Journal:Improving the creation and reporting of structured findings during digital pathology review
| Full article title | Improving the creation and reporting of structured findings during digital pathology review |
|---|---|
| Journal | Journal of Pathology Informatics |
| Author(s) | Cervin, Ida; Molin, Jesper; Lundstrom, Claes |
| Author affiliation(s) | Chalmers University of Technology, Linköping University, Sectra AB |
| Primary contact | Email: Available w/ login |
| Year published | 2016 |
| Volume and issue | 7 |
| Page(s) | 32 |
| DOI | 10.4103/2153-3539.186917 |
| ISSN | 2153-3539 |
| Distribution license | Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported |
| Website | http://www.jpathinformatics.org |
| Download | http://www.jpathinformatics.org/temp/JPatholInform7132-492631_134103.pdf (PDF) |
Abstract
Background: Today, pathology reporting consists of many separate tasks, carried out by multiple people. Common tasks include dictation during case review, transcription, verification of the transcription, report distribution, and reporting the key findings to follow-up registries. Introduction of digital workstations makes it possible to remove some of these tasks and simplify others. This study describes the work presented at the Nordic Symposium on Digital Pathology 2015, in Linköping, Sweden.
Methods: We explored the possibility of having a digital tool that simplifies image review by assisting note-taking, and with minimal extra effort, populates a structured report. Thus, our prototype sees reporting as an activity interleaved with image review rather than a separate final step. We created an interface to collect, sort, and display findings for the most common reporting needs, such as tumor size, grading, and scoring.
Results: The interface was designed to reduce the need to retain partial findings in the head or on paper, while at the same time be structured enough to support automatic extraction of key findings for follow-up registry reporting. The final prototype was evaluated with two pathologists, diagnosing complicated partial mastectomy cases. The pathologists experienced that the prototype aided them during the review and that it created a better overall workflow.
Conclusions: These results show that it is feasible to simplify the reporting tasks in a way that is not distracting, while at the same time being able to automatically extract the key findings. This simplification is possible due to the realization that the structured format needed for automatic extraction of data can be used to offload the pathologists' working memory during the diagnostic review.
Keywords: Digital pathology, structured reporting, usability, workflow
Background
The pathologist's review process with a microscope involves many activities such as reviewing the glass slides, taking notes, dictating the report, or referring to a colleague for advice.[1] Slide review is the most common activity during a session, but a considerable amount of time is spent on the other activities.[2] The main result of the review is distributed as a pathology report with the referring physician as the primary audience.[3] However, the result is also disseminated at multidisciplinary conferences, and certain parameters are reported to national registries for systematic follow-up. Structured (or synoptic) reports have been used successfully to improve this communication with external parties. The structure can act as a checklist and in that way improve the completeness of reporting or can be used as an electronic form to reduce the turn-around time since the additional dictation and transcription steps can be avoided.[4][5] The usability of structured reporting within pathology has, however, not been evaluated. Within radiology, the same benefits have been observed, but with regards to usability, structured reports have been associated with inappropriate interfaces for the reporting needs, and it has been questioned whether the structured format interferes with the image interpretation process.[6] Hence, it is important to take these issues into consideration when developing similar systems. On the other hand, the above studies see the reporting as a separate task alongside the review, and it is therefore only systems of that type that have been evaluated.
With digital pathology, it is instead possible to merge the reporting with the image review since the tasks are performed within the same system. This new possibility has so far received little attention, whereas the focus instead has been on new capabilities in terms of added features such as remote slide interpretation, digital image analysis, improved teleconferencing tools, and improved teaching capabilities.[7]
However, workflow improvements in terms of better integration between systems could be just as important. To improve these aspects, existing working patterns and assumptions need to be challenged. A few viewpoint papers and editorials present different visions of the future of digital pathology by employing this point of view: Krupinski[8] describes the future workstation as the pathologist's cockpit, where all the data needed to produce pathology reports are gathered in one interface including important workflow metrics in a digital dashboard to monitor the overall workflow. Fine[9] outlines a future scenario where digital image analysis algorithms are used to triage cases before review. The pathologist is then chauffeured between the most relevant areas of a case, where measurements are automatically performed. The pathologist can then select which measurements to accept and include into the final report. Hipp et al.[10] compared digital pathology to computational chess, and hypothesized that the best performing computer-aided diagnostic systems will be those that are best able to combine their performance with the pathologists, rather than the systems having best stand-alone performance.
