Journal:MaPSeq, a service-oriented architecture for genomics research within an academic biomedical research institution
| Full article title | MaPSeq, a service-oriented architecture for genomics research within an academic biomedical research institution |
|---|---|
| Journal | Informatics |
| Author(s) | Reilly, J.; Ahalt, S.; McGee, J.; Owen, P.; Schmitt, C.; Wilhelmsen, K. |
| Author affiliation(s) | University of North Carolina at Chapel Hill |
| Primary contact | Phone: +1 919-445-9619 (Wilhelmsen) |
| Editors | Bryant, A. |
| Year published | 2015 |
| Volume and issue | 2 (3) |
| Page(s) | 20–30 |
| DOI | 10.3390/informatics2030020 |
| ISSN | 2227-9709 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | http://www.mdpi.com/2227-9709/2/3/20/htm |
| Download | http://www.mdpi.com/2227-9709/2/3/20/pdf (PDF) |
Abstract
Genomics research presents technical, computational, and analytical challenges that are well recognized. Less recognized are the complex sociological, psychological, cultural, and political challenges that arise when genomics research takes place within a large, decentralized academic institution. In this paper, we describe a Service-Oriented Architecture (SOA) — MaPSeq — that was conceptualized and designed to meet the diverse and evolving computational workflow needs of genomics researchers at our large, hospital-affiliated, academic research institution. We present the institutional challenges that motivated the design of MaPSeq before describing the architecture and functionality of MaPSeq. We then discuss SOA solutions and conclude that approaches such as MaPSeq enable efficient and effective computational workflow execution for genomics research and for any type of academic biomedical research that requires complex, computationally-intense workflows.
Keywords: service-oriented architecture; genomics; massively parallel sequencing; computational workflow; academic biomedical research; decentralized organization; distributed decision-making
Introduction
Genomics research presents well-recognized technical, computational, and analytical challenges.[1][2][3][4] For example, while the technology for massively parallel genomic sequencing has progressed to the point where large amounts of data can be generated at a rapid pace and for a reasonable cost, the analytical burden presented by this massive amount of data can quickly overwhelm the genomic analyst. Indeed, the analysis and interpretation of genetic findings is generally considered the rate-limiting step in the translation of genomic sequencing data into clinical practice and patient care.[4]
Less recognized challenges to research in genomics and any biomedical field are the sociological, psychological, cultural, and political barriers, many of which arise from the organizational structure within which the research takes place. Indeed, research organizations tend to fall somewhere on a continuum between completely centralized and completely decentralized.[5][6][7][8] Each of these extremes has advantages and disadvantages. Centralized organizations traditionally function within a simple organizational design, with singular decision-making, top-level operational control, a consolidated budget, strong/clear communication channels, uniform culture and politics, and a high degree of efficiency, but at the expense of flexibility. Decentralized organizations, in contrast, generally operate within a complex organizational design, with distributed decision-making, local operational control, regionalized budgets, numerous weak or broken communication channels, inconsistent (and sometimes conflicting) culture and politics, and a high degree of flexibility, but at the expense of efficiency. The conceptualization, design, development, and implementation of information technology (IT) solutions for research in genomics and any biomedical field must therefore involve careful consideration of not only the needs of the user base, but also the organizational structure within which the research takes place.
Herein, we present a Service-Oriented Architecture (SOA) application — termed MaPSeq — that was conceptualized and designed to address the organizational challenges of computation-intensive biomedical research within a decentralized academic institution. In this article, we first describe the challenges that contributed to the conceptualization and design of MaPSeq. We then provide an overview of the technical architecture and capabilities of MaPSeq. Finally, we provide a discussion of service-oriented solutions such as MaPSeq.
