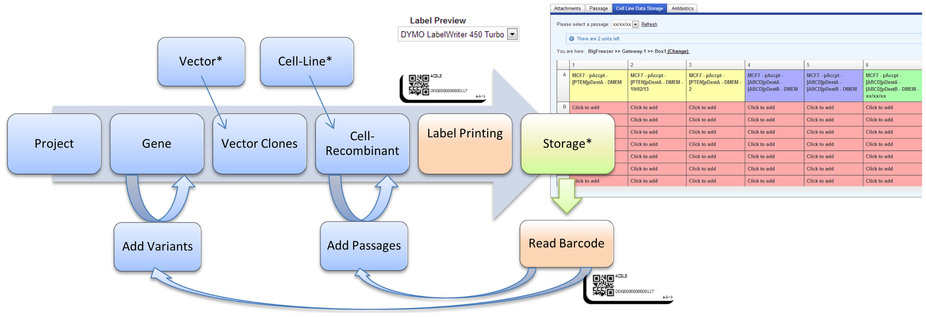
OLF offers a modern web-interface that is clearly organized and intuitive (Figure 1), and allows for responsive user interaction. The Compass-powered search engine allows users to locate required information quickly and conveniently. Users can also develop effective laboratory work-flows using the sample tracking feature together with barcode labels and mobile devices (Figure 2). OLF validates all user entered data for validity and will, where applicable, provide a list of viable options in form of select boxes. In this way, OLF effectively avoids ambiguity and ensures consistency of sample data. Finally, OLF comes with online documentation that introduces the system to users, administrators, and software developers (R10).
|
|
Example for practical implementation and user acceptance - OLF at the NanoCAN Center
Within the past three years since the introduction of the first version of OLF in 2010 at the Lundbeckfonden Center of Excellence NanoCAN, around 780 genes, 1,200 vector constructs, and 300 cell lines have been added to the system, along with 1,500 associated sample locations. Data are stored on a Microsoft SQL Server 2008 installation with a database size of approximately 7 MB. The more than 20 scientists engaged in functional genomics projects were introduced to the system through a one hour feature presentation, which enabled them to use OLF productively. The system was reported to be intuitive, albeit only one user had previous experience in using a LIMS system. Missing functionality considered useful for increased productivity and user convenience, such as handling of barcoded labels, were added to the system subsequently.
Recommended system configuration
OLF relies on a database backend for storing sample related data. However, only primitive data types, such as numbers or text fields, are persisted to the database itself, while all documents and files are stored in a folder and merely linked in the database. In this way, we expect a database size of less than 10–20 MB for most use cases. For smaller laboratories with less than 20 members, the file-based SQL database that is part of OLF's standalone version is appropriate. Since no significant processing of data is required, OLF is expected to be responsive even on systems with a single CPU core. For larger laboratories, however, we recommend using a system with multiple CPU cores and a dedicated database management system, such as MySQL for efficiently dealing with concurrent access in a responsive manner. Due to its dynamic nature, OLF has a large memory footprint and we strongly recommend providing a minimum of 768 MB of RAM on Linux and 1024 MB on Windows.
Methods
Grails web-application framework
We considered the Java based framework Grails to be the most promising candidate for a building a web-application. Grails is a VMWare/SpringSource product and builds on the company's experience in industry-standard frameworks such as Hibernate or Spring, which also form the core of Grails. In Grails, plug-ins deliver high-quality solutions for non-trivial web application tasks, e.g. database search (Compass and Apache Lucene), spreadsheet im- and export (Apache POI) and a user/security management (SpringSecurity). Grails embraces the paradigms convention over configuration and separation of concerns to keep the code concise and clean. Furthermore, Grails hides the complexity of data persistence with an object relational modeling technique, which encapsulates all database interactions and models them through Java domain classes. These characteristics allow faster development and integration of new features.