The above visionary descriptions of future systems are clearly valuable for the development of the digital pathology field. In our work, we share their objective to redefine and improve the pathology workflows by removing assumptions that stem from the traditional use of microscopes, but no longer are necessary in a digital environment. Our work goes beyond theoretical visions as we conducted a design-based research study: A task analysis leading to a prototype development and finally a user evaluation of the prototype. The main assumption we challenged was that with microscopic review, the image review and the reporting in the form of dictation are considered to be separate tasks. Instead, we explored the possibilities that arise when building a reporting system that assists the pathologist during the review, as well as prepopulating a large part of the pathology report with structured items generated during the review.
Methods
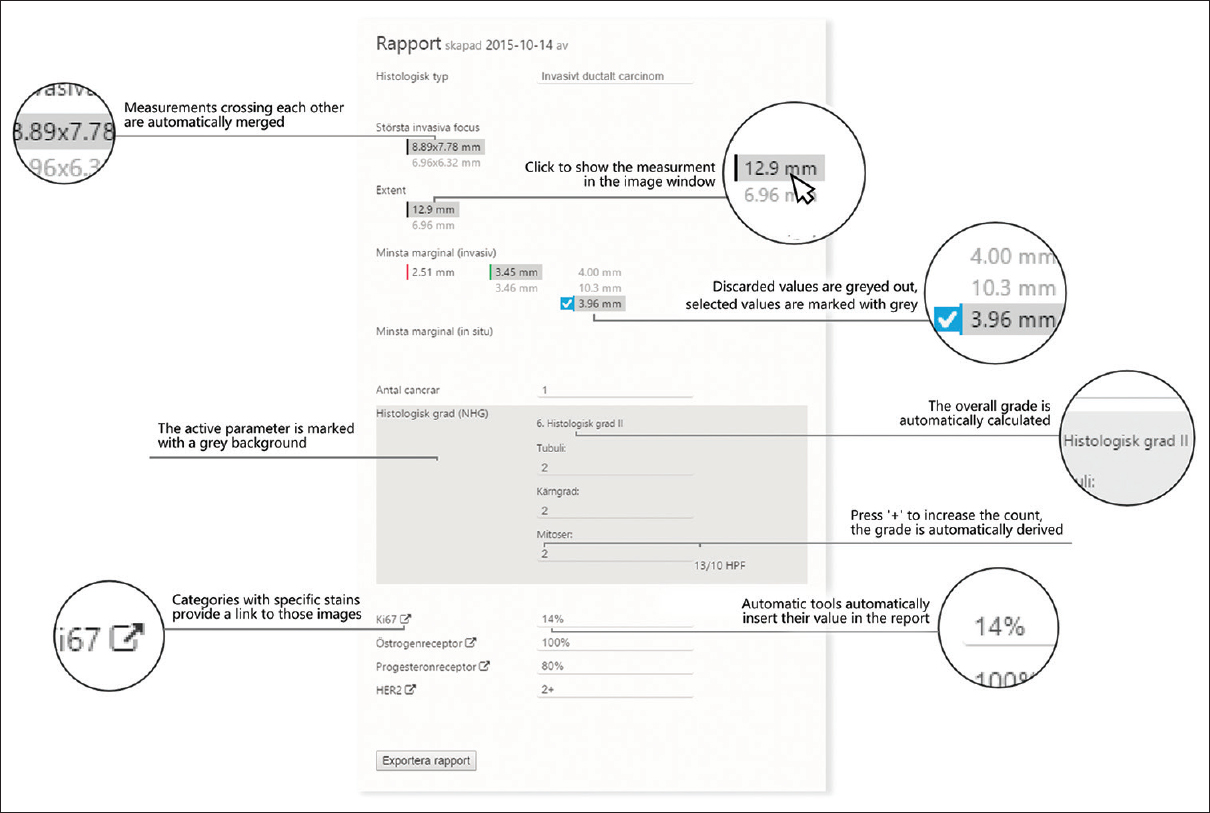
This work constitutes a design-based research study. In design-based research, knowledge is derived from the design process, typically by developing a prototype for a specific problem domain and then retrospectively analyze the artifacts observed throughout the design process.[11] In this study, we developed a prototype to specifically manage reportable findings, both to assist image review and open up for the possibility to extract findings to follow-up registries. The prototype was built in the Sectra IDS7/px viewer (Sectra AB, Linköping, Sweden) as shown in [Figure 1]. Below, we present the design activities that were performed within the project, which we then analyzed using thematic analysis.