Challenges driving the conceptualization and SOA design of MaPSeq
The design of MaPSeq was motivated by challenges that arose during the implementation of a genomic sequencing project titled “North Carolina Clinical Genomic Evaluation by NextGen Exome Sequencing” (NCGENES). This project, which is funded by the National Human Genome Resource Institute, aims to conduct whole exome sequencing of 500 patient samples drawn from multiple disease categories. NCGENES is a complex project, with both research and clinical arms. Soon after the project was initiated, the research and clinical teams realized that there were numerous barriers and roadblocks that needed to be overcome in order to achieve the analytical goals of the project. (See Table 1 for overview.)
| ||||||||||||||||||||
Challenge 1
Academic institutions face the challenge of balancing the needs of large, funded, research projects that typically support the development of an informatics infrastructure with the needs of smaller, often unfunded, research projects that cannot afford significant development costs. Furthermore, few research projects are sufficiently funded to support future development needs. Our institution faced these challenges when trying to balance the needs of the NCGENES investigative team with those of other investigative teams and anticipate future needs. The scale, general applicability, and complexity of massively parallel sequencing favored the development of an SOA approach to support both current and future needs related to genomic and non-genomic computationally-intense serial workflows.
Challenge 2
As is typical for an academic institution, our genomics infrastructure developed in an ad hoc manner, with multiple investigative teams working independently across the university campus. The result was a burgeoning, uncoordinated cluster of distributed compute resources. Compounding this challenge were the numerous network idiosyncrasies that prevented administrators within one network from accessing compute resources within a different network; thus, access privileges to campus compute resources were determined locally and required on-site (rather than remote) access.
Challenge 3
Decision-making at large academic institutions tends to be decentralized, with numerous decision makers enforcing different (and often conflicting) policies and procedures. This organizational structure inevitably leads to political and cultural conflicts and resistance to change, particularly when “external” IT teams attempt to change the processes in place among “central” investigative teams. Political and cultural resistance to the NCGENES project was encountered early on as the investigative team identified many barriers to the automation of human user-controlled workflow processes. While the existing human user-run workflows met the needs of small genomic sequencing projects and user groups, these workflows were inefficient for the computationally-demanding, whole-exome sequencing needs of NCGENES. Moreover, the use of a human contact as the point of access to an existing workflow created a roadblock to the execution of NCGENES, reduced the efficiency of genomic analysis, and threatened the security of sensitive patient data.
Existing solutions
Numerous Workflow Management Systems and workflow pipelines for genomic analysis exist, including COSMOS[9], Ergatis[10], i2b2[11], LONI[12], NG6[13], NGSANE[14], Orione[15], RUbioSeq[16], SeqInCloud[17], STATegra EMS[18], TREVA[19], and Pegasus.[20] Our team evaluated each of these systems for their ability to overcome the challenges described above. We found that existing solutions could address some, but not all, of the roadblocks and barriers that were hindering progress on the NCGENES project and that a new solution was needed. While all of the existing workflow systems and pipelines have proven to be effective, each has limitations.[21] MaPSeq is not unique in this regard, but it is responsive to the key features of a decentralized research organization. Specifically, as an SOA, MaPSeq allows for integration with multiple clients and distributed systems, whether local, open source, or commercial, and provides tailored, reusable, automated service solutions that address the varying and evolving needs and preferences of decentralized decision-makers. MaPSeq is scalable and can support both small- and large-scale projects and thus is responsive to the computational needs of all investigators. MaPSeq is efficient and allows for seamless, opportunistic use of distributed compute resources. Finally, the service-oriented, automated approach requires little coordination or communication among individual user groups and thus avoids local nuances in politics and culture.
