Difference between revisions of "Journal:Analyzing huge pathology images with open source software"
Shawndouglas (talk | contribs) (Added content. Saving and adding more.) |
Shawndouglas (talk | contribs) (Added content. Saving and adding more.) |
||
| Line 64: | Line 64: | ||
The LargeTIFFTools and NDPITools are based on the open source TIFF<ref name="LefflerLib12">{{cite web |url=http://www.remotesensing.org/libtiff/ |title=LibTIFF – TIFF Library and Utilities |author=Sam Leffler, S.; the authors of LibTIFF |date=2012}}</ref> and JPEG<ref name="LaneTheInd13">{{cite web |url=http://www.ijg.org/ |title=The Independent JPEG Group’s JPEG software |author=Lane, T.G.; Vollbeding, G. |date=2013}}</ref> or libjpeg-turbo<ref name="Lanelibjpeg12">{{cite web |url=http://libjpeg-turbo.virtualgl.org/ |title=libjpeg-turbo |author=Lane, T.G.; Vollbeding, G.; the authors of the libjpeg-turbo software |date=2012}}</ref> libraries. The NDPITools plugins for ImageJ are based on the Java API of ImageJ<ref name="RasbandImageJ12" /><ref name="SchneiderNIH12">{{cite journal |title=NIH Image to ImageJ: 25 years of image analysis |journal=Nature Methods |author=Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. |volume=9 |pages=671-675 |year=2012 |doi=10.1038/nmeth.2089 |pmid=22930834}}</ref> and on the open source software Image-IO <ref name="SachaImage04">{{cite web |url=http://ij-plugins.sourceforge.net/plugins/imageio/ |title=Image IO Plugin Bundle |author=Sacha, J. |date=2004}}</ref>, and use the Java Advanced Imaging 1.1.3 library.<ref name="SunJava06">{{cite web |url=http://www.oracle.com/technetwork/java/current-142188.html |archiveurl=http://www.webcitation.org/query.php?url=http://www.oracle.com/technetwork/java/current-142188.html&refdoi=10.1186/1746-1596-8-92 |title=Java Advanced Library 1.1.3 |author=Sun Microsystems, Inc |date=2006 |archivedate=11 July 2013}}</ref> | The LargeTIFFTools and NDPITools are based on the open source TIFF<ref name="LefflerLib12">{{cite web |url=http://www.remotesensing.org/libtiff/ |title=LibTIFF – TIFF Library and Utilities |author=Sam Leffler, S.; the authors of LibTIFF |date=2012}}</ref> and JPEG<ref name="LaneTheInd13">{{cite web |url=http://www.ijg.org/ |title=The Independent JPEG Group’s JPEG software |author=Lane, T.G.; Vollbeding, G. |date=2013}}</ref> or libjpeg-turbo<ref name="Lanelibjpeg12">{{cite web |url=http://libjpeg-turbo.virtualgl.org/ |title=libjpeg-turbo |author=Lane, T.G.; Vollbeding, G.; the authors of the libjpeg-turbo software |date=2012}}</ref> libraries. The NDPITools plugins for ImageJ are based on the Java API of ImageJ<ref name="RasbandImageJ12" /><ref name="SchneiderNIH12">{{cite journal |title=NIH Image to ImageJ: 25 years of image analysis |journal=Nature Methods |author=Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. |volume=9 |pages=671-675 |year=2012 |doi=10.1038/nmeth.2089 |pmid=22930834}}</ref> and on the open source software Image-IO <ref name="SachaImage04">{{cite web |url=http://ij-plugins.sourceforge.net/plugins/imageio/ |title=Image IO Plugin Bundle |author=Sacha, J. |date=2004}}</ref>, and use the Java Advanced Imaging 1.1.3 library.<ref name="SunJava06">{{cite web |url=http://www.oracle.com/technetwork/java/current-142188.html |archiveurl=http://www.webcitation.org/query.php?url=http://www.oracle.com/technetwork/java/current-142188.html&refdoi=10.1186/1746-1596-8-92 |title=Java Advanced Library 1.1.3 |author=Sun Microsystems, Inc |date=2006 |archivedate=11 July 2013}}</ref> | ||
===Basic functions=== | |||
The basic functions are the following. They can be performed even on a computer with a modest amount of RAM (see below the “Performance” discussion). | |||
1. splitting a tiled TIFF file into multiple TIFF files, one for each of the tiles (tiffsplittiles program); | |||
2. extracting (“cropping”) quickly a given rectangle of a supposedly tiled TIFF file into a TIFF or JPEG file (tifffastcrop program); | |||
3. splitting one or several TIFF file(s), possibly very large, into mosaic(s), without fully decompressing them in memory (tiffmakemosaic program); | |||
4. converting a NDPI file into a standard multiple-image TIFF file, tiled if necessary, using upon request the BigTIFF format introduced in version 4.0.0 of the TIFF library<ref name="LaneTheInd13" /><ref name="BTDesign12">{{cite web |url=http://www.remotesensing.org/libtiff/bigtiffdesign.html |title=BigTIFF Design |date=2012}}</ref><ref name="BTProp08">{{cite web |url=http://www.awaresystems.be/imaging/tiff/bigtiff.html |title=The BigTIFF File Format Proposal |date=2008}}</ref>, and encoding magnification and focus levels as TIFF “image description” fields (ndpi2tiff program); | |||
