Journal:Predicting biomedical metadata in CEDAR: A study of Gene Expression Omnibus (GEO)
| Full article title | Predicting biomedical metadata in CEDAR: A study of Gene Expression Omnibus (GEO) |
|---|---|
| Journal | Journal of Biomedical Informatics |
| Author(s) | Panahiazar, Maryam; Dumontier, Michel; Gevaert, Olivier |
| Author affiliation(s) | Stanford University |
| Primary contact | Email: olivier dot gevaert at stanford dot edu |
| Year published | 2017 |
| Volume and issue | 72(8) |
| Page(s) | 132–139 |
| DOI | 10.1016/j.jbi.2017.06.017 |
| ISSN | 1532-0464 |
| Distribution license | Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International |
| Website | http://datascience.codata.org/articles/10.5334/dsj-2017-001/ |
| Download | http://datascience.codata.org/articles/10.5334/dsj-2017-001/galley/620/download/ (PDF) |
Abstract
A crucial and limiting factor in data reuse is the lack of accurate, structured, and complete descriptions of data, known as metadata. Towards improving the quantity and quality of metadata, we propose a novel metadata prediction framework to learn associations from existing metadata that can be used to predict metadata values. We evaluate our framework in the context of experimental metadata from the Gene Expression Omnibus (GEO). We applied four rule mining algorithms to the most common structured metadata elements (sample type, molecular type, platform, label type and organism) from over 1.3 million GEO records. We examined the quality of well supported rules from each algorithm and visualized the dependencies among metadata elements. Finally, we evaluated the performance of the algorithms in terms of accuracy, precision, recall, and F-measure. We found that PART is the best algorithm outperforming Apriori, Predictive Apriori, and Decision Table.
All algorithms perform significantly better in predicting class values than the majority vote classifier. We found that the performance of the algorithms is related to the dimensionality of the GEO elements. The average performance of all algorithm increases due of the decreasing of dimensionality of the unique values of these elements (2697 platforms, 537 organisms, 454 labels, 9 molecules, and 5 types). Our work suggests that experimental metadata such as present in GEO can be accurately predicted using rule mining algorithms. Our work has implications for both prospective and retrospective augmentation of metadata quality, which are geared towards making data easier to find and reuse.
Graphical abstract
Keywords
Data mining; Prediction; Metadata; GEO; CEDAR
1. Introduction
Biomedical data is increasingly being viewed as a valuable commodity that can be mined for new insights beyond that for which it was created. Large community-focused databases such as the Gene Expression Omnibus (GEO)[1] or the database of Genotypes and Phenotypes (dbGAP)[2] offer a wealth of omics’ data that have been used in developing diagnostic, prognostic, and therapeutic models.[3][4] One crucial and limiting factor in the reuse of data lies in having access to accurate descriptions about the data – known as metadata. Community standards to describe an experiment (e.g. Minimum Information About a Microarray Experiment; MIAME[5]) are being widely promoted to highlight essential metadata, but creating good metadata can be challenging.[6][7]
Indeed, metadata is often of low quality, and many entries are absent, erroneous or inconsistent. The largest database of gene expression studies, the GEO microarray database, contains 50,000 studies, over 1.3 million samples, and is still growing [1]. Yet the description of these samples suffers from a lack of consistency and completeness. For example, a preliminary analysis revealed that are 32 different ways to specify the age in GEO (e.g. age, Age, Age years, age year). Yet, these metadata are essential for researchers to find and reuse datasets of interest. When metadata are incomplete or inaccurate, researchers will miss relevant hits while being forced to sift through irrelevant results - resulting in lower productivity and potentially weaker scientific analyses. These issues are often attributed to lack of appropriate supporting infrastructure.[8]
Metadata authoring applications such as ISA-Tools[9] or RightField[10] can be used to codify guidelines that specify multiple metadata elements and require users to use a set of controlled terms, such as terms from specified ontologies contained in the NCBO BioPortal.[11] Yet even with such tools, authoring good metadata is tedious and error-prone, and could benefit from more automation. The development of more effective platforms for metadata authoring and discovery is one of the goals of the Center for Expanded Data Annotation and Retrieval (CEDAR).[7][8]
In this study, we examine the utility of supervised machine learning to predict metadata from existing metadata. This will help metadata submitter during the submission process. Predicting metadata could be a guideline for template authors during the process of metadata definition. This facility will not only significantly facilitate the template definition task but also will make the resulting templates more comprehensive and reflective of the actual data. In CEDAR we also take advantage of emerging community-based standard templates for describing different kinds of biomedical datasets, and we investigate the use of computational techniques to help investigators to assemble templates and to fill in their values.[7]
Learning value sets from data will help ensure that template authors do not miss important value sets that appear frequently in the data. Thus, data submitters will be able to find the terms they need, hence improving the quality of the metadata.
We use the increasing amounts of structured metadata to learn from as the project progresses and learn value sets conditional on the experimental level metadata. This incorporation of structural knowledge into the learning technology will allow us to infer common metadata patterns and their value sets in the context of technology platform, organism, molecule, label or sample type. Our key goal is to facilitate as much of the metadata collection process as possible, by suggesting possible value sets for the fields based on available data. This process will limit the value options, will reduce the burden of entering metadata terms and will significantly shorten the time that is needed for investigators to enter metadata.
We found that experimental metadata such as present in GEO can be accurately predicted using rule mining algorithms. Our work has implications for both prospective and retrospective augmentation of metadata quality, which are geared towards making data easier to find and reuse.
2. Background
Supervised learning uses classification algorithms to learn from data and make predictions. The goal of supervised learning is to build a model of the distribution of class labels from instances.[12] The classifier can then assign class labels to instances in which the values of the predictor features are known, but the value of the class label is unknown. Numerous supervised classification techniques have been developed including decision trees, artificial neural networks, and statistical techniques such as bayesian networks.[12] Machine learning has been widely applied across domains including the biomedical domain[13], such as protein function prediction[14], clinical outcome prediction[15] and survival analysis.[16]
As we mentioned earlier, this study specifically is about metadata and association between them. Therefore, using machine learning will be helpful to mine the data, learn from the data, and find this association. In our study, we wanted to find the correlation between metadata elements and their values. Association rules are the main technique for data mining to find these correlations. Sharma et al., compared association rule mining algorithms (e.g. AIS and FP-Growth, and Apriori).[17] Each algorithm has advantages and disadvantages according to their comparison. For example, AIS requires multiple scanning of the database, only rules that have one item in right side can be generated, and too many candidate itemsets are generated. FP-Growth also has some disadvantages such as the resulting FP-Tree is not unique for the same logical database and it cannot be used in interactive mining system. Apriori is scanning the complete database multiple times but still, it is easy to implement. Predictive Apriori algorithm overcomes this disadvantage of the Apriori algorithm with scanning the beast n rules instead of scanning all rules. PART algorithm uses partial decision trees to generate the decision list that is shown in the output, but only this final list is what is used to make classifications and with that, we have better performance.
