Journal:Use of application containers and workflows for genomic data analysis
| Full article title | Use of application containers and workflows for genomic data analysis |
|---|---|
| Journal | Journal of Pathology Informatics |
| Author(s) | Schulz, Wade L.; Durant, Thomas; Siddon, Alexa J.; Torres, Richard |
| Author affiliation(s) | Yale University School of Medicine, VA Connecticut Healthcare System |
| Primary contact | Email: Log in to source site to view |
| Year published | 2016 |
| Volume and issue | 7 |
| Page(s) | 53 |
| DOI | 10.4103/2153-3539.197197 |
| ISSN | 2153-3539 |
| Distribution license | Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported |
| Website | http://www.jpathinformatics.org |
| Download | http://www.jpathinformatics.org/temp/JPatholInform7153-6111104_165831.pdf (PDF) |
Abstract
Background: The rapid acquisition of biological data and development of computationally intensive analyses has led to a need for novel approaches to software deployment. In particular, the complexity of common analytic tools for genomics makes them difficult to deploy and decreases the reproducibility of computational experiments.
Methods: Recent technologies that allow for application virtualization, such as Docker, allow developers and bioinformaticians to isolate these applications and deploy secure, scalable platforms that have the potential to dramatically increase the efficiency of big data processing.
Results: While limitations exist, this study demonstrates a successful implementation of a pipeline with several discrete software applications for the analysis of next-generation sequencing (NGS) data.
Conclusions: With this approach, we significantly reduced the amount of time needed to perform clonal analysis from NGS data in acute myeloid leukemia.
Keywords: Big data, bioinformatics workflow, containerization, genomics
Introduction
The amount of data available for research is growing at an exponential rate. The recent push for open data has also rapidly increased the availability of biomedical datasets for secondary analysis. Examples include the Yale Open Data Access project[1], a repository of clinical trial data, and The Cancer Genome Atlas (TCGA)[2], a project that makes genomic data accessible to researchers after initial findings are released. While these data sets promote ongoing research, the ability to efficiently store, move, and analyze such large repositories is often a bottleneck to analysis.[3]
In addition to the massive growth in volume and availability, novel analyses — including advanced statistical methods and machine learning — often require significant resources for efficient processing. One example of this in biomedical research is the analysis of next generation sequencing (NGS) data. NGS is also known as massively parallel or high-throughput sequencing, as it simultaneously sequences many fragments of DNA, thereby producing enormous amounts of information. These datasets often require several preprocessing steps followed by detailed analysis. In addition to being resource intensive, the reproducibility of computational experiments using these data is often limited due to the complexity of system and software configuration.[4] Some application frameworks have made advances to improve the reproducibility of individual applications and analysis pipelines[5][6], but significant work remains to increase this reliability, particularly for experiments performed in resource-limited environments or on computational clusters.
The deployment of complex computational systems is not unique to bioinformatics. As such, there has been significant progress in building virtualization layers for operating systems and more recently, software applications.[7][8] A current example of this includes the Docker platform (Docker, San Francisco, CA, U.S.A.), which allows for the creation and configuration of software containers for deployment on a range of systems.[9][10] While the use of these technologies has limitations, it also has the potential improve the usability of many software applications in computational biology. As such, several studies and initiatives have begun to focus on the use of Docker in bioinformatics and computer science research.[11][12][13] In this paper, we demonstrate the potential benefits of containerized applications and application workflows for computational genomics research.
Technical background
To augment an ongoing study related to tumor heterogeneity, we obtained access to acute myeloid leukemia (AML) NGS data from TCGA.[14] Aligned NGS data from TCGA are available through the Cancer Genomics Hub (cgHub). The data set of interest consisted of approximately 12 terabytes (TBs) of whole genome sequencing (WGS) data and another 12 TB of whole exome sequencing data. Our analysis required the identification of somatic variants followed by a prediction of tumor heterogeneity using publicly available software tools. Unfortunately, many bioinformatics tools have specific software dependencies and natively run on only a subset of operating systems.[15] In addition, many applications are unable to run their computations in parallel, thus limiting analysis throughput. While increasing the number of servers or individual server resources can improve analysis speed, overall processing may still be less efficient due to these limitations.