5. creating a standard TIFF file for all or part of the magnification levels and focus levels present in the given NDPI file (the user can ask for specific magnification and focus levels and for a specific rectangular region of the image), and, upon request, creating a mosaic for each image which doesn’t fit into RAM or for all images (ndpisplit program). The names of the created files are built on the name of the source file and incorporate the magnification and focus levels (and, in the case of mosaic pieces, the coordinates inside the mosaic). | |||
==References== | ==References== | ||
Revision as of 23:05, 26 September 2015
| Full article title | Analyzing huge pathology images with open source software |
|---|---|
| Journal | Diagnostic Pathology |
| Author(s) | Deroulers, Christophe; Ameisen, David; Badoual, Mathilde; Gerin, Chloé; Granier, Alexandre; Lartaud, Marc |
| Author affiliation(s) | Université Paris Diderot, Université Paris-Sud, Montpellier RIO Imaging, CIRAD |
| Primary contact | Email: deroulers@imnc.in2p3.fr |
| Year published | 2013 |
| Volume and issue | 8 |
| Page(s) | 92 |
| DOI | 10.1186/1746-1596-8-92 |
| ISSN | 1746-1596 |
| Distribution license | Creative Commons Attribution 2.0 Generic |
| Website | http://www.diagnosticpathology.org/content/8/1/92 |
| Download | http://www.diagnosticpathology.org/content/pdf/1746-1596-8-92.pdf (PDF) |
|
|
This article should not be considered complete until this message box has been removed. This is a work in progress. |
Abstract
Background
Digital pathology images are increasingly used both for diagnosis and research, because slide scanners are nowadays broadly available and because the quantitative study of these images yields new insights in systems biology. However, such virtual slides build up a technical challenge since the images occupy often several gigabytes and cannot be fully opened in a computer’s memory. Moreover, there is no standard format. Therefore, most common open source tools such as ImageJ fail at treating them, and the others require expensive hardware while still being prohibitively slow.
Results
We have developed several cross-platform open source software tools to overcome these limitations. The NDPITools provide a way to transform microscopy images initially in the loosely supported NDPI format into one or several standard TIFF files, and to create mosaics (division of huge images into small ones, with or without overlap) in various TIFF and JPEG formats. They can be driven through ImageJ plugins. The LargeTIFFTools achieve similar functionality for huge TIFF images which do not fit into RAM. We test the performance of these tools on several digital slides and compare them, when applicable, to standard software. A statistical study of the cells in a tissue sample from an oligodendroglioma was performed on an average laptop computer to demonstrate the efficiency of the tools.
Conclusions
Our open source software enables dealing with huge images with standard software on average computers. They are cross-platform, independent of proprietary libraries and very modular, allowing them to be used in other open source projects. They have excellent performance in terms of execution speed and RAM requirements. They open promising perspectives both to the clinician who wants to study a single slide and to the research team or data centre who do image analysis of many slides on a computer cluster.
Virtual slides
The virtual slide(s) for this article can be found here: http://www.diagnosticpathology.diagnomx.eu/vs/5955513929846272
Keywords: Digital pathology; Image processing; Virtual slides; Systems biology; ImageJ; NDPI
Background
Virtual microscopy has become routinely used over the last few years for the transmission of pathology images (the so-called virtual slides), for both telepathology and teaching.[1][2] In more and more hospitals, virtual slides are even attached to the patient’s file.[3][4] They have also a great potential for research, especially in the context of multidisciplinary projects involving e.g. mathematicians and clinicians who do not work at the same location. Quantitative histology is a promising new field, involving computer-based morphometry or statistical analysis of tissues.[5][6][7][8][9] A growing number of works report the pertinence of such images for diagnosis and classification of diseases, e.g. tumours.[10][11][12][13][14] Databases of clinical cases[15] will include more and more digitized tissue images. This growing use of virtual microscopy is accompanied by the development of integrated image analysis systems offering both virtual slide scanning and automatic image analysis, which makes integration into the daily practice of pathologists easier. See Ref. 16[16] for a review of some of these systems.