In previously published manuscript[18], we proposed a framework to predict structured metadata terms from unstructured metadata for improving quality and quantity of metadata, using the Gene Expression Omnibus (GEO) microarray database. Our framework consists of classifiers trained using term frequency-inverse document frequency (TF-IDF) features and a second approach based on topics modeled using a Latent Dirichlet Allocation model (LDA) to reduce the dimensionality of the unstructured data. Our results based on GEO database showed that structured metadata can be predicted with TF-IDF more accurate than LDA. And both TF-IDF and LDA are outperforming the majority vote baseline as well. Overall this is a promising approach for metadata prediction that is likely to be applicable to other datasets and has implications for researchers interested in biomedical metadata curation and metadata prediction. Considering that metadata is structured and unstructured in GEO and other resources, we decided to find the correlation between structured metadata. In this study, we found the correlation between selected structured metadata elements versus in previous work we predicted structure metadata from the free text. Structure metadata has a potential to be predicted and suggested to metadata template author or metadata submitter during the submission process based on each other.
Several studies have been done regarding GEO metadata prediction. For instance Buckberry et al.[19] presented a method for predicting the sex of samples in gene expression microarray datasets. They believe that the metadata associated with many publicly available expression microarray datasets often lacks sample sex information, therefore limiting the reuse of these data in new analyses or larger meta-analyses where the effect of sex is to be considered. The package called massiR provides a method for researchers to predict the sex of samples in microarray datasets. “This package implements unsupervised clustering methods to classify samples into male and female groups, providing an efficient way to identify or confirm the sex of samples in mammalian microarray datasets”.[19] As it is clear this study is just about particular field in GEO data and it is specialized to predict the sex of the samples.
In this study, we propose methods to predict structured metadata. This method is applicable to any structured metadata in biomedical field. We use association rule mining (ARM) algorithms due to their interpretability and good performance.[20] ARM is a method for discovering relations between variables in large databases.[21] ARM was defined by Agrawal in the early 90s in relation to a so called market basket analysis using APRIORI.[20] Since then, multiple studies have used this technique successfully to model data.[22] For example, ARM has been used to predict infection detection[23], to detect common risk factors in pediatric diseases[24], to understand the interaction between proteins[25], to discover frequent patterns in gene data[22], and to understand what drugs are co-prescribed with antacids [26]. To the best of our knowledge, ARM has not yet been applied for predicting experimental metadata.[26]
3. Objective
We hypothesized that there are strong correlations between metadata elements and their values that can be used to predict metadata. The goal of this study is to predict the metadata based on the correlation between them. For example, there is a correlation between platforms, organism, and type. For GPL570 as a platform and Homo Sapiens as an organism a possible type of the study is RNA. We used four algorithms: Apriori, Predictive Apriori, Decision Table and PART (see below). We used these algorithms to find the association between metadata elements and to predict the value of each element of interest. We then evaluated our approach using a standard cross-validation of experimental metadata from GEO, a primary repository of gene expression data.
4. Materials and methods
4.1. Metadata
Our work focused on GEO[1], a large and well known database of gene expression data which contains experimental metadata authored by the original data submitters. We used the “GEOmetadb” package[27] in R[28] to query and obtain the metadata for microarray experiments. GEOmetadb implements an SQLite database that stores all the metadata associated with all GEO data types including GEO samples (GSM), GEO platforms (GPL), GEO data series (GSE). GEO itself stores curated gene expression DataSets (GDS) that allows non-technical users to identify and visualize differentially expressed genes in a given study. However, GEO DataSet curation is not standardized across studies which preclude more powerful methods such as integrated meta-analysis across multiple experiments to find robust gene signatures. GDS have not been considered in this study.
The GEO database as of October 2015 contains 1,368,682 individual sample records in 50,000 studies or series. It includes 1.4 million samples now (June 2016), which is decreased to 1.2 million samples after removing elements that occur less than 250 times. A series is identified with a series id (i.e. GSExxxxx) and each series consist of one or more samples. A sample (identified with GSMxxxxx) describes the set of molecules that are being probed and references a platform (i.e. GPLxxxxx) used to representing the molecular data.[1] Each study is annotated with up to 32 metadata fields representing the conditions under which the sample was handled. There are 32 fields (16 for each channel of study including ch1 and ch2).
After discussion with the researchers in the field we considered five common structured elements for this study including (sample type, molecular type, platform, label type and organism (Table 1) from 16 elements (title, gsm, series-id, gpl, status, submission data, last-update-date, type, sources name, organism, characteristics, molecule, label, treatment protocols, extract-protocol, label -protocol). Other elements are date related (e.g. last-update-date) or they are considered as unstructured (e.g. title) metadata. Therefore, we removed free text and date related information. We also removed the studies with more than half missing value. We explained the prediction for unstructured metadata such as title of the study in our previous work. We define a structured element as a metadata element which contains a single concept, such as the organism from which the material was derived. More specifically, GEO metadata includes 5 sample types (e.g. RNA, genomic), 9 types of molecules that were extracted from the biological material (e.g., total RNA, cytoplasmic RNA), 12,431 different platforms (e.g., GPL13653 for Affymetrix GeneChip Rat Genome U34A Array), 1,641 compounds used to label the samples (e.g., biotin, Cy3) and 2434 organisms (e.g. mus musculus). We removed elements that occur less than 250 times to avoid the long tail, resulting in modeling 2697 platforms, 5 types, 537 organisms, 9 molecule, and 454 labels (Table 2). We also made sure we did not reduce the number of type and molecule with this set up threshold, which they were not that many to begin with.
| ||||||||||||||
| ||||||||||||||||||||||||||||
4.2. Association rule mining algorithms
In this section, we describe the four different Association Rule Mining Algorithms (ARM) algorithms including Apriori, Predictive Apriori, Decision Table and PART. These algorithms have been used to learn the rules and find the possible associations between five structural GEO elements and their values. We compared all four algorithms with the majority vote classifier representing the baseline model.
An association rule is an implication expression of the form X to Y, where X and Y are disjoint itemsets. The strength of an association rule can be measured in terms of its support and confidence. Support determines how often a rule is applicable to a given data set, while confidence determines how frequently items in Y appear in transactions that contain X.[17]
The Apriori algorithm identifies association rules by identifying frequently occurring item sets.[20] An item set is called frequent when its support is above a defined minimum support. An item set X of length L is frequent if and only if all subsets of X with length L - 1 are frequent. For every frequent item set T and every non-empty subset S of T, Apriori outputs a rule of the form S ⇒ (T - S) if and only if the confidence of that rule is above the user specified threshold. To run the algorithm some parameters had to be defined (e.g. T = 0: The metric type which has been used to rank the rules. (default = confidence); C = 0.9: The minimum confidence of a rule; D = 0.05: The delta by which the minimum support is decreased in each iteration; U = 1.0: Upper bound for minimum support; M = 0.1: The lower bound for the minimum support). Apriori is easy to implement, but it is computationally and memory intensive.