MaPSeq technical architecture and capabilities
Overview of MaPSeq architecture
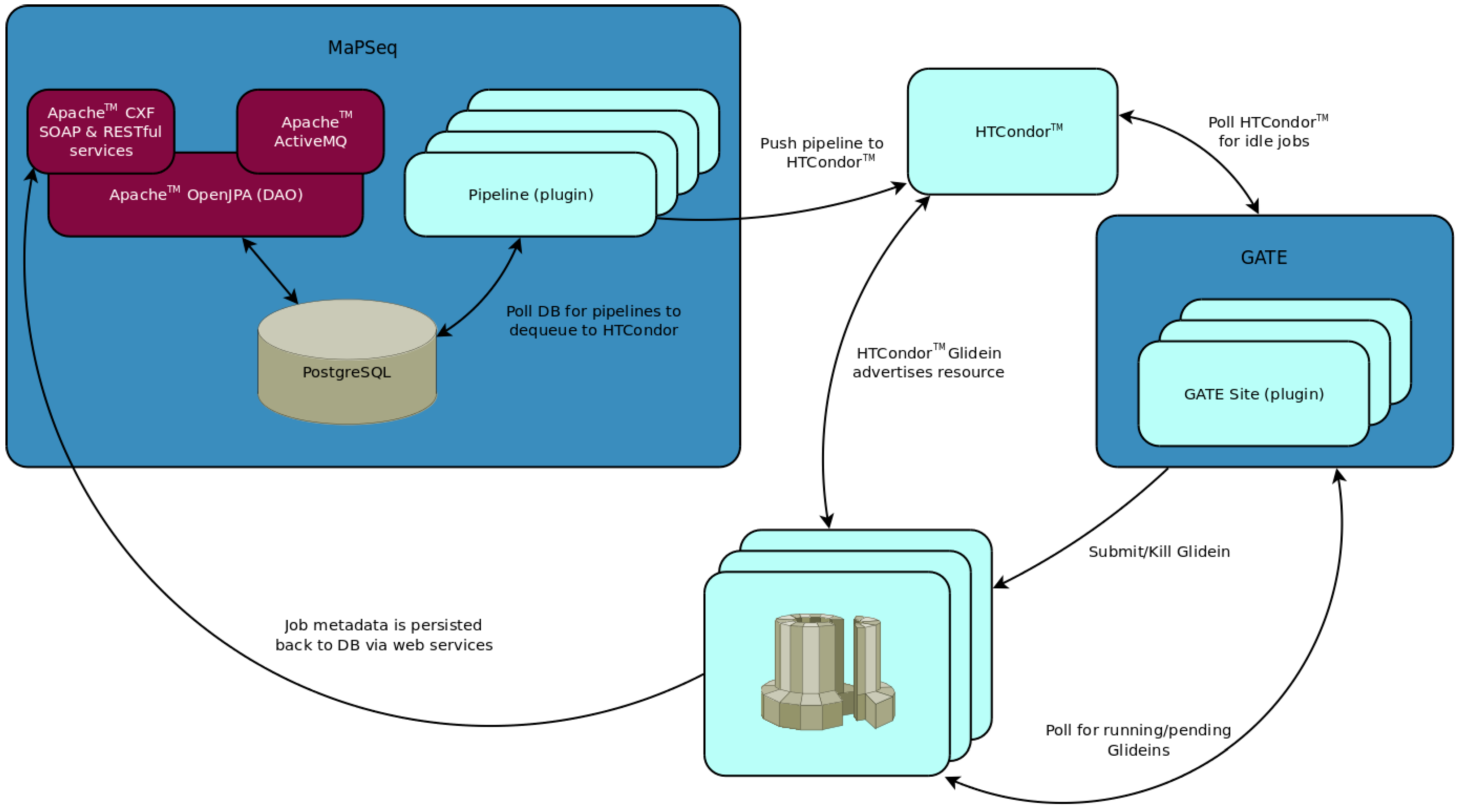
MaPSeq was designed as an open source, plugin-based SOA solution[22][23][24] that provides modifiable services to make opportunistic use of multiple institutional and cloud-based compute resources in order to efficiently complete the multitude of steps involved in the analysis of large-scale, genomic sequencing data (see Figure 1). The plugin framework of MaPSeq is based on the Open Services Gateway initiative (OSGi). This framework was chosen because of its modular agile architecture and the ability to remotely manage workflow pipelines in an on-demand manner and within a sandboxed environment. Moreover, the investigative team had relevant prior experience with the Open Science Grid Engagement Program, which aims to facilitate collaborative research through advanced distributed computing technologies.
MaPSeq and, its sister technology, the Grid Access Triage Engine (GATE), are built on top of ApacheTM Karaf, which is an OSGi-based lightweight container for application deployment. MapSeq works together with GATE to provide extensible capabilities for the analysis of genomic sequencing data, including: pipeline execution and management; meta-scheduling of workflow jobs; opportunistic compute-node utilization and management; secure messaging and data transfer; and client access via web services.