As previously noted, the Docker platform allows for virtualized application deployments within a lightweight, Linux-based wrapper called a container.[9][11][15] This approach is similar to operating system virtualization but at the application level. Containerization enables developers to create virtual environments that have only the minimum necessary libraries, which users can quickly deploy on their own infrastructure in a secure, reproducible fashion. In addition, the isolation offered by this approach means that a more robust, parallelized workflow can be created for some applications that do not natively support multi-threading or parallel processing. While this type of workflow implementation is not beneficial for all use cases, scenarios where compute capacity on a single node exceeds what a single application can utilize are likely to benefit from such an approach.
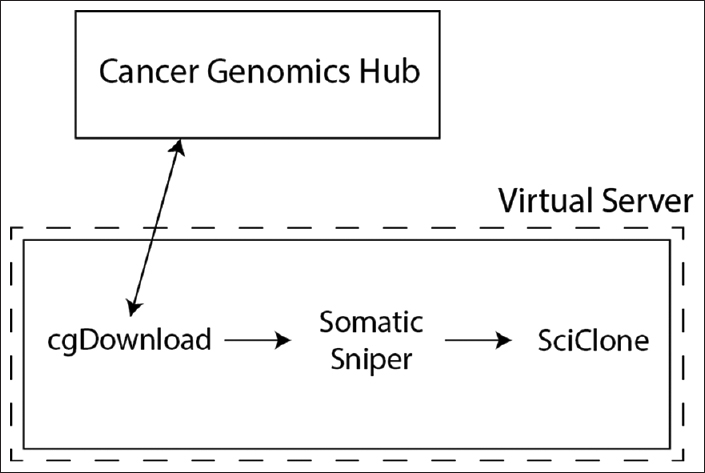
To efficiently predict tumor heterogeneity from TCGA data, we implemented two key technologies: a Python workflow using the Luigi library (Spotify, Stockholm, Sweden)[16] and Docker containers for each key application in our analysis pipeline. While neither of these technologies is unique to genomic analysis or bioinformatics, Luigi is a general workflow orchestration library, and Docker allows for application visualization, as previously noted. For this use case, we were restricted to a single computational node with eight cores and 128 gigabyte (GB) of memory for analysis. As shown in [Figure 1], aligned WGS data were obtained for paired germline and tumor specimens from cgHub (now the genomic data commons) using the cgDownload utility.[17] We identified somatic variants with the SomaticSniper application[18] and generated clonal predictions with the SciClone library.[19] Sequential data acquisition and analysis took approximately four hours to complete per paired sequence but had significant variability between specimens, since file size and the number of genetic mutations varied significantly.
|
Approach
Because of the large number of specimens and the time needed for sequential analysis, our first approach to improving pipeline efficiency was the implementation of a Python workflow using the Luigi library. This library is commonly used for workflow automation and has gained significant traction within bioinformatics through the development of the Sci: Luigi library.[20] Implementation of our pipeline with this workflow library allowed us to include automated fault tolerance, primarily for issues that resulted in sequence download failures due to brief losses in network connectivity. In addition, the workflow could run continuously and unsupervised, which markedly reduced the amount of hands-on time needed for analysis.