Predictive Apriori[29] is a variant of Apriori that searches for the best ’n’ rules using a support-based corrected confidence value. Since we just look at the best n rules is this algorithm, to run the algorithm we need to set the particular class attribute to predict as well (C = the class index for the chosen element to predict from 1 to 5) in each run. Predictive Apriori maximizes the accuracy and minimizes the number of searches as compared to Apriori. A rule is added if the expected predictive accuracy of the rule is among the ’n’ best and it is not subsued by a rule with at least the same expected predictive accuracy.[30]
A Decision Table[31] is a compact and easy to understand method to show the relationship between a series of conditions and resultant actions. It is based on a decision tree, where each node represents a feature and each branch represents a value that the node can assume. To run the algorithm some other parameters had to be defined (e.g. D = 1 to set the forward search and N = 5 which is the number of non-improving nodes to consider before terminating search). A Decision Table can be translated into a set of rules by creating a separate rule for each path from the root to a leaf in the tree constructing an optimal binary.
Finally, PART[32] is an algorithm that uses partial trees to generate near-optimal decision list. This list is what is used to make classifications. Once a partial tree has been build, a single rule is extracted from it. To run the PART algorithm considering previous parameters we also set minimum number of instances per leaf equal to M = 2. The difference between heuristics for PART and heuristics for Decision Table is that the latter evaluate the average quality of a number of disjointed sets (one for each value of the feature that is tested), while PART only evaluate the quality of the set of instances that is covered by the candidate rule.
4.3. Experimental setup and evaluation framework
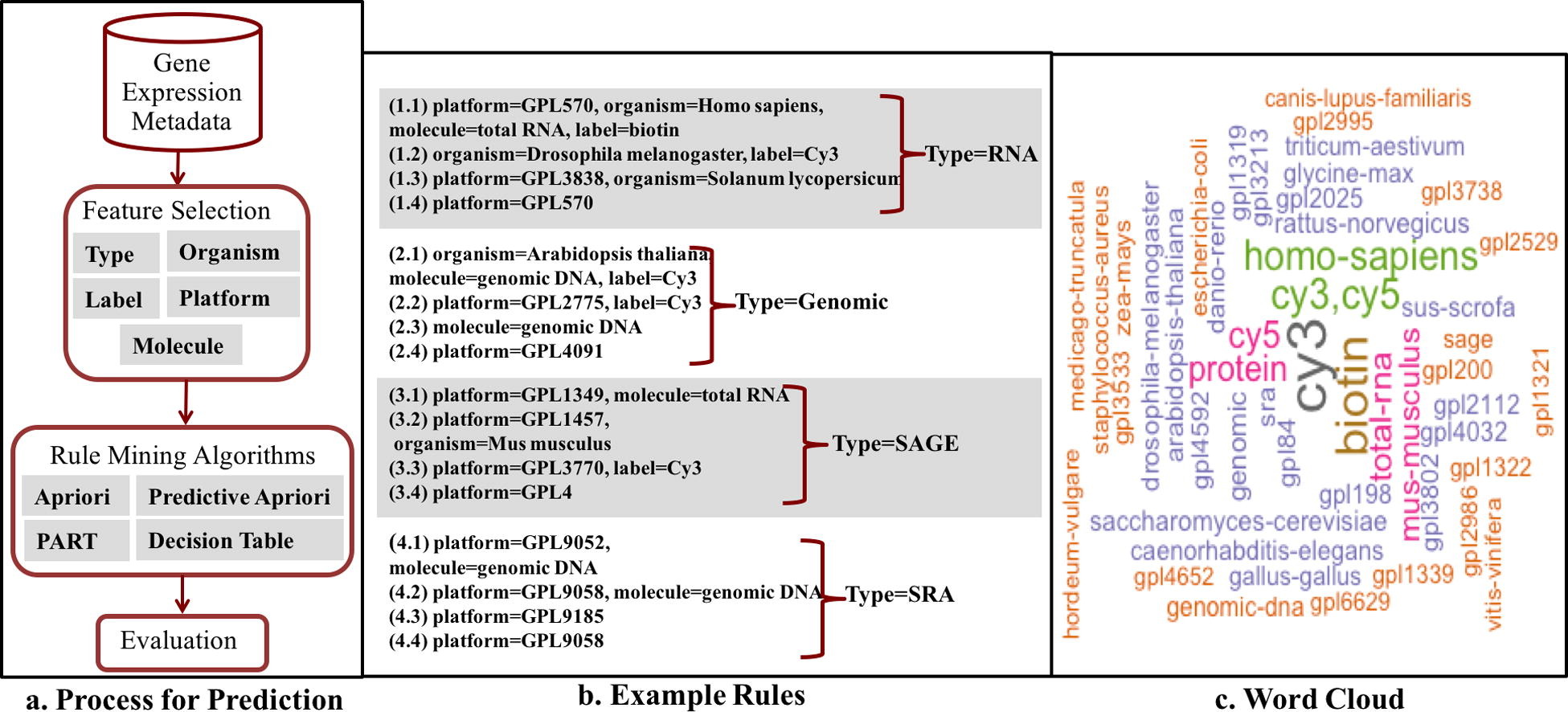
We used the four ARM algorithms to discover rules from our GEO dataset (Fig. 1). We predicted each feature based on the other features (e.g. ’type’ was predicted using molecule, label, platform, and organism). An example of a rule is: if organism = Homo Sapiens, molecule = total RNA then type = RNA. We performed 90:10 cross-validation in which we used 90% of the sample data for training and 10% for testing. Since the same sample can be used in another series, we partitioned the dataset by superseries such that samples that belong to the same study are either all in the training set or all in the test set. We assessed classifier performance based on the standard metrics for accuracy, precision, recall and F-measure.[33] The summary of the process of metadata prediction is shown in Fig. 1a. We then identified predictive features by counting the number of times a feature was selected as a feature in the model. We visualized the dependencies between all features as a network.
|
4.4. Results
In this section, we discuss rules discovered with each of the four ARM algorithms over the experimental metadata from the GEO database. We report on the performance of each algorithm, and discuss associations within the rulesets.