|
MaPSeq pipelines
MaPSeq pipelines (Figure 1) are OSGi-based plugins comprised of a number of bundles and/or services. At a minimum, a MaPSeq pipeline consists of: (1) a Java Message Service destination that exposes a mechanism whereby a user can trigger a pipeline; (2) a workflow designed as a Directed Acyclic Graph (DAG) and consisting of a collection of programmatic tasks; (3) an executor that dequeues the workflows at a customizable frequency (e.g., two workflows every five minutes, ten workflows every three minutes, etc.); and (4) a metadata file that describes all of the aforementioned features and tracks their status. Complex pipelines can be broken into numerous smaller sub-pipelines to enable symbolic check-pointing or fault tolerance. For example, a genomic analysis pipeline can be logically split into two sub-pipelines: an alignment sub-pipeline and a variant calling sub-pipeline. This approach enables a researcher to, for example, modify a step in the variant calling sub-pipeline and re-run that sub-pipeline without the need to re-run the alignment sub-pipeline, thereby reducing the runtime burden. Additionally, this approach allows the sub-pipelines to be reused in other pipelines, thus fostering software re-usability. Of note, all pipelines are project-specific and defined by the needs of the project and research team such that pipeline development is tailored to a specific application.
HTCondor
HTCondor (Figure 1) serves as a central manager and provides meta-scheduling for MaPSeq via the DAG Manager (DAGMan). MaPSeq workflows are comprised of numerous modules that form the vertices of a DAG. The DAGs can be exported for submission to HTCondor using DAGMan. MaPSeq provides a suite of modules that wrap third-party libraries (e.g., GATK, Picard, etc.) for execution on the grid and that include a number of lifecycle events. These lifecycle events check for valid inputs and outputs, successful execution, and provenance of job metadata, thus ensuring consistency and rapid detection of errors. HTCondor manages serial execution of MaPSeq modules, as well as job-to-machine resource negotiation or “matchmaking”. The matchmaking process identifies job requirements (e.g., four cores and 4 GB memory required), as defined by the job metadata, and pairs those requirements with available machine attributes (e.g., eight cores and 32 GB memory available). After a MaPSeq module is executed, that module, or job wrapper, persists the job metadata over web services into a PostgreSQL database. HTCondor Glideins are used to provision compute resources for the execution of jobs, as described below.
GATE
GATE (Figure 1) is a homegrown OSGi-based system that serves as a sister technology for MaPSeq. Whereas MaPSeq uses plugins to execute workflow pipelines, GATE uses plugins to access compute resources. GATE continuously monitors a local HTCondor instance for idle jobs and profiles compute resources for availability. If an idle job is detected, then GATE uses plugins to submit an HTCondor Glidein to the most appropriate compute resource, which then joins the local HTCondor pool. GATE defers matchmaking to the HTCondor Negotiator, which uses daemons to perform the matchmaking. GATE grows and shrinks the number of Glideins by assessing the number of running and idle local jobs against the number of running and idle Glidein jobs on the compute resource grid. After a Glidein is activated, it registers back to the HTCondor Central Manager as an available resource. This approach enables jobs to be both site-specific and site-agnostic.
Security, interfaces, and administration
Of significance, both MaPSeq and GATE use Secure SHell (SSH) technology, running with daemons, for authentication and data transfer. This level of security is particularly important for applications such as genomics that involve the movement of sensitive patient data.