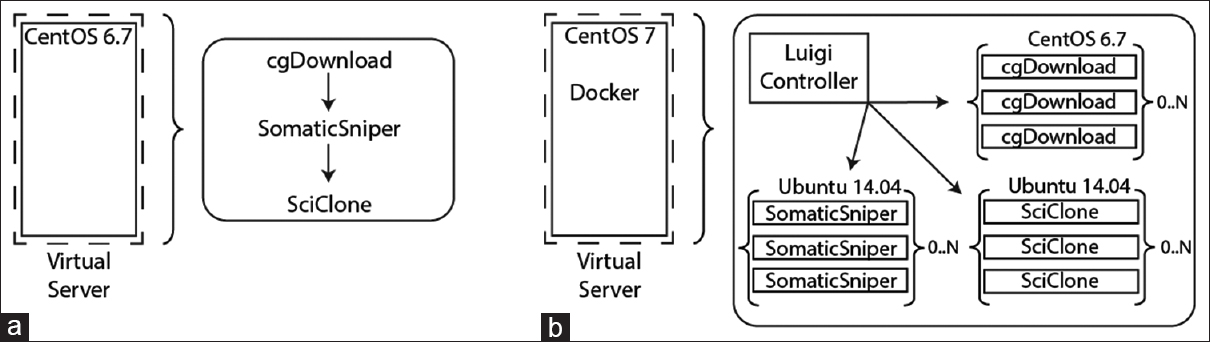
Workflow automation led to a significant improvement in efficiency; however, the inability to parallelize the software packages limited analysis throughput. While the cgDownload utility does support multi-threading, local bandwidth and disk size limitations made bulk downloading of the entire TCGA data set difficult. In addition, a bulk download followed by a full analysis of the data set would mean that network capacity would be saturated while the computational resources remained idle, followed by the opposite scenario. To maintain high, simultaneous utilization of all local hardware resources, including bandwidth, memory, and processor capacity, we deployed the cgDownload utility, SomaticSniper, and SciClone within isolated Docker containers and executed each container with our Luigi workflow [Figure 2]. This approach allowed us to horizontally and vertically scale each application to take full advantage of our local hardware. In addition, each application could be deployed on a single node running CentOS7 (Red Hat, Raleigh, NC, USA) within a container running its natively-supported operating system: CentOS6 for cgDownload and Ubuntu (Canonical, London, UK) for SomaticSniper and SciClone.
|
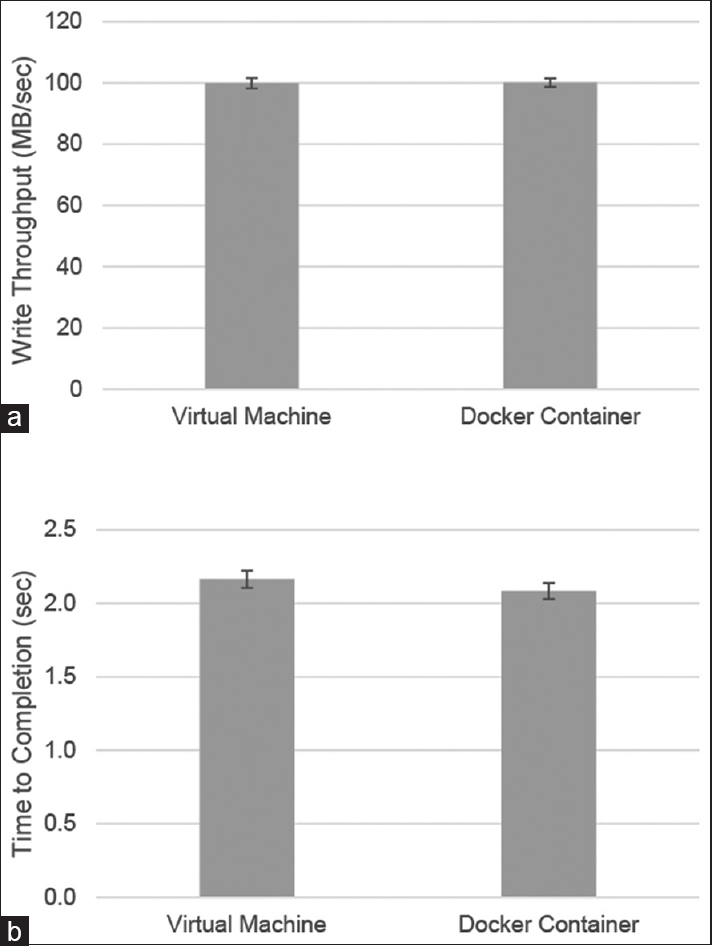
This approach to application deployment can offer significant performance benefits. However, any virtualization technology has the potential to offset these gains due to resource overhead in the virtualization layer. To assess the impact of Docker virtualization on two key metrics, disk throughput, and processing efficiency, we used two benchmarking tools to evaluate performance on the virtual server as well as in a Docker container. To benchmark disk input/output performance, we used the dd command line tool, a standard Linux utility that can be used to read and write files and to gather performance statistics. The dd utility was used to write a 1 GB file and showed similar performance in both a virtual machine and a Docker container within this same environment [Figure 3a]. Similarly, results from sysbench, an open-source benchmarking utility originally created by MySQL AB, found that the time needed to calculate 10,000 primes in either environment was equivalent [Figure 3b]. When combined with evidence from other studies[21], these results demonstrate that Docker has a minimal overhead for these components.