OpenLabFramework
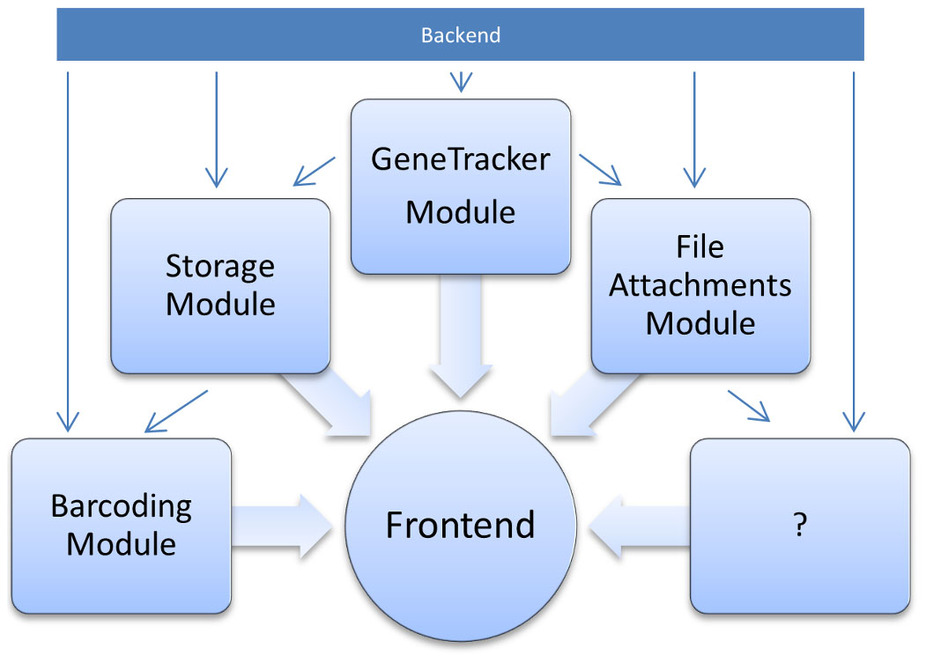
As illustrated in Figure 3, OLF has a modular structure, in which a back-end plug-in provides the necessary base classes, as well as user, project and security management. Content plug-ins can then add arbitrary classes and view templates, and integrate with other plug-ins. All plug-ins are finally merged in the front-end application, which utilizes the scaffolding mechanism of Grails to dynamically create Ajax-driven views for user interaction.
|
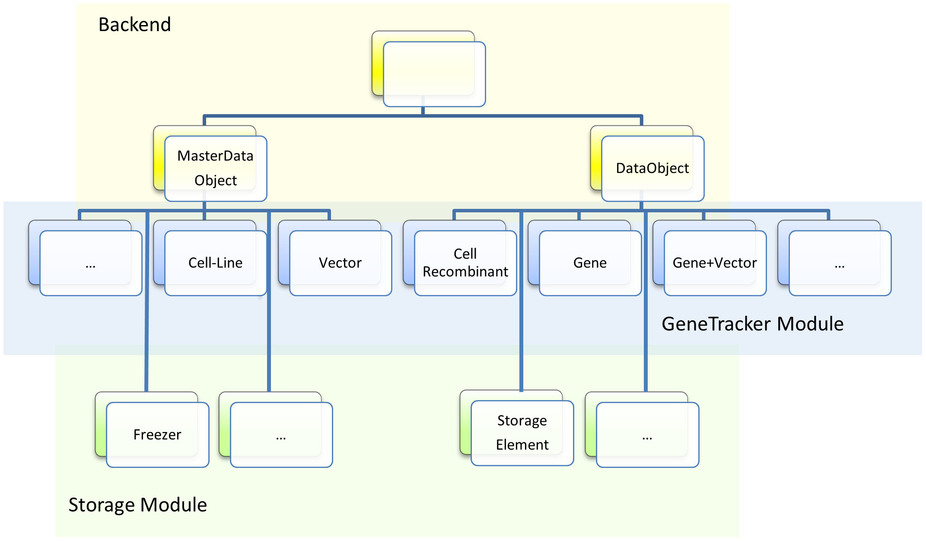
The back-end of OLF introduces two base classes called MasterDataObject and DataObject (Figure 4). MasterDataObjects can be extended by classes representing master data that are maintained by system administrators, e.g. wildtype cell lines or vector systems in the GeneTracker module, or freezers and storage locations in the Storage module. More dynamic content is expressed through DataObject classes, which can be created and modified by regular users, e.g. genes or genetically engineered cell lines in the GeneTracker module, or StorageElements, which contain location data for other DataObject instances, in the StorageModule. This hierarchy makes OLF highly generic, but also allows fine-grained interactions between more specialized classes.
|
OLF strictly follows established patterns found in software architecture. Code is separated into different functional layers according to the model-view-controller pattern. In addition, business logic related code is bundled in service classes and tag libraries to avoid code duplication. Furthermore, OLF introduces the concept of content modules, which can be attached to existing pages. Content modules can either contribute new links to OLF's menu, render additional add-ins or tabs, or provide additional links called operations (Figure 1). By providing a clear structure and suitable interfaces, other developers can thus easily contribute to OLF.