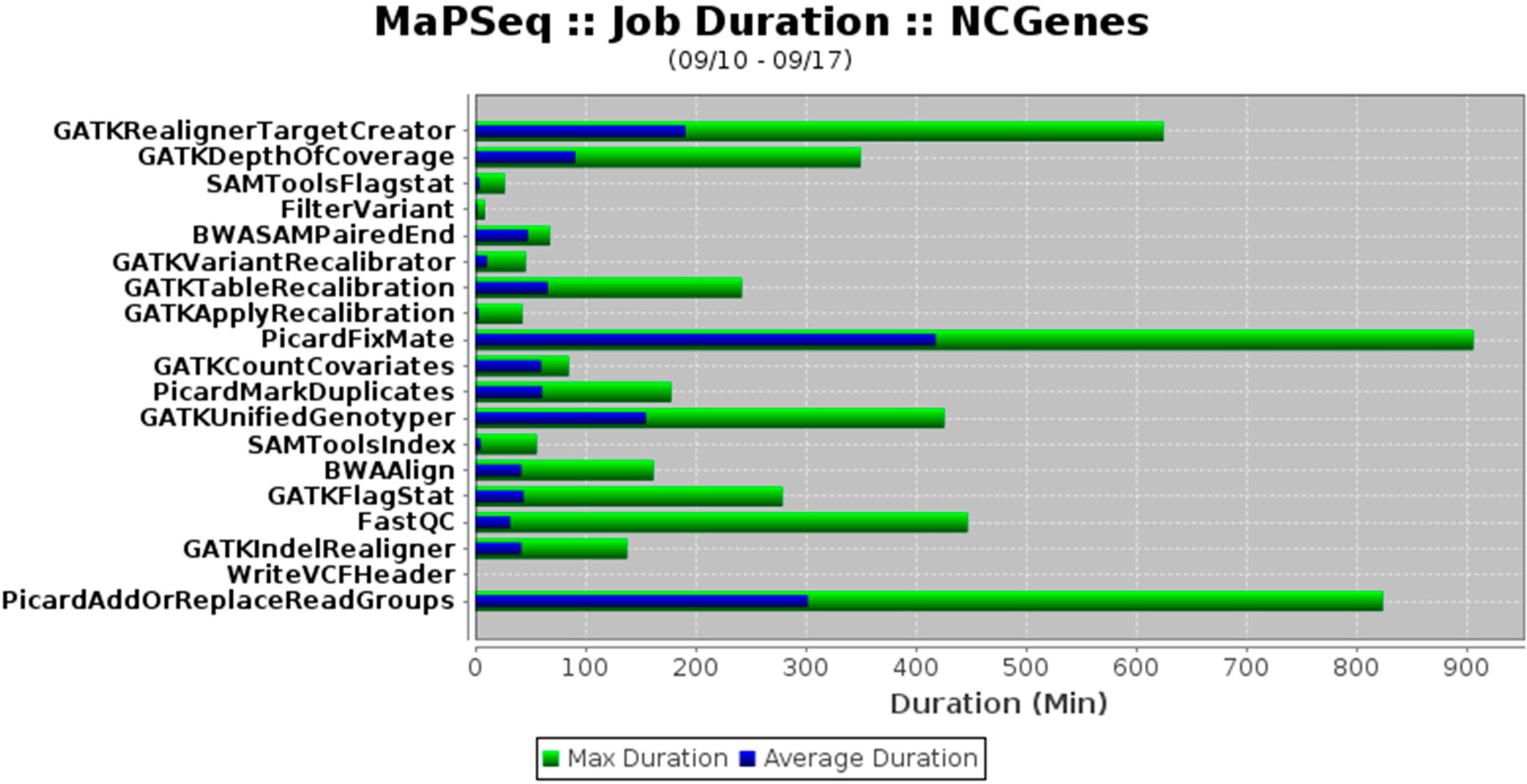
Clients can interface with MaPSeq using Apache™ CXF (Figure 1), which is an industry-standard web service. Both Simple Object Access Protocol (SOAP) and Representational State Transfer (RESTful) services are supported by Apache CXF. Pipeline invocations are triggered via a JavaScript Object Notation (JSON)-formatted message to an ApacheTM ActiveMQ destination. The JSON message contains the mapping between a MaPSeq-managed sample file instance and a workflow run instance. A pipeline-specific “message listener” then determines if the message is legitimate for subsequent processing. For genomic sequencing data, this process may involve verification that an object layer in the data file specifies that the data file contains raw sequencing data and sufficient metadata. A rich set of MaPSeq reports can be generated and sent to a client via email, for review and detection of potential problems (see example in Figure 2).
Apache Karaf is unique among containers in that it embeds an SSH daemon to enable a client to administratively manage pipeline deployment within a sandboxed environment. MaPSeq pipelines can be added, removed, or altered without having to stop the container, thereby provisioning a continuous, uninterrupted environment to execute new pipelines while existing pipelines are running. This accessibility allows for a pipeline developer to independently iterate on pipeline improvements.