Over five thousand rules were generated from the analysis of the GEO database. We divided the rules into two kinds of rules: (1) complex rules having at least two elements in the antecedent, and (2) simple rules having only one element in the antecedent and one in the consequent. Fig. 1b. highlights rules to predict four metadata elements: RNA, Genomic, SAGE, and SRA. For example rule 1.1 is a complex rule to predict sample type using values from the other 4 features. This rule predicts RNA as type when the platform is GPL570 (i.e. the Affymetrix Human Genome U133 Plus 2.0 platform), the label is Biotin, the type of molecule that was extracted from the biological material is total RNA, and the sample was obtained from humans (Homo sapiens). In contrast, rule 3.4 is a simple rule that predicts the sample type as SAGE, when the platform used is GPL4. For the most common sample type, RNA, the generated rules have more variety with varying rule complexity (e.g. rule 1.1 with length 5 compared to rule 1.4, a simple rule). For the metadata element type, the value SRA is only predicted with the length of up to 3 (e.g. rules 4.1, 4.2). Next, Fig. 1c. provides insight into reoccurring values in rules generated by the PART algorithm. For instance, the label Cy3 is most frequently used.
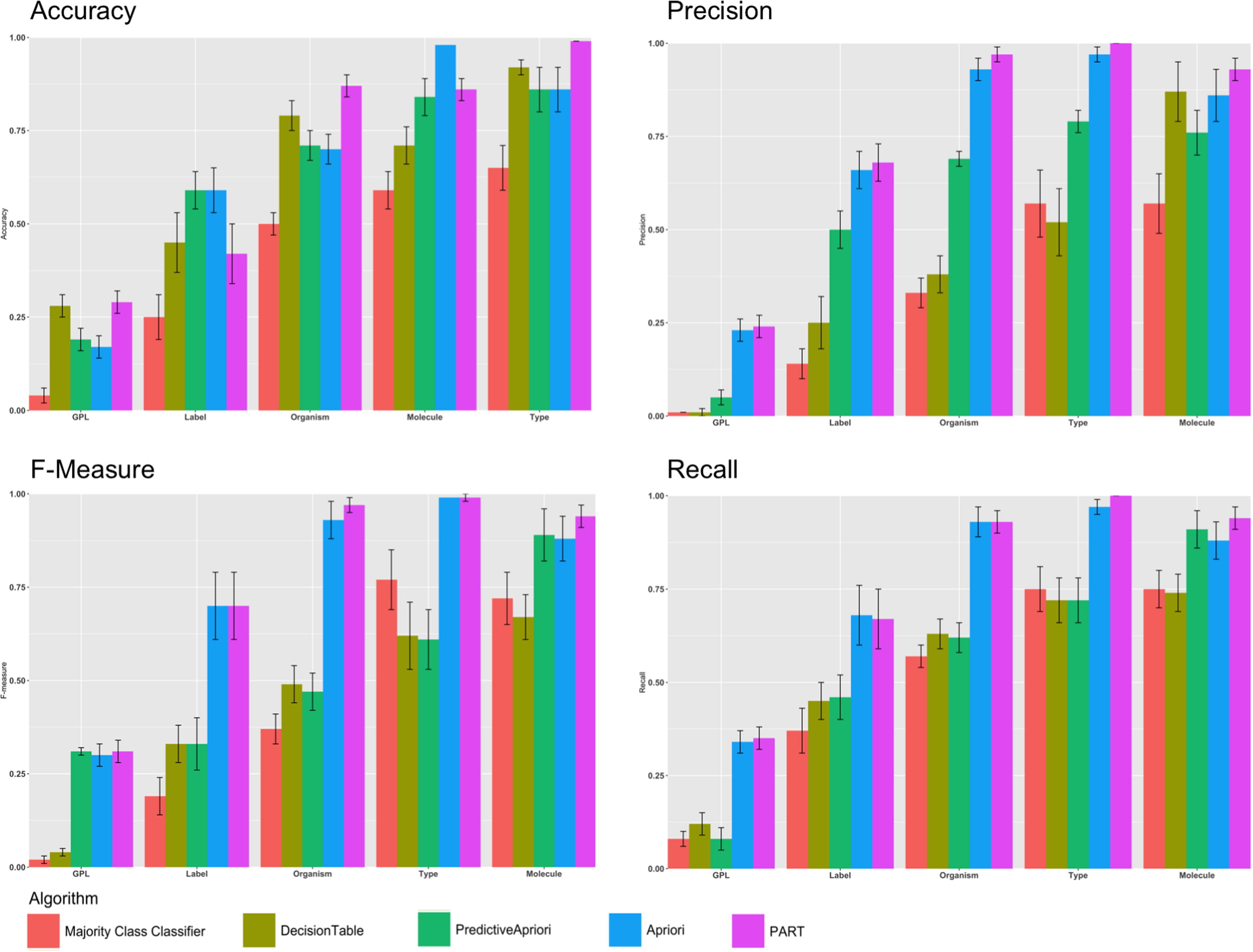
Next, we sought to understand how each of the four rule mining algorithms performed for each of the five selected features drawn from the GEO dataset. Fig. 2 shows the performance using F-measure, precision, recall and accuracy for each of the four algorithms and the majority vote baseline. Our results indicate that PART is the best classifier. Also, only PART and Decision Table consistently outperformed the majority vote classifier for predicting all features that we examined. PART and Decision Table outperformed Apriori and Predictive Apriori for Label, Organism, and Type. As shown in Fig. 2 for each performance measurement we considered the confidence interval. We calculated the confidence interval for 10 iterations for each algorithm. As an example, Table S2 in supplementary materials shows the details regarding the calculation of traditional confidence interval for all algorithms.