|
Design activities
The design activities were performed together with developers at Sectra and three pathologists working at two Swedish hospitals. The design activities included field trips, analysis of anonymized pathology reports, paper prototypes, video prototypes, software prototypes, and a user study. The activities were performed during a time period of six months during the autumn of 2014 as a master's thesis project.[12] The main activities can be summarized as follows:
Analysis of pathology reports
Pathology reports were studied to determine if any naturally occurring structure was present. Sectra provided anonymized pathology reports from three different regions in Sweden. The reports were from different types of pathology diagnoses and were therefore sorted using text pattern-matching algorithms (regular expressions in C++) to separate different types of reports from each other. The separation took place using certain keywords associated with different types of cancer diagnosis as regular expression. The keywords were determined using the Swedish quality and standardization committee's recommended parameters for certain kind of cancer types.
Prototyping
A pathologist's workflow was studied in real life to get an understanding of the report workflow and image review. This, together with the Swedish quality and standardization committee's recommended parameters for invasive breast cancer, was the base for the interface. An early design of the interface was produced as a paper prototype and it mainly focused on measurements, tasks including cell counting, and reporting and summation of histological grading. The paper prototype was translated into the first web-based prototype which was presented to two pathologists in a demonstrational video. Their feedback was used as the basis for the final prototype.
User study
The final prototype was evaluated in a study with two pathologists. They were tasked to perform diagnostic reviews of two large, single-tumor, partial breast mastectomies. One review was performed with the developed prototype and the other with only pen and paper for note-taking. To create a deep understanding of the workflow implications, a qualitative approach was taken where evaluation results were derived from observations and interviews.
Data sources
One important part of the knowledge generated in this work is the contents of the documents created in the design process. To elicit this knowledge, the information was systematically gathered, encoded, and categorized. All documents created throughout the design activities of the project were filed and organized in physical and digital folders. We prescreened the documents to extract the ones containing data generated within the project. All printed publications and reports that already have been referred to in the background section were excluded from the analysis. The documents were printed or copied before being analyzed, and the resulting set comprised 53 pages of text and 23 pages of images and sketches. The information encoding was performed in several steps. The initial codes were derived by parallel coding of the documents until no new codes were derived; in total, 11 pages were parallel-coded. The initial codes were then summarized into four themes, two themes that mainly were derived from the design process and two themes from the evaluation study. All the documents were then recoded with the four themes using colored highlighters. The highlighted areas were analyzed and summarized.
Results
The results consist of three main components: Insights from design activities, the developed prototype, and the user study. The design insights, divided into two different themes, will be presented first. We then present the final prototype and connect the design choices to the elicited insights. Finally, the results from the final evaluation study of the prototype are described.
Design insights
Two themes were derived from the development of the prototype. The first theme describes the internal and external communication and collaboration where the pathology report is involved. The second theme analyzes the generation and structuring of the data to complete the final report for distribution. Main findings are presented below, illustrated by quotes from the participating pathologists. The quotes have been translated from Swedish, and each quote ends with the quoting pathologist within parentheses.