|
Discussion
Genomics research within an academic environment presents numerous challenges. In addition to the computational and technical challenges inherent in genomics research[1][2][3][4], there are complex sociological, psychological, cultural, and political challenges that affect operations within academic institutions and indeed many other types of organizations.[25][26][27][28][29] Moreover, academic biomedical research institutions tend to be decentralized in their organizational structure. Whereas centralized organizations tend to function within a simple organizational design, with singular decision-making, top-level operational control, a consolidated budget, strong/clear communication channels, uniform culture and politics, and a high degree of operational efficiency, decentralized organizations, in contrast, operate within a complex organizational design, with distributed decision-making, localized operations and budgets, weak communication channels, nuances in culture and politics across academic units, and minimal operational efficiency.[5][6][7][8]
MaPSeq provides a reusable, service-oriented solution that addresses the diverse and evolving computational needs of decentralized decision-makers and scales to support both small- and large-scale projects. The automated approach requires little coordination or communication among individual user groups and thus avoids human roadblocks that may otherwise decrease efficiency. By leveraging the OSGi framework and Apache Karaf, MaPSeq allows for quick development iterations on MaPSeq pipeline plugins; pipelines can be created, altered, deployed, triggered, and removed without having to stop and restart the container. Finally, the use of HTCondor as a meta-scheduler and the addition of GATE as a sister technology allow MaPSeq to extend compute cluster capacity and make opportunistic use of distributed compute resources across the university campus.
In an environment of legacy systems, distributed and uncoordinated decision-making and compute resources, diverse and evolving user needs, and political and cultural resistance to change, centralized technical solutions will not promote efficient and effective biomedical research. SOA solutions provide the flexibility, scalability, extensibility, accessibility, interoperability, generalizability, achievability, and functionality required to attain efficient and effective, transformative biomedical research within a decentralized organization.
Limitations
Like any scientific workflow pipeline, MaPSeq is not without limitations.[21] First, while the underlying technology is open source and freely available, there is a considerable learning curve involved in implementation of the technology. Second, GATE is a homegrown solution and requires institution-specific adaptation before it can be adopted for use. Third, the MaPSeq solution must be continuously assessed against the evolving needs of relevant stakeholders, including users, patients, investigators, institutional administrators, and policy makers.
Conclusions
SOA solutions such as MaPSeq are well suited to overcome the many challenges to biomedical research that are inherent in a decentralized academic institution. MaPSeq has transformed genomics research at our institution and currently supports several large genomics research projects, as well as a few small ones. While MaPSeq was originally termed as an acronym for “Massively Parallel Sequencing” and designed to support genomics research, we note that the general architecture and approach can be adapted for other complex or computationally-intense workflows.
Finally, we note that MaPSeq (version 5.0) is available through a University of North Carolina Open Source Public License (version 1.1, ©2004). The only prerequisites are Java 1.7+, Apache™ Maven 3, and a network connection (full technical specifications and installation/operational instructions can be found at[30], with an accompanying RENCI technical report at reference[31]).
Acknowledgements
This project was conceptualized and implemented by RENCI and the UNC High-Throughput Sequencing Facility, in collaboration with Information Technology Services Research Computing and the Lineberger Comprehensive Cancer Center at the University of North Carolina at Chapel Hill and with funding from the National Institutes of Health (1R01-DA030976-01, 1U01-HG006487-01, 5UL1-RR025747-03, 1U19-HD077632-01, and 1U01-HG007437-01). The authors acknowledge the contributions of Corbin Jones, Associate Professor in the Department of Biology, and Jeff Roach, Senior Scientific Research Associate for Research Computing, Information Technology Services, University of North Carolina at Chapel Hill, to the design and implementation of MaPSeq. Karamarie Fecho, provided writing support for this manuscript, and RENCI provided funding for that support.
Author contributions
Jason Reilly designed and implemented MaPSeq with assistance from Phillips Owen as a replacement of earlier work by Charles Schmitt and based on prior work by John McGee, Kirk Wilhelmsen oversaw the implementation of MapSeq. Stanley Ahalt provided general guidance and facilities support for the development and implementation of MaPSeq.
Conflicts of interest
The authors declare no conflict of interest.
References
- ↑ 1.0 1.1 Koboldt, D.C.; Ding, L.; Mardis, E.R.; Wilson, R.K. (2010). "Challenges of sequencing human genomes". Briefings in Bioinformatics 11 (5): 484-498. doi:10.1093/bib/bbq016. PMC PMC2980933. PMID 20519329. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2980933.
- ↑ 2.0 2.1 Kahn, S.D. (2011). "On the future of genomic data". Science 331 (6018): 728-729. doi:10.1126/science.1197891. PMID 21311016.