Discussion
Numerous commercial and open-source laboratory information management systems (LIMS) exist today. However, since commercial licenses are expensive and lack the possibility to be adapted to specific needs without additional costs, academic laboratories usually focus on finding an open-source solution to their sample management issues. Moreover, none of the existing solutions seems optimal for all given tasks (Table 2). Naturally, most LIMS are dedicated to a specific field of research and are thus not generally suited for other fields. Some solutions, on the other hand, focus on certain general aspects of laboratory work, such as sample tracking, protocols, or work-flows.
OpenLabFramework (OLF) was developed to address the need for an open-source LIMS solution for covering vector constructs and cell line library sample tracking. Acknowledging that many LIMS remain limited to their research domain, we created OLF in a strictly modular and extensible fashion, with dedicated modules for sample tracking, barcoded label printing, and file management. We expect that OLF can be adapted to other research fields and biomaterials with minimal developmental effort by implementing a content module similar to GeneTracker, which is then complemented by plugging in additional features as needed.
We built OLF using Grails and its extensive plug-in eco-system. This allows OLF to satisfy basic software development requirements, such as flexibility, reliability and simplicity. The use of a solid framework allows developers to focus on user-specific requirements, as for instance the support for mobile devices, and to keep the application up-to-date, since it will improve together with the underlying framework. The use of Grails, which can be considered the most dynamic and flexible web-application framework available in Java, together with its plug-ins, poses a significant simplification when adding new web-application features. This advantage separates OLF from comparable LIMS, which also embrace the concept of modularity or the use of web-application frameworks.
The introduction of OLF allows controlling sample logistics effectively, which is a particular challenge upon movement, turn-over of lab staff, and improper labeling of samples. OLF may further increase productivity by including modern technologies so far disregarded by most other open-source LIMS, such as printing and reading barcode labels. The high degree of automation and standardization that can be achieved by this may substantially reduce user-caused errors in sample assignment. A web-layer for mobile devices provides an additional advantage. In this way, samples can now be removed from physical and virtual storage at the same time, thus limiting the risk of forgetting this step after the work with the sample is completed. As illustrated in Figure 2, the implementation of OLF in a laboratory environment can lead to a significantly more productive work-flow.
Finally, unlike most LIMS, OLF is not bound to a specific database or web-container. OLF can be coupled with a large number of database management systems, including non-SQL solutions like MongoDB. If a suitable server is not available, OLF can be installed locally or even be deployed to the cloud with little effort, as demonstrated in our demo application. This flexibility will reduce technical hurdles in the introduction of OLF to a new laboratory.
OLF offers efficient and user-friendly management of sample information and location in the field of high-throughput biology and functional genomics. Being extensible, it can be adapted to satisfy additional requirements with little developmental effort. One requirement currently neglected is the extensive integration of additional web tools, which would make OLF more attractive to end-users. Although result data files can be uploaded and linked to samples, we envision that direct interfacing with laboratory equipment would make result data management significantly more convenient. The implementation of RESTful web services could expose OLF data sets and functionality to other tools and information systems. Along with this, the reporting capabilities could be improved, allowing for customized reports, where information from several instances is pooled. This would also help in establishing data analysis directly within OLF or through integration of additional tools (R15). Another important aspect worth considering is that OLF might in some instances require a more fine-grained access to the data. This functionality could be added through additional user roles or access control lists. Finally, the creation of a dedicated app for mobile platforms, such as Android or iOS, would improve the mobile user experience.
Conclusions
In order to retain an overview over large sample libraries typically found in nowadays laboratories, an efficient system for management and tracking of samples is required. OpenLabFramework (OLF) has been developed with focus on vector construct and cell line libraries. Thanks to its modularity, however, it can be adapted to new scenarios. OLF can be deployed using different database management systems either locally, to a server, or to the cloud. The incorporation of modern technologies, such as mobile devices and printing of barcode labels may increase productivity even further. These properties together may be considered characteristic for a next-generation LIMS and should provide the potential for widespread adaptation.