Communication and collaboration
Pathologists communicate and collaborate externally mainly with the referring physician and internally with lab personnel. When starting to review a case, both the internal gross description and the external request have information that needs to be transferred to the report. This transfer creates a double effort since values are reported several times, as in this field note:
The request, with isolated parameters, can be used to populate a part of the report automatically. Today, parameters are reported several times, such as tumor size which is reported both in the gross description and the report (P1).
The ways of communication within the department do not guarantee that documented values are retained, and the values the pathologists dictate are not guaranteed to be transcribed perfectly. Therefore, they need to keep track of important values themselves:
I keep notes on every single case to be able to control the transcription of the medical secretary (P3).
The pathologists feel a need to create a clear and concise structure in the report to facilitate for the referring physician:
Our system today is very limited, it is not possible to use bold font (…), which makes it unclear what is a heading and what is a response (P1).
Reports should not be too talkative, and additions of descriptive text should only be done when something is hard to assess (P3).
Even though descriptive text preferably should be used as little as possible, it might be necessary in complex cases with a lot of uncertainties:
Sometimes, things do not fit in and you have to write "explanations" and descriptions, therefore it would be preferable to have the option to write descriptive text together with certain parameters (P1).
In the analysis of naturally occurring structure in existing reports, it became clear that the same type of parameters are expressed in various ways. An example of these unnecessary variations is the reporting of vascular invasion in breast cancer reports, which can be expressed as "not detected," "yes," "no," "could not be seen," and "could be seen," when it easily could be expressed with a simple "yes" or "no." Variation in how to express parameters leaves the referring physician with an undesirable interpretation task and an associated risk for misinterpretation:
(…) a benefit is that it is easier to get a more similar answer regardless of who is reporting. Otherwise, it is easy to get personal variations (…) (P1).
The analysis of the reports quickly yielded the insight that many parameters need to be reported in large and complex cases. However, to be able to report one value, you sometimes need to produce many values and then choose one:
The tumor is sectioned into multiple slices, and measurements are performed on all the slices (to determine the size). Then, I choose the largest measurement (…) (P1).
A large amount of measurements increase the need to structure the information that the pathologist generate to remember what parts of the review that have already been performed:
(The pathologist) usually makes a sketch of the whole specimen on a macro level based on the gross description. In this study, (the pathologist) takes notes about the performed measurements (…) (P3).
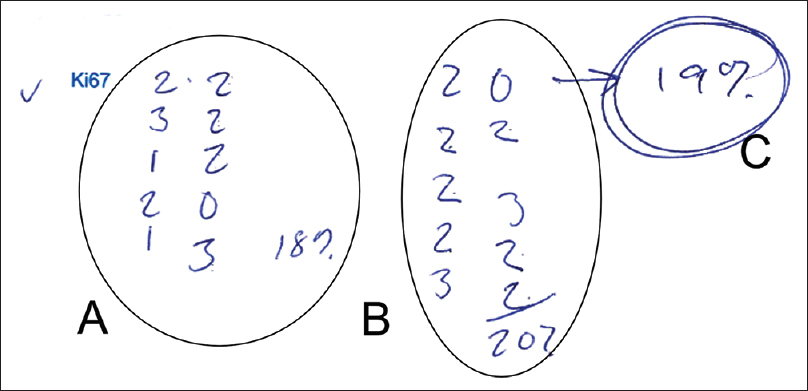
Some manual measurements are quite complicated, and it becomes necessary to take notes to deal with the overload of information. In [Figure 2], a pathologist uses a procedure to quantify the number of positive KI-67 cells by counting 10 nuclei at a time and write down how many were positive and then summarizing the result and calculating the total percentage.
|
This management of data is also used to support the overall reviewing workflow, for example, by keeping track of what slides contain tumor or have already been reviewed.
The main considerations derived from observing the current reporting workflow can be summarized as follows:
- It is important to communicate an understandable report to the referring physician. A more structured format creates a more universal understanding.
- The pathologist often needs to take notes during image review to be able to create, evaluate, and choose what to report.