- ↑ 3.0 3.1 Green, R.C.; Rehm, H.L.; Kohane, I.S. (2014). "Chapter 9: Clinical genome sequencing". In Willard, H.F.; Ginsburg, G.S.. Genomic and Personalized Medicine (2nd ed.). Oxford, UK: Academic Press. pp. 102–122. ISBN 9780123822277.
- ↑ 4.0 4.1 4.2 Dewey, F.E.; Grove, M.E.; Pan, C. et al. (2014). "Clinical interpretation and implications of whole-genome sequencing". JAMA 311 (10): 1035-1045. doi:10.1001/jama.2014.1717. PMC PMC4119063. PMID 24618965. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4119063.
- ↑ 5.0 5.1 Orlikowski, W.J.; Barley, S.R. (2001). "Technology and institutions: What can research on information technology and research on organizations learn from each other?". MIS Quarterly 25 (2): 145-165. doi:10.2307/3250927. PMC PMC2980933. PMID 20519329. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2980933.
- ↑ 6.0 6.1 Heiden, S. (10 December 2007). "Centralization Versus Decentralization: A Closer Look at How to Blend Both". Chief Learning Officer. MediaTec Publishing, Inc. http://www.clomedia.com/articles/centralization_versus_decentralization_a_closer_look_at_how_to_blend_both. Retrieved 16 April 2015.
- ↑ 7.0 7.1 Jain, P. (15 February 2013). "To Centralize Analytics or Not, That is the Question". Forbes. Forbes.com, LLC. http://www.forbes.com/sites/piyankajain/2013/02/15/to-centralize-analytics-or-not/. Retrieved 16 April 2015.
- ↑ 8.0 8.1 Ingram, D. (2015). "Centralized Vs. Decentralized Organizational Design". Houston Chronicle. Hearst Newspapers, LLC. http://smallbusiness.chron.com/centralized-vs-decentralized-organizational-design-11476.html. Retrieved 13 July 2015.
- ↑ Gafni, E.; Luquette, L.J.; Lancasster, A.K. et al. (2014). "COSMOS: Python library for massively parallel workflows". Bioinformatics 30 (20): 2956-2958. doi:10.1093/bioinformatics/btu385. PMC PMC4184253. PMID 24982428. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4184253.
- ↑ Orvis, J.; Crabtree, J.; Galens, K. et al. (2010). "Ergatis: A web interface and scalable software system for bioinformatics workflows". Bioinformatics 26 (12): 1488-1492. doi:10.1093/bioinformatics/btq167. PMC PMC2881353. PMID 20413634. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2881353.
- ↑ Kohane, I.S.; Churchill, S.E.; Murphy, S.N. (2011). "A translational engine at the national scale: Informatics for integrating biology and the bedside". Journal of the American Medical Informatics Association 19 (2): 181-185. doi:10.1136/amiajnl-2011-000492. PMC PMC3277623. PMID 22081225. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3277623.
- ↑ Dinov, I.D.; Torri, F.; Macciardi, F. et al. (2011). "Applications of the pipeline environment for visual informatics and genomics computations". BMC Bioinformatics 12: 304. doi:10.1186/1471-2105-12-304. PMC PMC3199760. PMID 21791102. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3199760.
- ↑ Mariette, J.; Escudié, F.; Allias, N. et al. (2012). "Integrated next generation sequencing storage and processing environment". BMC Genomics 13: 462. doi:10.1186/1471-2164-13-462. PMC PMC3444930. PMID 22958229. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3444930.
- ↑ Buske, F.A.; French, H.J.; Smith, M.A. et al. (2014). "NGSANE: A lightweight production informatics framework for high-throuput data analysis". Bioinformatics 30 (10): 1471-1472. doi:10.1093/bioinformatics/btu036. PMC PMC4016703. PMID 24470576. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4016703.
- ↑ Cuccuru, G.; Orsini, M.; Pinna, A. et al. (2014). "Orione, a web-based framework for NGS analysis in microbiology". Bioinformatics 30 (13): 1928-1929. doi:10.1093/bioinformatics/btu135. PMC PMC4071203. PMID 24618473. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4071203.