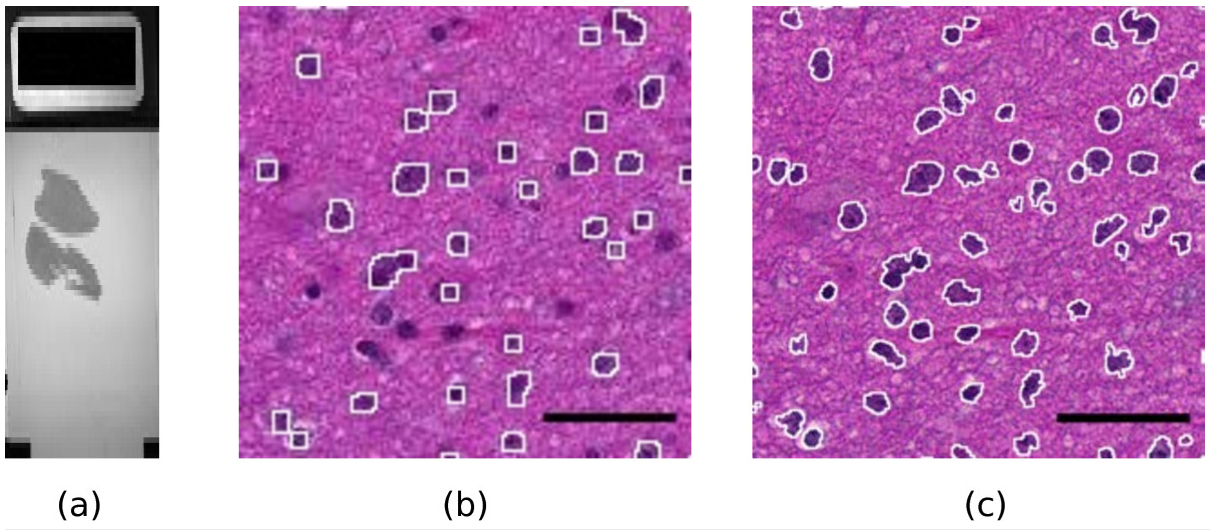
Modern slide scanners produce high magnification microscopy images of excellent quality[1], for instance at the so-called “40x” magnification. They allow much better visualization and analysis than lower magnification images. As an example, Figure 1 shows two portions of a slide at different magnifications, 10x and 40x. The benefit of the high magnification for both diagnosis and automated image analysis is clear. For instance, the state of the chromatin inside the nucleus and the cell morphology, better seen at high magnification, are essential to help the clinician distinguish tumorous and non-tumorous cells. An accurate, non-pixelated determination of the perimeters of the cell nuclei is needed for morphometry and statistics.
Figure 1. A sample slide. (a): macroscopic view of the whole slide (the black rectangle on the left is 1x2 cm).
(b,c): Influence of the magnification on the quality of results. (b): a portion of the slide scanned at magnification
level 10x. The white contours show the result of an automatic detection of the dark cell nuclei with the ImageJ
software. A significant fraction of the cell nuclei is missed and the contours are rather pixelated. (c): the same
portion of the slide scanned at magnification 40x. The white contours show the result of the same automatic
detection. Almost all cell nuclei are detected and the shapes of the contours are much more precise.
Scale bar: 4 μm.
However, this technique involves the manipulation of huge images (of the order of 10 billions of pixels for a full-size slide at magnification 40x with a single focus level) for which the approach taken by most standard software, loading and decompressing the full image into RAM, is impossible (a single slice of a full-size slide needs of the order of 30 GiB of RAM). As a result, standard open-source software such as ImageJ[17], ImageMagick[18] or GraphicsMagick[19] completely fails or is prohibitively slow when used on these images. Of course, commercially available software exists[16], but it is usually quite expensive, and very often restricted to a single operating system. It uses proprietary source code, which is a problem if one wants to control or check the algorithms and their parameters when doing image analysis for research.