|
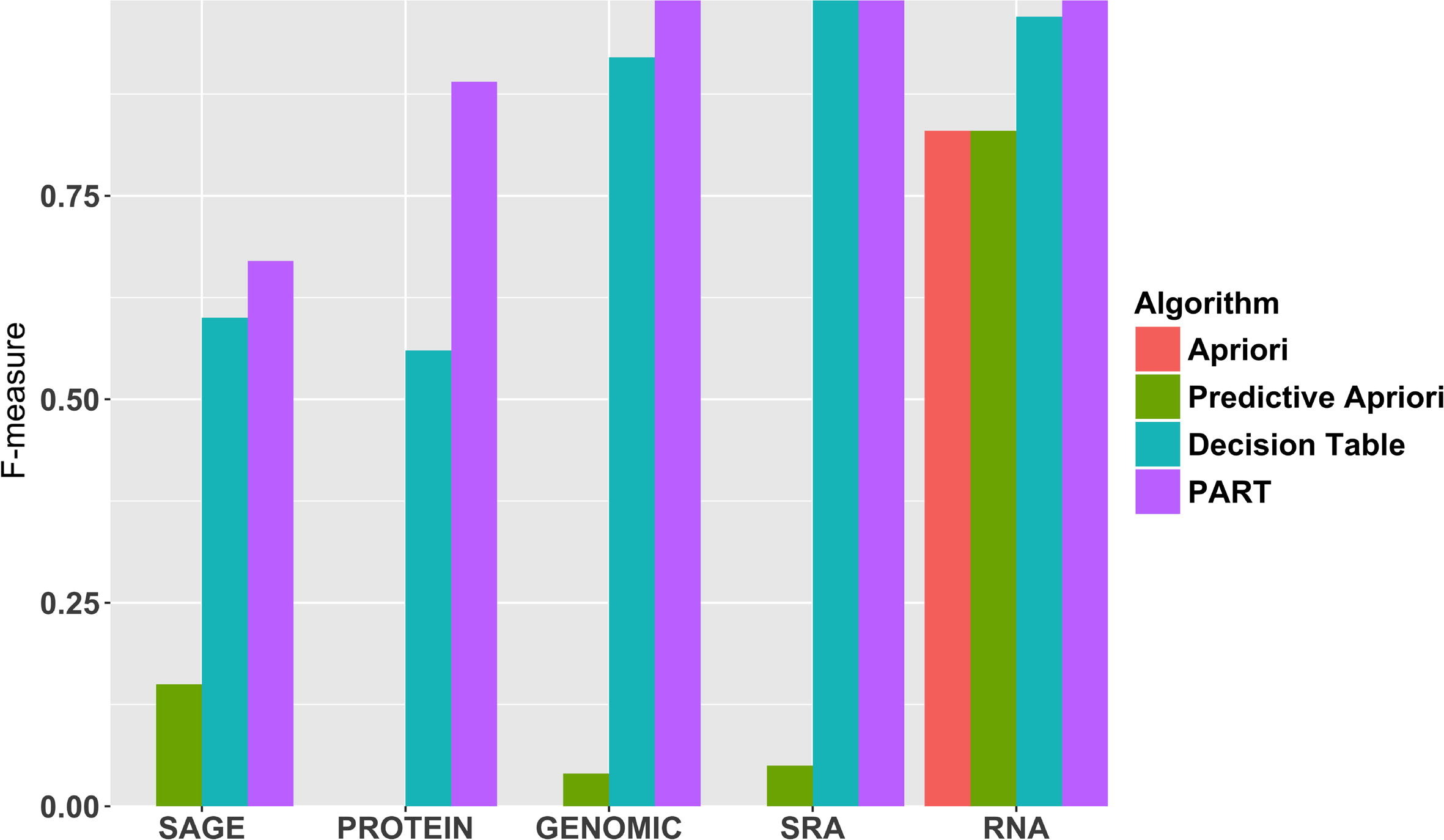
Next, Fig. 3 shows the F-measure to predict the metadata element type using all four algorithms. Our results suggest that the accuracy of predicting specific metadata values can vary significantly for each algorithm. For instance, ‘RNA’, ‘SRA’, and ‘GENOMIC’ is near perfectly predicted by PART, while lower performance is seen for predicting the ‘PROTEIN’ and ‘SAGE’ types. The Decision Table follows the same trend as PART, but is less successful for each metadata value for this metadata element. Apriori and Predictive Apriori predict ‘RNA’, but largely fail for the other values. Apriori generates too many unnecessary candidates. A candidate itemset is unnecessary if at least one of its subsets is infrequent. This is the major reason that we have low performance in Apriori in general.[34] We report the F-measure for all values for all metadata elements in the supplementary materials (Table S1).
|
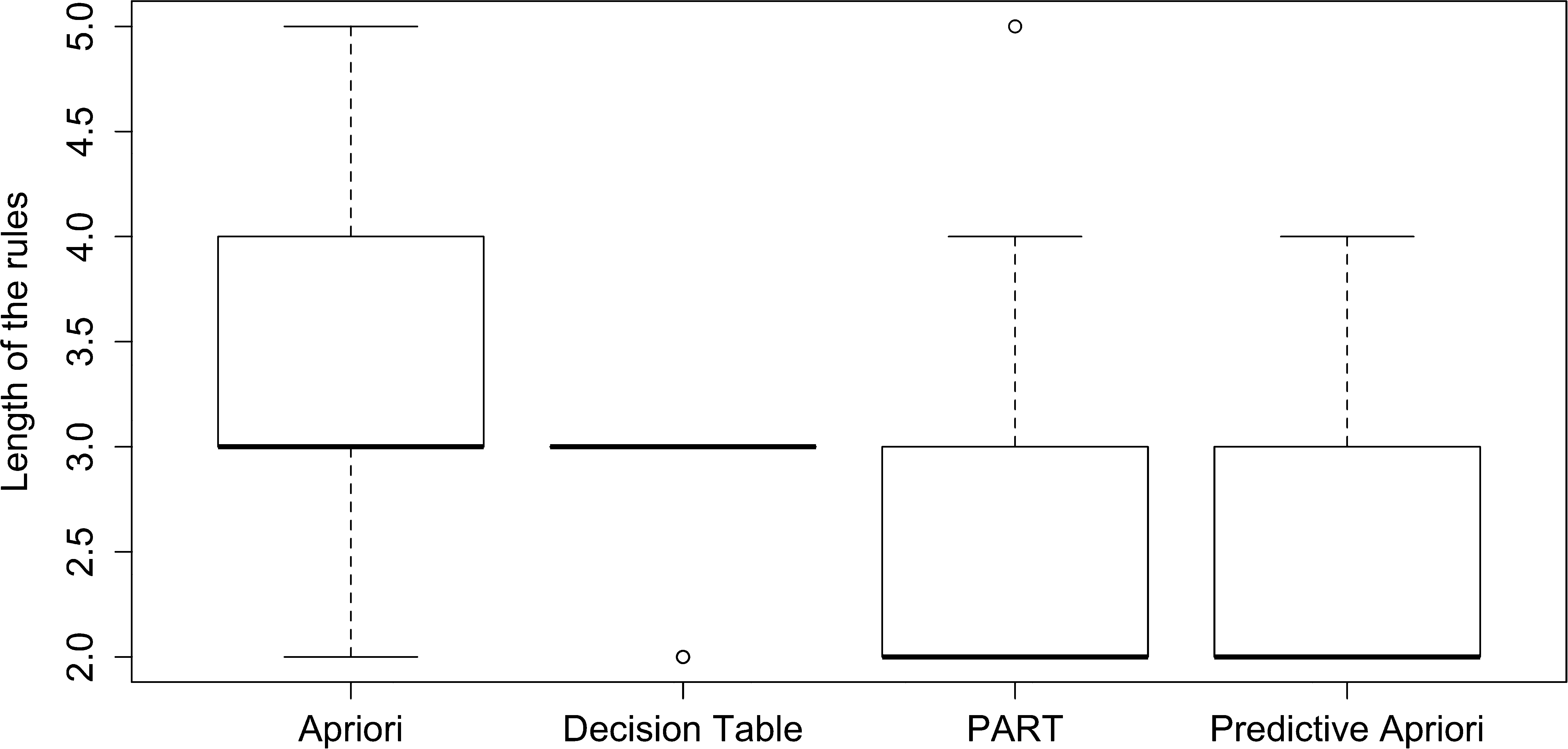
Next, we analyzed the rules to assess whether performance was influenced by length of rule. Fig. 4 shows the rule length for all algorithms. We find that the median length of rules is lowest for PART and Predictive Apriori (length 2), while nearly all of the Decision Table rules have a length of 3. Apriori appears to have the greatest variety in length of rules.
|
Finally, we investigated the associations that exist between GEO metadata, at least as uncovered by each classifier.