OLF embraces open-source in the hope of attracting not only laboratories in need of a LIMS, but also a community of software developers willing to adapt OLF to new scenarios. We intend to contribute in the future by developing further modules, e.g. for seamless evaluation of experimental data by integration of third party tools. Consequent utilization of community-proven open-source libraries make OLF's backend already highly reliable. With the support of its own community, OLF as a whole is expected to reach the same high quality standard in the future.
Additional information
Availability and Requirements
Project name: OpenLabFramework
URL: https://github.com/NanoCAN/OpenLabFramework
Wiki: https://github.com/NanoCAN/OpenLabFramework/wiki
Demo: http://www.nanocan.dk/openlabframework/demo (user: admin, password: demo0815)
Operating system(s): Platform independent
Programming language: Java, Groovy, Java-script
Other requirements: Java 1.6 or higher, Tomcat 6.0 or higher
License: GNU GPL v3
Acknowledgements
This work was supported by the Lundbeckfonden grant for the NanoCAN Center of Excellence in Nanomedicine, the Region Syddanmarks ph.d.-pulje and Forskningspulje, the Fonden Til Lægevidenskabens Fremme, and co-financed by the INTERREG 4 A-program Syddanmark-Schleswig-K.E.R.N. with funds from The European Regional Development Fund.
Author information
Affiliations
Lundbeckfonden Center of Excellence in Nanomedicine NanoCAN, University of Southern Denmark, Odense, DK
Markus List, Steffen Schmidt, Jakub Trojnar, Mads Thomassen, Torben A. Kruse & Jan Mollenhauer
Institute of Molecular Medicin (IMM), University of Southern Denmark, Odense, DK
Markus List, Steffen Schmidt, Jakub Trojnar & Jan Mollenhauer
Clinical Institute (CI), University of Southern Denmark, Odense, DK
Markus List, Mads Thomassen, Torben A. Kruse & Qihua Tan
Epidemiology, Biostatistics and Biodemography, Institute of Public Health, University of Southern Denmark, Odense, DK
Qihua Tan
Department of Biochemistry and Molecular Biology (BMB), University of Southern Denmark, Odense, DK
Jakub Trojnar
io-consultants GmbH & Co. KG, Heidelberg, DE
Jochen Thomas
Department of Mathematics and Computer Science (IMADA), University of Southern Denmark, Odense, DK
Jan Baumbach
Contributions
M.L. and J.Th. developed the software requirement specification. S.S. and J.Tr. specified requirements specific to the biomedical domain and contributed extensively to bug-tracking and regular testing of OLF. ML implemented the application. M.T., T.K., Q.T., J.B. and J.M. jointly supervised the project, designed graphics and work-flow diagrams. All authors contributed equally to drafting the manuscript. All authors read and approved the final manuscript.
Competing interests
The authors declare no competing financial interests.
Corresponding author
Correspondence to Markus List.
Supplementary information
PDF files 1.Supplementary Information: OpenLabFramework Supplemental Material
References
- ↑ 1.0 1.1 Sanchez-Villeda, H.; Schroeder, S.; Polacco, M. et al. (2003). "Development of an integrated laboratory information management system for the maize mapping project". Bioinformatics 19 (16): 2022-2030. doi:10.1093/bioinformatics/btg274. PMID 14594706.
- ↑ 2.0 2.1 2.2 Jayashree, B.; Reddy, P.T.; Leeladevi, Y. et al. (2006). "Laboratory information management software for genotyping workflows: Applications in high throughput crop genotyping". BMC Bioinformatics 7: 383. doi:10.1186/1471-2105-7-383. PMC PMC1559653. PMID 16914063. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1559653.
- ↑ 3.0 3.1 3.2 Voegele, C.; Alteyrac, L.; Caboux, E. et al. (2010). "A sample storage management system for biobanks". Bioinformatics 26 (21): 2798-2800. doi:10.1093/bioinformatics/btq502. PMID 20807837.
- ↑ 4.0 4.1 Morris, C.; Pajon, A.; Griffiths, S.L. et al. (2011). "The Protein Information Management System (PiMS): A generic tool for any structural biology research laboratory". Acta Crystallographica Section D 67 (4): 249–260. doi:10.1107/S0907444911007943. PMC PMC3069740. PMID 21460443. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3069740.