|
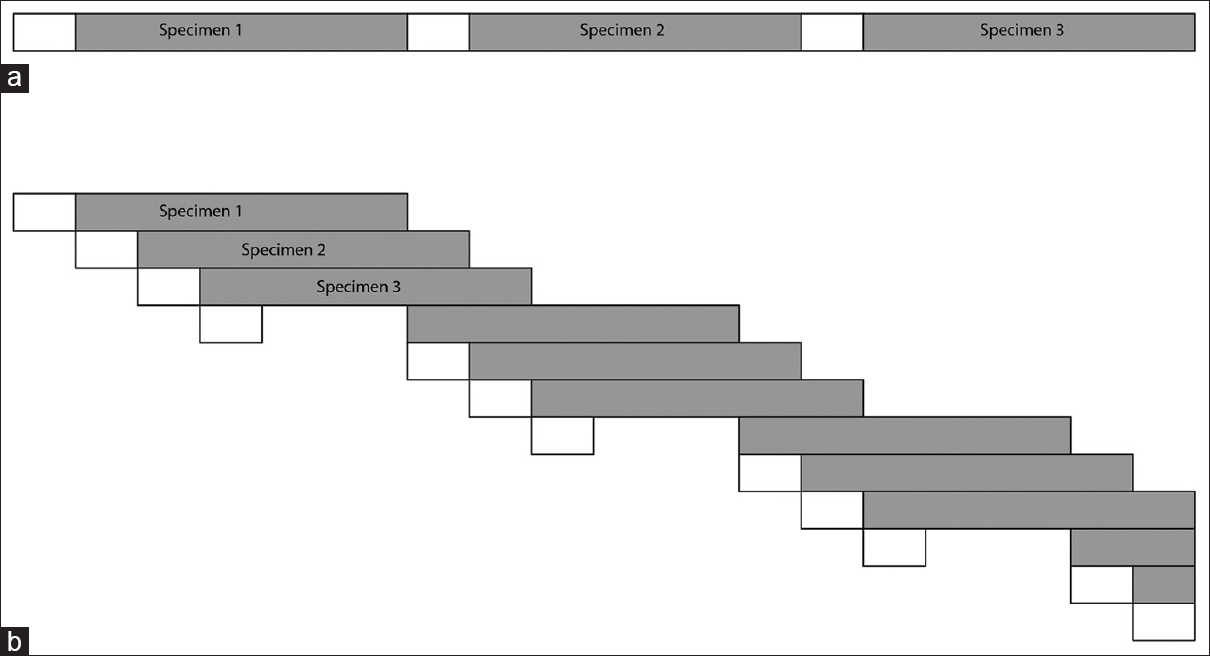
Since we found similar benchmarking results for both the virtual machine and Docker containers, we next executed our workflow to analyze the AML WGS results from TCGA. Using this approach, we were able to stagger data download and processing to take advantage of all system resources [Figure 4]. One limitation to this approach is that none of the tools described here provide built-in resource monitoring. This pipeline was well-suited to parallel analysis since each application had specific, isolated resource needs such as network, storage, or compute capacity, which could be monitored with custom code within the Python workflow. Use of this automated staggered workflow with Docker containers allowed use to analyze fifty specimens from the TCGA data set within approximately 3 days. It is difficult to provide statistical performance metrics since many factors such as data volume, and network bandwidth can significantly alter the overall pipeline performance. However, the general approach of staggered, parallel computation should provide increased processing efficiency for workloads such as this.