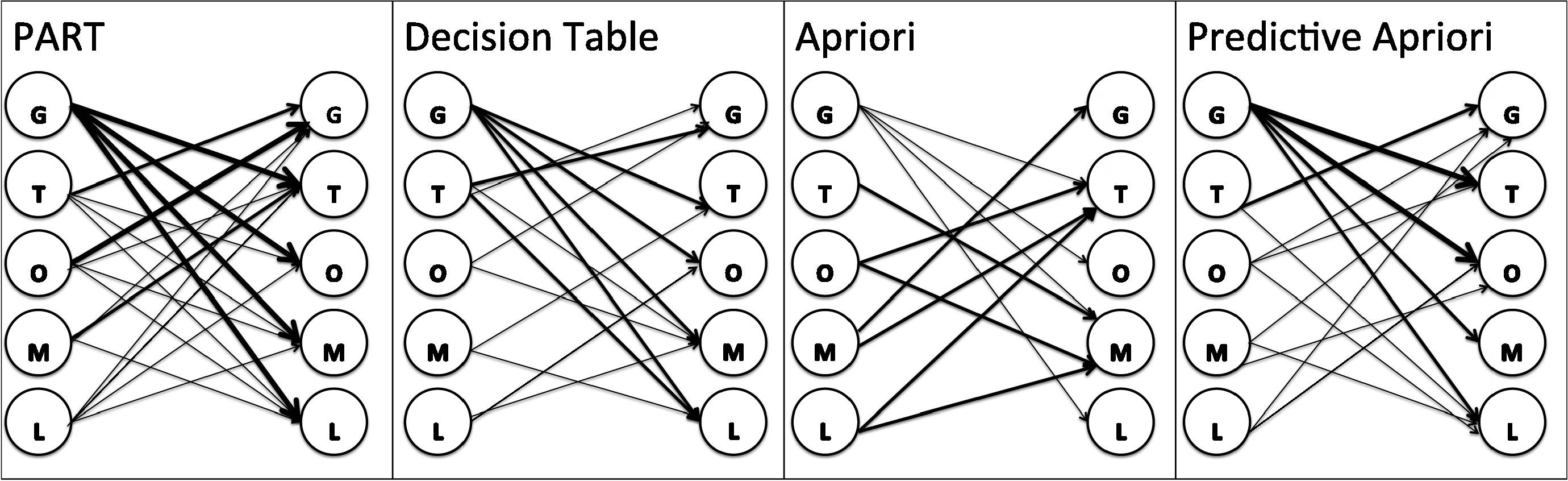
Fig. 5 shows the association network for rules generated by all algorithms. The association network shows the dependency between elements in each algorithm. On the other hand which elements can predict other elements. This association between elements also shows which element is more predictable based on other elements and reveals the power of each element to predict other elements. For example, in PART algorithm the platform (GPL) has a power to predict all other elements. It means we can predict the possible organism, molecule, type and label which are associated with the particular platform. As it shown in Fig. 5, there are tick arrows from platform to other elements, which shows the strong power of prediction of other elements based on the platform. The same description assigned to other algorithm based on the arrows in the network in Fig. 5.
|
5. Discussion
In this work, we explored the use of ARM algorithms to predict structured metadata. Our results, based on the analysis of a subset of GEO’s metadata elements, support the hypothesis that associations between certain metadata elements exist and can be used by ARM algorithms in a predictive manner. Our goal is to simplify the authoring of metadata as much as possible for metadata submitter with predicting the metadata value and recommend that to the metadata submitter during the submission process. We show that algorithms, which have been used in this study, particularly PART and Decision Tables, perform better than using the most frequently occurring metadata value for a particular metadata element (i.e. majority vote classifier). We found differences in the length of rules generated by different algorithms and the quality of their predictions. While our work focused on the metadata in the GEO database, we anticipate that our approach can be applied to other databases of experimental metadata with similar levels of success.
Our research has important implications for initiatives aimed at improving the quantity and quality of metadata in a prospective and retrospective manner. Several efforts are devoted to prospective metadata authoring – they specify metadata that can, should, and minimally must be provided. BioSharing.org[6] catalogs guidelines, standards, and the policies for databases, journals, and funders. Metadata authoring applications such as ISA-Tools[9] or RightField[10] can be used to codify guidelines and enable users to author metadata using ontologies from the NCBO BioPortal.[11] Authoring good metadata is tedious and error-prone, and could benefit from more automation. Our work shows that a subset of metadata elements can be predicted with sufficiently high accuracy. Thus, our predictive approach could be useful for metadata authoring. It could vastly reduce the amount of metadata authoring a submitter must do, but also potentially improve the quantity and quality of metadata. Generating higher quality metadata with less effort is a key part of our NIH BD2K Center for Data Annotation and Retrieval (CEDAR)[7], which is developing intelligent tools for metadata authoring and discovery.[8] We believe that the application of ARM and other machine learning algorithms will greatly accelerate metadata authoring, and improve the quality of research data submissions; failure to do so will likely continue the present situation wherein guidelines are variably applied.[35] Additionally, metadata prediction can be useful retrospectively. Our predictive framework can be used to highlight metadata values that differ from our predictions and may need to be more closely examined. We also anticipate that we could use the approach to predict missing metadata, subject again to further validation by professional users in the field or possibly through crowdsourcing, which has been applied to find and categorize errors in Linked Data.[36] Our work is not without limitations. First, a key limitation in ARM algorithms lies in the vast number of discovered rules and the arbitrary thresholds applied to limit these rules. The main drawback is that the arbitrary thresholds may reduce the amount of information and affect the performance of the classifier specifically when we have the high variety of the values (e.g. values for the platform). Existing approaches employ different parameters to search for interesting rules.[22][37][38] This fact and a large number of rules make it difficult to compare the output of ARM algorithms. Several methods for solving this problem such as rule reduction methods, association rule refinement and association rules for supervised classification have been proposed.[38] Most studies suggest the latest one is the more effective one.[22][30][38] Second, our method is currently focused on learning rules from structured metadata. However, databases of experimental metadata often contain textual descriptions which could not be used directly in our approach. In previous work, we showed that experimental metadata could be predicted using classifiers trained with term frequency-inverse document frequency (TF-IDF) based models.[18]
Finally, while our work showed promise in predicting some of the metadata values in GEO, it remains to be seen how well the approach will be with other experimental databases. We expect that our approach will work well with well structured data sets such as the Sequence Read Archive (SRA), but perhaps do less well on data sets with less metadata. Further study on data sets comprised of different sizes, different varieties of the values for each element, and different combination of structured and unstructured elements is needed. It is also unclear whether data from one database can be usefully combined with data from other databases to improve prediction.