- ↑ 5.0 5.1 Ponko, S.C.; Bienvenue, D. (2012). "ProteinTracker: An application for managing protein production and purification". BMC Research Notes 5: 224. doi:10.1186/1756-0500-5-224. PMC PMC3436699. PMID 22574679. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3436699.
- ↑ 6.0 6.1 Droit, A.; Hunter, J.M.; Rouleau, M. et al. (2007). "PARPs database: A LIMS systems for protein-protein interaction data mining or laboratory information management system". BMC Bioinformatics 8: 483. doi:10.1186/1471-2105-8-483. PMC PMC2266781. PMID 18093328. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2266781.
- ↑ 7.0 7.1 Morisawa, H.; Hirota, M.; Toda, T. (2006). "Development of an open source laboratory information management system for 2-D gel electrophoresis-based proteomics workflow". BMC Bioinformatics 7: 430. doi:10.1186/1471-2105-7-430. PMC PMC1599757. PMID 17018156. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1599757.
- ↑ 8.0 8.1 Haebel, P.W.; Arcus, V.L.; Baker, E.N.; Metcalf, P. (2001). "LISA: An intranet-based flexible database for protein crystallography project management". Acta Crystallographica Section D 57 (Pt 9): 1341-1343. doi:10.1107/S0907444901009295. PMID 11526339.
- ↑ 9.0 9.1 9.2 Triplet, T.; Butler, G. (2012). "The EnzymeTracker: An open-source laboratory information management system for sample tracking". BMC Bioinformatics 13 (15): 1341-1343. doi:10.1186/1471-2105-13-15. PMC PMC3353834. PMID 22280360. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3353834.
- ↑ 10.0 10.1 Olhovsky, M.; Williton, K.; Dai, A.Y. et al. (2011). "OpenFreezer: A reagent information management software system". Nature Methods 8 (8): 612–613. doi:10.1038/nmeth.1658. PMID 21799493.
- ↑ 11.0 11.1 Stocker, G.; Fischer, M.; Rieder, D. et al. (2009). "iLAP: a workflow-driven software for experimental protocol development, data acquisition and analysis". BMC Bioinformatics 10: 390. doi:10.1186/1471-2105-10-390. PMC PMC2789074. PMID 19941647. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2789074.
- ↑ 12.0 12.1 Melo, Alexandre; Alessandra Faria-Campos; Daiane M DeLaat; Rodrigo Keller; Vinícius Abreu; Sérgio Campos (2010). "SIGLa: an adaptable LIMS for multiple laboratories". BMC Genomics 11 (Suppl 5): S8. doi:10.1186/1471-2164-11-S5-S8. PMC PMC3045801. PMID 21210974. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3045801.
- ↑ 13.0 13.1 Burgarella, S.; Cattaneo, D.; Pinciroli, F.; Masseroli, M. (2005). "MicroGen: a MIAME compliant web system for microarray experiment information and workflow management". BMC Bioinformatics 6 (Suppl 4): S6. doi:10.1186/1471-2105-6-S4-S6. PMC PMC1866379. PMID 16351755. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1866379.
- ↑ 14.0 14.1 Krestyaninova, M.; Zarins, A.; Viksna, J. et al. (2009). "A system for information management in biomedical studies – SIMBioMS". Bioinformatics 25 (20): 2768-2769. doi:10.1093/bioinformatics/btp420. PMC PMC2759553. PMID 19633095. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2759553.
- ↑ 15.0 15.1 Truong, C.V.C.; Groeneveld, L.F.; Morgenstern, B.; Groeneveld, E. (2011). "MolabIS - An integrated information system for storing and managing molecular genetics data". BMC Bioinformatics 12: 425. doi:10.1186/1471-2105-12-425. PMC PMC3268772. PMID 22040322. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3268772.
- ↑ Mollenhauer, J.; Stamou, D.; Flyvbjerg, A. et al. (2010). "David versus Goliath". Nanomedicine 6 (4): 504–509. doi:10.1016/j.nano.2010.04.002. PMID 20417315.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added. In Table 2, checkmarks and Xs were replaced with Ys and Ns. The "Methods" section has also been moved from the end to a more logical position before the Discussion and Conclusion.