|
Conclusion
he complex nature of genomic data and the tools used to analyze these data sets makes efficient processing difficult with standard environments. As noted above, the use of emerging technologies such as Docker in combination with automated workflows may significantly improve the efficiency of data processing in bioinformatics. With the growing number of open data projects, use of these techniques will be necessary to take advantage of available computational resources.
While performance and pipeline efficiency were key components of this implementation, Docker containers also allow for application isolation from the host operating system. Since many bioinformatics tools have complex sets of dependencies and are difficult to build from source, the ability to deploy containers with different operating systems and dependency versions to the same host decreases the amount of effort needed to being analysis. For example, the cgDownload utility is distributed as a compiled binary for use on CentOS 6.7, but can only be deployed on CentOS 7 when built from source, which requires a significant amount of manual configuration. As shown in [Figure 2], the use of containers allowed the deployment of each utility on its natively supported operating system, which improves stability and decreases the potential for dependency conflicts among software applications.
Several other tools exist for the orchestration of containerized applications, such as Kubernetes and Docker Swarm. For complex platforms, these tools can be used to deploy containers across hardware clusters and to integrate networking and storage resources between containers. However, these applications work strictly at the container level and do not inherently provide application-level workflows as presented here. Additional implementation experience about the use of these tools within high-performance clusters may provide valuable insights about the scalability of these tools within bioinformatics workflows.
The above findings demonstrate the promise of emerging technologies to improve the efficiency of genomic analysis. Because of the subsequent increase in analysis throughput, use of these tools means that big data analyses can be done even with limited local computational capacity. Finally, use of container technology can improve pipeline and experimental reproducibility since preconfigured applications can be readily deployed to nearly any host system. While many factors can impact reproducibility, the use of containers limits variability due to differences in software environment or application configuration when appropriately deployed. The continued use of emerging technology and novel approaches to software architecture has the potential to increase the efficiency of computational analysis in bioinformatics.
Financial support and sponsorship
ACLPS Paul E. Strandjord Young Investigator Grant.
Conflicts of interest
There are no conflicts of interest.
References
- ↑ Krumholz, H.M.; Waldstreicher, J. (2016). "The Yale Open Data Access (YODA) Project--A mechanism for data sharing". New England Journal of Medicine 375 (5): 403–405. doi:10.1056/NEJMp1607342. PMID 27518657.
- ↑ Collins, F.S.; Barker, A.D. (2007). "Mapping the cancer genome: Pinpointing the genes involved in cancer will help chart a new course across the complex landscape of human malignancies". Scientific American 296 (3): 50–7. PMID 17348159.
- ↑ Fan, J.; Han, F.; Liu, H. (2014). "Challenges of big data analysis". National Science Review 1 (2): 293–314. doi:10.1093/nsr/nwt032. PMC PMC4236847. PMID 25419469. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4236847.
- ↑ Nekrutenko, A.; Taylor, J. (2012). "Next-generation sequencing data interpretation: Enhancing reproducibility and accessibility". Nature Reviews Genetics 13 (9): 667–72. doi:10.1038/nrg3305. PMID 22898652.
- ↑ Blankenberg, Daniel; Von Kuster, Gregory; Coraor, Nathaniel; Ananda, Guruprasad; Lazarus, Ross; Mangan, Mary; Nekrutenko, Anton; Taylor, James (2010). "Galaxy: a web-based genome analysis tool for experimentalists". Current Protocols in Molecular Biology 19 (Unit 19.10.1–21). doi:10.1002/0471142727.mb1910s89. PMC PMC4264107. PMID 20069535. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4264107.