6. Conclusion
We have shown that predicting metadata using ARM algorithms is possible using an existing large biomedical database such as GEO. Future work will focus on expanding this application to other databases such as Biosample datasets (e.g. SRA), more comprehensive metadata as well as aggregation with other models from our previous works on both structured and unstructured metadata.[18] GEO database includes both structured and unstructured metadata as well as other resources. We will extend our methods from previous work such as LDA and TF-IDF to other unstructured data (e.g. abstract of the related manuscript associated with the studies) to improve additional information to improve classification. However, an ensemble classifier could be considered to combine predictions given by different methods, i.e. from rule-based algorithms trained on structured metadata and from other machine learning methods trained on textual features. Predictive metadata can be used both prospectively to facilitate metadata authoring, and retrospectively to improve, correct and augment existing metadata in biomedical databases.
Conflict of interest
There is not conflict of interest.
Acknowledgement
This research is supported by CEDAR, The Center for Expanded Data Annotation and Retrieval (U54 AI117925) awarded by the National Institute of Allergy and Infectious Diseases through funds provided by the trans-NIH Big Data to Knowledge (BD2K) initiative. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Supplementary material
- Supplementary Table S1: Download Comma Separated Value file (3KB)
- Supplementary Table S2: Download Comma Separated Value file (327KB)
References
- ↑ 1.0 1.1 1.2 1.3 Barrett, T.; Wilhite, S.E.; Ledoux, P. et al. (2013). "NCBI GEO: Archive for functional genomics data sets - Update". Nucleic Acids Research 41 (DB1): D991-5. doi:10.1093/nar/gks1193. PMC PMC3531084. PMID 23193258. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3531084.
- ↑ Tryka, K.A.; Hao, L.; Sturcke, A. et al. (2014). "NCBI's Database of Genotypes and Phenotypes: dbGaP". Nucleic Acids Research 42 (DB1): D975-9. doi:10.1093/nar/gkt1211. PMC PMC3965052. PMID 24297256. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3965052.
- ↑ Fan, C.; Prat, A.; Parker, J.S. et al. (2011). "Building prognostic models for breast cancer patients using clinical variables and hundreds of gene expression signatures". BMC Medical Genomics 4: 3. doi:10.1186/1755-8794-4-3. PMC PMC3025826. PMID 21214954. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3025826.
- ↑ Sutherland, A.; Thomas, M.; Brandon, R.A. et al. (2011). "Development and validation of a novel molecular biomarker diagnostic test for the early detection of sepsis". Critical Care 15 (3): R149. doi:10.1186/cc10274. PMC PMC3219023. PMID 21682927. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3219023.
- ↑ Brazma, A.; Hingamp, P.; Quackenbush, J. et al. (2001). "Minimum information about a microarray experiment (MIAME)-toward standards for microarray data". Nature Genetics 29 (4): 365–71. doi:10.1038/ng1201-365. PMID 11726920.
- ↑ 6.0 6.1 Field, D.; Sansone, S.; Delong, E.F. et al. (2010). "Meeting Report: BioSharing at ISMB 2010". Standards in Genomic Sciences 3 (3): 254-8. doi:10.4056/sigs/1403501. PMC PMC3035313. PMID 21304729. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3035313.
- ↑ 7.0 7.1 7.2 7.3 Musen, M.A.; Bean, C.A.; Cheung, K.H. et al. (2015). "The center for expanded data annotation and retrieval". JAMIA 22 (6): 1148–52. doi:10.1093/jamia/ocv048. PMC PMC5009916. PMID 26112029. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5009916.
- ↑ 8.0 8.1 8.2 Panahiazar, M.; Dumontier, M.; Gevaert, O. (2015). "Context aware recommendation engine for metadata submission". First International Workshop on Capturing Scientific Knowledge: 3–7. https://www.isi.edu/ikcap/sciknow2015/papers/Panahiazar.pdf.
- ↑ 9.0 9.1 Rocca-Serra, P.; Brandizi, M.; Maguire, E. et al. (2010). "ISA software suite: Supporting standards-compliant experimental annotation and enabling curation at the community level". Bioinformatics 26 (18): 2354-6. doi:10.1093/bioinformatics/btq415. PMC PMC2935443. PMID 20679334. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2935443.
- ↑ 10.0 10.1 Wolstencroft, K.; Owen, S.; Horridge, M. et al. (2011). "RightField: Embedding ontology annotation in spreadsheets". Bioinformatics 27 (14): 2021–2022. doi:10.1093/bioinformatics/btr312.
- ↑ 11.0 11.1 Noy, N.F.; Shah, N.H.; Whetzel, P.L. et al. (2009). "BioPortal: Ontologies and integrated data resources at the click of a mouse". Nucleic Acids Research 37 (WS1): W170-3. doi:10.1093/nar/gkp440. PMC PMC2703982. PMID 19483092. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2703982.
- ↑ 12.0 12.1 Kotsiantis, S.B. (2007). "Supervised machine learning: A review of classification techniques". Informatica 31 (3): 249–268. http://www.informatica.si/index.php/informatica/article/view/148.
- ↑ Bellazzi, R.; Zupan, B. (2008). "Predictive data mining in clinical medicine: Current issues and guidelines". International Journal of Medical Informatics 77 (2): 81–97. doi:10.1016/j.ijmedinf.2006.11.006. PMID 17188928.
- ↑ Xiong, W.; Liu, H.; Guan, J.; Zhou, S. (2013). "Protein function prediction by collective classification with explicit and implicit edges in protein-protein interaction networks". BMC Bioinformatics 14 (Suppl 12): S4. doi:10.1186/1471-2105-14-S12-S4. PMC PMC3848795. PMID 24267980. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3848795.
- ↑ Daemen, A.; Gevaert, O.; De Moor, B. (2007). "Integration of clinical and microarray data with kernel methods". Conference Proceedings of the IEEE Engineering in Medicine and Biology Society 2007: 5411–5. doi:10.1109/IEMBS.2007.4353566. PMID 18003232.