- Many values, especially measurements, are produced during image review, more than those ending up in the report. These need to be categorized and easily overviewed to decide on what to report.
Prototype functionality
We used the design insights to create a final prototype apt for evaluation in actual use. The prototype was implemented as a plug-in functionality to the existing digital review software, as shown in [Figure 1]. The main idea with the prototype was to leverage the possibility to closely integrate image review and reporting to fulfill the pathologists' needs. From the pilot studies and the development of the final prototype, we derived four different design principles to support the conception of different functions, as follows.
The first design principle was to support at least two types of personal work routines: to work through a case slide by slide or to take one reportable item after another. For example, we implemented support both to make length measurements and then add that to a certain category or to first select a category and then perform the measurement belonging to that category.
The second principle was to divide all possible steps during the generation of a final reportable value into separate steps. We implemented this as support for first collecting as many measurements as you wanted into a certain category, then sorting by clicking each value to quickly go through a collection, followed by selecting which values to report. This way it was possible to reuse the same type of interaction pattern for different types of measurements.
The third principle was to, as much as possible, automatically generate calculations that the pathologist might need to perform. This was implemented for the breast case by automatically summarizing the subscores for the Nottingham histological grade, and to convert the mitotic count to the corresponding grade. Thus, instead of letting the user initiate each calculation, the principle adopted was to generate many calculations and let the pathologist select among them when populating the report.
The fourth and final principle was to support the situation where the template did not cover everything that needed to be reported. This was implemented by supporting the addition of a descriptive text comment to all the reportable parameters.
Each feature is described according to these principles in [Figure 3].
|
Prototype evaluation
The final prototype was then evaluated in a final study with two pathologists diagnosing two partial breast mastectomies. The pathologists performed the same review in two modes. The first mode was using only pen and paper, and the second was using the prototype as well as pen and paper if they wanted to. However, the actual outcome in the second mode was that the pen and paper were hardly used at all by any of the pathologists. This evaluation was recorded by using screen capture and by recording an interview after they had reviewed the cases. These data were also analyzed thematically.
A noteworthy result is that the pathologists' personal work routines were changed as an effect of incorporating the prototype into their review. Using the prototype, one pathologist adopted an approach to finalize all measurements for each slide at once, documenting multiple parameters for each slide. In contrast, the work style of this pathologist with pen and paper was to report each parameter separately across all slides, and then repeating this pattern for each parameter. This was noticed and perceived as a positive change by the pathologist and is described in the following field note:
It is possible to perform multiple tasks in a more structured way. Usually, there is a risk to be interrupted and you have to make sure to perform one task at a time (…). It is hard to keep all things in mind and therefore you work more systematically. The prototype creates an order to help me determine what to do, like "on this slide I will measure both minimum margin and invasiveness" and then I go to the next (slide) and do the same (…) (P1).
Another possible change in workflow was that the selection process for a measurement differed when the prototype was used. Without the prototype, one pathologist used separate notes and put the slides side by side to compare the measurements with each other to be able to choose what value to report. Putting slides side by side is another functionality made possible by the digital case review.
In the pen and paper case, one pathologist dealt with the overload of information when trying to figure out which tumor section was the largest, as seen in the scribbles shown in [Figure 4].
|
When the prototype was used, the pathologists replaced this way of taking notes and made the decision solely with the prototype's help, as in this field note:
(…) If you have measured invasive focus on these slides, you can just revisit the measurements (with annotation linking) and then choose which one to report instead of choosing a measurement and then re-evaluating it with the next measurement. This is better since you can perform the measurements and then evaluate and determine which one to actually report (P1).
The need of actively categorizing and filtering data was reduced using the prototype. The pathologists expressed that all values were displayed in a manner that was easy to overview. The structure made the pathologist feel confident with all collected data during image review, as illustrated by this field note:
I think it is very good that everything just ends up in the report to avoid mixing up numbers, etc., (P2).
One need that could not be met by the existing prototype was the categorization of multiple tumors to be able to distinguish these from each other. This need could not be met by using the descriptive text fields that were available.