- ↑ Hatakeyama, M.; Opitz, L.; Russo, G.; Qi, W.; Schlapbach, R.; Rehrauer, H. (2016). "SUSHI: An exquisite recipe for fully documented, reproducible and reusable NGS data analysis". BMC Bioinformatics 17: 228. doi:10.1186/s12859-016-1104-8.
- ↑ Dudley, J.T.; Butte, A.J. (2010). "In silico research in the era of cloud computing". Nature Biotechnology 28 (11): 1181–5. doi:10.1038/nbt1110-1181. PMC PMC3755123. PMID 21057489. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3755123.
- ↑ Howe, B. (2012). "Virtual appliances, cloud computing, and reproducible research". Computing in Science & Engineering 14 (4): 36–41. doi:10.1109/MCSE.2012.62.
- ↑ 9.0 9.1 "Docker". Docker, Inc. https://www.docker.com/. Retrieved 21 November 2016.
- ↑ Anderson, C. (2015). "Docker". IEEE Software 32 (3): 102–105. doi:10.1109/MS.2015.62.
- ↑ 11.0 11.1 Boettiger, C. (2015). "An introduction to Docker for reproducible research". SIGOPS Operating Systems Review 49 (1): 71–79. doi:10.1145/2723872.2723882.
- ↑ Hung, L.H.; Kristiyanto, D.; Lee, S.B.; Yeung, K.Y. (2016). "GUIdock: Using Docker containers with a common graphics user interface to address the reproducibility of research". PLoS One 11 (4): e0152686. doi:10.1371/journal.pone.0152686. PMC PMC4821530. PMID 27045593. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4821530.
- ↑ Moreews, F.; Sallou, O.; Ménager, H. et al. (2015). "BioShaDock: A community driven bioinformatics shared Docker-based tools registry". F1000Research 4: 1443. doi:10.12688/f1000research.7536.1. PMC PMC4743153. PMID 26913191. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4743153.
- ↑ Cancer Genome Atlas Research Network (2013). "Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia". New England Journal of Medicine 368 (22): 2059-74. doi:10.1056/NEJMoa1301689. PMC PMC3767041. PMID 23634996. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3767041.
- ↑ 15.0 15.1 Piccolo, S.R.; Frampton, M.B. (2016). "Tools and techniques for computational reproducibility". GigaScience 5 (1): 30. doi:10.1186/s13742-016-0135-4. PMC PMC4940747. PMID 27401684. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4940747.
- ↑ "Spotify/Luigi". GitHub, Inc. https://github.com/spotify/luigi. Retrieved 21 November 2016.
- ↑ "Genomic Data Commons". National Cancer Institute. https://gdc.cancer.gov/. Retrieved 21 November 2016.
- ↑ Larson, D.E.; Harris, C.C.; Chen, K. et al. (2012). "SomaticSniper: Identification of somatic point mutations in whole genome sequencing data". Bioinformatics 28 (3): 311-7. doi:10.1093/bioinformatics/btr665. PMC PMC3268238. PMID 22155872. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3268238.
- ↑ Miller, C.A.; White, B.S.; Dees, N.D. et al. (2014). "SciClone: inferring clonal architecture and tracking the spatial and temporal patterns of tumor evolution". PLoS Computational Biology 10 (8): e1003665. doi:10.1371/journal.pcbi.1003665. PMC PMC4125065. PMID 25102416. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4125065.
- ↑ "Pharmbio/SciLuigi". GitHub, Inc. https://github.com/pharmbio/sciluigi. Retrieved 21 November 2016.
- ↑ Preeth, E.N.; Mulerickal, F.J.; Paul, B.; Sastri, Y. (2015). "Evaluation of Docker containers based on hardware utilization". Proceedings of the 2015 International Conference on Control Communication and Computing India 2015: 697–700. doi:10.1109/ICCC.2015.7432984.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added.