- ↑ Panahiazar, M.; Taslimitehrani, V.; Pereira, N.; Pathak, J. (2007). "Using EHRs and Machine Learning for Heart Failure Survival Analysis". Studies in Health Technology and Informatics 2015: 5411–5. doi:10.1109/IEMBS.2007.4353566. PMID 18003232.
- ↑ 17.0 17.1 Kumbhare, T.A.; Chobe, S.V. (2014). "An overview of association rule mining algorithms". International Journal of Computer Science and Information Technologies 5 (1): 927–930. http://ijcsit.com/docs/Volume%205/vol5issue01/ijcsit20140501201.pdf.
- ↑ 18.0 18.1 18.2 Posch, L.; Panahiazar, M.; Dumontier, M.; Gevaert, O. (2016). "Predicting structured metadata from unstructured metadata". Database 2016: baw080. doi:10.1093/database/baw080. PMC PMC4892825. PMID 128637268. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4892825.
- ↑ 19.0 19.1 Buckberry, S.; Bent, S.J.; Bianco-Miotto, T.; Roberts, C.T. (2014). "massiR: a method for predicting the sex of samples in gene expression microarray datasets". Bioinformatics 30 (14): 2084-5. doi:10.1093/bioinformatics/btu161. PMC PMC4080740. PMID 24659105. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4080740.
- ↑ 20.0 20.1 20.2 Agrawal, R.; Srikant, R. (1994). "Fast Algorithms for Mining Association Rules in Large Databases". Proceedings of the 20th International Conference on Very Large Data Bases: 487–99. ISBN 1558601538.
- ↑ Piatestsky-Shapiro, G. (1991). "Chapter 13: Discovery, analysis, and presentation of strong rules". In Piatestsky-Shapiro, G.; Frawley, W.. Knowledge Discovery in Databases. MIT Press. pp. 229–248. ISBN 9780262660709.
- ↑ 22.0 22.1 22.2 22.3 Ordonez, C. (2006). "Comparing association rules and decision trees for disease prediction". Proceedings of the International Workshop on Healthcare Information and Knowledge Management: 17–24. doi:10.1145/1183568.1183573. ISBN 1595935282.
- ↑ Brossette, S.E.; Sprague, A.P.; Hardin, J.M. et al. (1998). "Association rules and data mining in hospital infection control and public health surveillance". JAMIA 5 (4): 373-81. PMC PMC61314. PMID 9670134. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC61314.
- ↑ Downs, S.M.; Wallace, M.Y. (2000). "Mining association rules from a pediatric primary care decision support system". Proceedings AMIA Symposium: 200–4. PMC PMC2243862. PMID 11079873. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2243862.
- ↑ Oyama, T.; Kitano, K.; Satou, K.; Ito, T. (2002). "Extraction of knowledge on protein-protein interaction by association rule discovery". Bioinformatics 18 (5): 705-14. PMID 12050067.
- ↑ Chen, T.J.; Chou, L.F.; Hwang, S.J. (2003). "Application of a data-mining technique to analyze coprescription patterns for antacids in Taiwan". Clinical Therapeutics 25 (9): 2453-63. PMID 14604744.
- ↑ Zhu, X.; Suk, H.I.; Shen, D. (2014). "A novel matrix-similarity based loss function for joint regression and classification in AD diagnosis". NeuroImage 100: 91–105. doi:10.1016/j.neuroimage.2014.05.078. PMC PMC4138265. PMID 12050067. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4138265.
- ↑ R Development Core Team (2011). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. ISBN 3900051070. https://www.r-project.org/.
- ↑ Scheffer, T.. "Finding Association Rules That Trade Support Optimally against Confidence". In De Raedt, L.; Siebes, A.. Principles of Data Mining and Knowledge Discovery. Lecture Notes in Computer Science. 2168. Springer, Berlin, Heidelberg. pp. 424–35. doi:10.1007/3-540-44794-6_35. ISBN 9783540425342.
- ↑ 30.0 30.1 Mutter, S.; Hall, M.; Frank, E.. "Using Classification to Evaluate the Output of Confidence-Based Association Rule Mining". In Webb, G.I.; Yu, X.. AI 2004: Advances in Artificial Intelligence. Lecture Notes in Computer Science. 3339. Springer, Berlin, Heidelberg. pp. 538–49. doi:10.1007/978-3-540-30549-1_47. ISBN 9783540240594.
- ↑ Kohavi, R.. "The power of decision tables". In Lavrac, N.; Wrobel, S.. Machine Learning: ECML-95. Lecture Notes in Computer Science. 912. Springer, Berlin, Heidelberg. pp. 174–89. doi:10.1007/3-540-59286-5_57. ISBN 9783540592860.
- ↑ Fürnkranz, J. (1999). "Separate-and-Conquer Rule Learning". Artificial Intelligence Review 13 (1): 3–54. doi:10.1023/A:1006524209794.
- ↑ Powers, D.M.W. (2011). "Evaluation: From precision, recall and F-measure to ROC, informedness, markedness and correlation". Journal of Machine Learning Technologies 2 (1): 37–63. doi:10.9735/2229-3981.2.1.
- ↑ Tank, D.M. (2014). "Improved Apriori Algorithm for Mining Association Rules". International Journal of Information Technology and Computer Science 6 (7): 15–23. doi:10.5815/ijitcs.2014.07.03.
- ↑ Tang, B.M.; Huang, S.J.; McLean, A.S. (2010). "Genome-wide transcription profiling of human sepsis: A systematic review". Critical Care 14 (6): R237. doi:10.1186/cc9392. PMC PMC3219990. PMID 21190579. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3219990.
- ↑ Acosta, M.; Zaveri, A.; Simperl, E. et al.. "Crowdsourcing Linked Data Quality Assessment". In Alani, H.; Kagal, L.; Fokoue, A. et al.. The Semantic Web – ISWC 2013. Lecture Notes in Computer Science. 8219. Springer, Berlin, Heidelberg. pp. 260–76. doi:10.1007/978-3-642-41338-4_17. ISBN 9783642413377.
- ↑ Narvekar, M.; Syed, S.F. (2015). "An Optimized Algorithm for Association Rule Mining Using FP Tree". Procedia Computer Science 45: 101–10. doi:10.1016/j.procs.2015.03.097.
- ↑ 38.0 38.1 38.2 Moreno, M.N.; Segrera, S.; López, V.F.. "Association Rules: Problems, solutions and new applications". In Ruiz, R.; Riquelme, J.C.; Aguilar-Ruiz, J.S.. III Taller de Minería de Datos y Aprendizaje (TAMIDA '05). Editorial Thomson. pp. 317–23. ISBN 8497324498.
Notes
Per the original license, this presentation is faithful to the original, with only minor differences in presentation/format.