Overall, the prototype was perceived as positive by both pathologists. They thought that the prototype would be useful for assessing partial mastectomy cases and that a similar approach could work for other types of cancer as well. Both pathologists also thought that the prototype could be useful in their everyday work and possibly improve their efficiency. Both pathologists expressed that the prototype would decrease time spent on image review, especially in cases with many measurements and variables to document and report.
Conclusions
This study is an example of the design process that takes place before a typical medical evaluation takes place, and highlights the different aspects that need to be considered when designing software functionality. Throughout the process, we generated different design principles and an understanding of the problem domain reporting in a digital pathology environment. One of the important findings in the results is that the report generation should be seen as process where the pathologists has a need to collect, sort, and display different types of measurements, and that it is possible to build a tool to aid the pathologists in this process. The use of the tool had an immediate effect on the personal working routines since it became possible to report multiple variables at the same time with improved convenience. A possible explanation for this effect could be that the pathologists' working memory was relieved, even though this hypothesis has not been evaluated in this study. Offloading working memory would be desirable, as it could have a positive effect on the quality of the review, especially for large cases that require a large amount of measurements.
The experienced convenience of the evaluated system depends heavily on a tight integration between the review of the slides and the way that reporting is performed. With a conventional microscope, it is today impossible to transfer measurements into structured data automatically. Instead, pen and paper are used to store intermediate values, and the final values are selected and organized during the dictation stage of the review. Pen and paper are flexible tools that humans can adapt to different situations, and it is therefore a challenge to create digital systems with the same flexibility. This study shows that at least for a narrow diagnostic scope, it is possible to reach sufficient flexibility. It should be noted, however, that a system appropriate for use within all domains of pathology is a much greater challenge to develop. We argue that to create a generic system, future work needs to be investigated whether it is possible to identify data operations that are relevant for all pathology domains, similar to what have been developed within the information visualization domain.[13] For example, we could reuse the design pattern to collect measurements under a heading, select those that should be reported, and filter out the rest, for many different types of measurements, not only length measurements as in this prototype. On the other hand, it is probably more difficult to generalize automatic calculation functionalities since these are more specific to the different grading protocols within different domains. In radiology, however, a project has generated a publicly available database where the reports for different radiology domains are published and the automatic calculation functionalities are defined within the templates.[14]
Another important aspect of the reporting is the collaboration. The main purpose of the final report is to enable efficient communication between the referring physician and the pathologist, very similar to the boundary object model.[15] Boundary objects are defined as objects that act as an interface between several communities of practice and satisfy the informational requirements of each of them. These types of communication objects also have a tendency to change appearance over time through a negotiation process, where the communities of practice agree on the objects' content and function. For example, if the pathologist agrees with the referring physician to always make an addition to the report template when something extraordinary is found in a case, all the other reports can become shorter, since those headings can be removed from the standard template. A structured reporting system needs to take these aspects into consideration, otherwise, the users will feel that the system hampers the communication with the referring physician and hampers the naturally occurring dynamic change that is an effect of the negotiation process.
This study was limited in scope and used a very limited number of participants. It is, therefore, not possible to draw any strong conclusions about whether most pathologists would prefer the type of reporting represented by the prototype, or that those reporting principles are better than other means. The aim was instead to generate hypotheses and a basic understanding of the problem area, and the results should therefore be treated as such. The findings are promising and indicate benefits within reach for future development along the proposed lines. Specifically, it would be interesting to develop this track to generalize the prototype design in this study to more disciplines within pathology and to evaluate the performance by comparing it with other means of reporting.
Acknowledgements
We would like to thank the participants of this study. We would also like to thank Anna Bodén, Gordan Maras, Sten Thorstenson, and Olle Westman, for their advice during the development of the prototype.
Financial support and sponsorship
This work was supported by VINNOVA (2014-04257) and the Swedish Research Council (2011-4138).
Conflicts of interest
All authors are employed by Sectra AB.