- ↑ Rubio-Camarillo, M.; Gómex-López, G.; Fernández, J.M. et al. (2013). "RUbioSeq: A suite of parallelized pipelines to automate exome variation and bisulfite-seq analyses". Bioinformatics 29 (13): 1687-1689. doi:10.1093/bioinformatics/btt203. PMC PMC3694642. PMID 23630175. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3694642.
- ↑ Mohamed, N.M.; Lin, H.; Feng, W.C. (2013). "Accelerating Data-Intensive Genome Analysis in the Cloud" (PDF). Virginia Tech. http://synergy.cs.vt.edu/pubs/papers/nabeel-bicob13-genome-analysis-cloud.pdf. Retrieved 16 April 2015.
- ↑ De Diego, R.H.; Boix-Chova, N.; Gómez-Cabrero, D. et al. (2014). "STATegra EMS: An experiment management system for complex next-generation omics experiments". BMC Systems Biology 8 (Suppl 2): S9. doi:10.1186/1752-0509-8-S2-S9. PMC PMC4101697. PMID 25033091. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4101697.
- ↑ Li, J.; Doyle, M.A.; Saeed, I. et al. (2014). "Bioinformatics pipelines for targeted resequencing and whole-exome sequencing of human and mouse genomes: A virtual appliance approach for instant deployment". PLOS ONE 9 (4): e95217. doi:10.1371/journal.pone.0095217. PMC PMC3994043. PMID 24752294. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3994043.
- ↑ Deelman, E.; Vahi, K.; Juve, G. et al. (2015). "Pegasus: A workflow management system for science automation". Future Generation Computer Systems 46 (5): 17–35. doi:10.1016/j.future.2014.10.008.
- ↑ 21.0 21.1 Bromberg, Y. (2013). "Building a genome analysis pipeline to predict disease risk and prevent disease". Journal of Molecular Biology 425 (21): 3993–4005. doi:10.1016/j.jmb.2013.07.038. PMID 23928561.
- ↑ Sprott, D.; Wilkes, L. (January 2004). "Understanding Service-Oriented Architecture". Microsoft Developer Network. Microsoft. https://msdn.microsoft.com/en-us/library/aa480021.aspx. Retrieved 16 April 2015.
- ↑ CIO Staff (19 March 2007). "SOA Definition and Solutions". CIO. CXO Media, Inc. http://www.cio.com/article/2439274/service-oriented-architecture/soa-definition-and-solutions.html. Retrieved 16 April 2015.
- ↑ Bailey, M. (2008). "Principles of Service Oriented Architecture". SlidePlayer. SlidePlayer.com, Inc. http://slideplayer.com/slide/701834/. Retrieved 16 April 2015.
- ↑ Williams, R.; Edge, D. (1996). "The social shaping of technology". Research Policy 25 (6): 865–899. doi:10.1016/0048-7333(96)00885-2.
- ↑ Lorenzi, N.M.; Riley, R.T.; Blyth, A.J.C. et al. (1997). "Antecedents of the people and organizational aspects of medical informatics: Review of the literature". Journal of the American Medical Informatics Association 4 (62): 79-93. doi:10.1136/jamia.1997.0040079. PMC PMC61497. PMID 9067874. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC61497.
- ↑ Jasperson, J.S.; Sambamurthy, V.; Zmud, R.W. (1999). "Social influence and individual IT use: Unraveling the pathways of appropriation moves". Proceedings of the 20th International Conference on Information Systems: 113–118. http://aisel.aisnet.org/icis1999/10/.
- ↑ Sassen, S. (2002). "Towards a sociology of information technology". Current Sociology 50 (3): 365-388. doi:10.1177/0011392102050003005.
- ↑ Schmidt, J.; Lyle, D. (2005). Integration Competency Center: An Implementation Methodology. Redwood City, CA: Informatics Corp. pp. 153. ISBN 9780976916307.
- ↑ "Massively Parallel Sequencing". University of North Carolina at Chapel Hill. 2015. http://jdr0887.github.io/MaPSeq-API/index.html. Retrieved 13 July 2015.
- ↑ "TR-14-03 MaPSeq, A Computational and Analytical Workflow Manager for Downstream Genomic Sequencing". RENCI. 3 June 2014. http://renci.org/technical-reports/mapseq-computational-and-analytical-workflow-manager/. Retrieved 13 July 2015.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added.