References
- ↑ Randell, R.; Ruddle, R.A.; Thomas, R.; Treanor, D. (2012). "Diagnosis at the microscope: A workplace study of histopathology". Cognition, Technology & Work 14 (14): 319–335. doi:10.1007/s10111-011-0182-7.
- ↑ Randell, R.; Ruddle, R.A.; Quirke, P.; Rhomas, R.G.; Treanor, D. (2012). "Working at the microscope: analysis of the activities involved in diagnostic pathology". Histopathology 60 (3): 504–510. doi:10.1111/j.1365-2559.2011.04090.x. PMID 22176210.
- ↑ Mossanen, M.; True, L.D.; Wright, J.L. et al. (2014). "Surgical pathology and the patient: A systematic review evaluating the primary audience of pathology reports". Human Pathology 45 (11): 2192-2201. doi:10.1016/j.humpath.2014.07.008. PMID 25149550.
- ↑ Casati, B.; Bjugn, R. (2012). "Structured electronic template for histopathology reporting on colorectal carcinoma resections: Five-year follow-up shows sustainable long-term quality improvement". Archives of Pathology & Laboratory Medicine 136 (6): 652–6. doi:10.5858/arpa.2011-0370-OA. PMID 22646273.
- ↑ Bjugn, R.; Casati, B.; Norstein, J. (2008). "Structured electronic template for histopathology reports on colorectal carcinomas: A joint project by the Cancer Registry of Norway and the Norwegian Society for Pathology". Human Pathology 39 (3): 359-67. doi:10.1016/j.humpath.2007.06.019. PMID 18187180.
- ↑ Weiss, D.L.; Langlotz, C.P. (2008). "Structured reporting: Patient care enhancement or productivity nightmare?". Radiology 249 (3): 739-47. doi:10.1148/radiol.2493080988. PMID 19011178.
- ↑ Farahani, N.; Parwani, A.V.; Pantanowitz, L. (2015). "Whole slide imaging in pathology: Advantages, limitations, and emerging perspectives". Pathology and Laboratory Medicine International 2015 (7): 23–33. doi:10.2147/PLMI.S59826.
- ↑ Krupinski, E.A. (2010). "Optimizing the pathology workstation "cockpit": Challenges and solutions". Journal of Pathology Informatics 1: 19. doi:10.4103/2153-3539.70708. PMC PMC2956171. PMID 21031008. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2956171.
- ↑ Fine, J.L. (2014). "21(st) century workflow: A proposal". Journal of Pathology Informatics 5 (1): 44. doi:10.4103/2153-3539.145733. PMC PMC4260324. PMID 25535592. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4260324.
- ↑ Hipp, J.; Flotte, T.; Monaco, J. et al. (2011). "Computer aided diagnostic tools aim to empower rather than replace pathologists: Lessons learned from computational chess". Journal of Pathology Informatics 2: 25. doi:10.4103/2153-3539.82050. PMC PMC3132993. PMID 21773056. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3132993.
- ↑ Obrenović, Z. (2011). "Design-based research: What we learn when we engage in design of interactive systems". Interactions 18 (5): 56–59. doi:10.1145/2008176.2008189.
- ↑ Cervin, I. (3 March 2015). "Reporting in digital pathology: Increasing efficiency and accuracy using structured reporting". Linköping University. http://www.diva-portal.org/smash/record.jsf?pid=diva2%3A792072&dswid=-3555.
- ↑ Shneiderman, B. (1996). "The eyes have it: A task by data type taxonomy for information visualizations". Proceedings of the IEEE Symposium on Visual Languages, 1996: 336-343. doi:10.1109/VL.1996.545307.
- ↑ Kahn, C.E. (2014). "Incorporating intelligence into structured radiology reports". SPIE Proceedings 9039. doi:10.1117/12.2043912.
- ↑ Lee, C.P. (2007). "Boundary negotiating artifacts: Unbinding the routine of boundary objects and embracing chaos in collaborative work". Computer Supported Cooperative Work 16 (3): 307–339. doi:10.1007/s10606-007-9044-5.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added.