In addition, many automated microscopes or slide scanners store the images which they produce into proprietary or poorly documented file formats, and the software provided by vendors is often specific to some operating system. This leads to several concerns. First, it makes research based on digital pathology technically more difficult. Even when a project is led on a single site, one has often to use clusters of computers to achieve large-scale studies of many full-size slides from several patients.[20] Since clusters of computers are typically run by open source software such as Linux, pathology images stored in non-standard file formats are a problem. Furthermore, research projects are now commonly performed in parallel in several sites, not to say in several countries, thanks to technology such as Grid[21], and there is ongoing efforts towards the interoperability of information systems used in pathology.[3][22] Second, proprietary formats may hinder the development of shared clinical databases[15] and access of the general public to knowledge, whereas the citizen should receive benefit of public investments. Finally, they may also raise financial concerns and conflicts of interest.[23]
There have been recent attempts to define open, documented, vendor-independent software[24][25], which partly address this problem. However, very large images stored in the NDPI file format produced by some slide scanners manufactured by Hamamatsu, such as the NanoZoomer, are not yet fully supported by such software. For instance, LOCI Bio-Formats[25] is presently unable to open images, one dimension of which is larger than 65k, and does not deal properly with NDPI files of more than 4 GiB. OpenSlide[24] does not currently support the NDPI format. NDPI-Splitter[26]needs to be run on Windows and depends on a proprietary library.
To address these problems, we have developed open source tools which achieve two main goals: reading and converting images in the NDPI file format into standard open formats such as TIFF, and splitting a huge image, without decompressing it entirely into RAM, into a mosaic of much smaller pieces (tiles), each of which can be easily opened or processed by standard software. All this is realized with high treatment speed on all platforms.
Implementation
Overview
The main software is implemented in the C programming language as separate, command-line driven executables. It is independent of any proprietary library. This ensures portability on a large number of platforms (we have tested several versions of Mac OS X, Linux and Windows), modularity and ease of integration into scripts or other software projects.
It is complemented by a set of plugins for the public domain software ImageJ[17], implemented in Java, which call the main executables in an automatic way to enable an interactive use.
The LargeTIFFTools and NDPITools are based on the open source TIFF[27] and JPEG[28] or libjpeg-turbo[29] libraries. The NDPITools plugins for ImageJ are based on the Java API of ImageJ[17][30] and on the open source software Image-IO [31], and use the Java Advanced Imaging 1.1.3 library.[32]
Basic functions
The basic functions are the following. They can be performed even on a computer with a modest amount of RAM (see below the “Performance” discussion).
1. splitting a tiled TIFF file into multiple TIFF files, one for each of the tiles (tiffsplittiles program);
2. extracting (“cropping”) quickly a given rectangle of a supposedly tiled TIFF file into a TIFF or JPEG file (tifffastcrop program);
3. splitting one or several TIFF file(s), possibly very large, into mosaic(s), without fully decompressing them in memory (tiffmakemosaic program);
4. converting a NDPI file into a standard multiple-image TIFF file, tiled if necessary, using upon request the BigTIFF format introduced in version 4.0.0 of the TIFF library[28][33][34], and encoding magnification and focus levels as TIFF “image description” fields (ndpi2tiff program);
5. creating a standard TIFF file for all or part of the magnification levels and focus levels present in the given NDPI file (the user can ask for specific magnification and focus levels and for a specific rectangular region of the image), and, upon request, creating a mosaic for each image which doesn’t fit into RAM or for all images (ndpisplit program). The names of the created files are built on the name of the source file and incorporate the magnification and focus levels (and, in the case of mosaic pieces, the coordinates inside the mosaic).
References
- ↑ 1.0 1.1 Diamond, J.; McCleary, D. (2009). "Virtual Microscopy". In Hannon-Fletcher, M.; Maxwell, P.. Advanced Techniques in Diagnostic Cellular Pathology. Chichester UK: John Wiley & Sons, Ltd. ISBN 9780470515976.
- ↑ Ameisen, D.; Yunès, J.B.; Deroulers, C.; Perrier, V.; Bouhidel, F.; Battistella, M.; Legrès, L.; Janin, A.; Bertheau, P. (2013). "Stack or Trash? Fast quality assessment of virtual slides". Diagnostic Pathology 8 (Suppl 1): S23. doi:10.1186/1746-1596-8-S1-S23. PMC PMC3849546. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3849546.
- ↑ 3.0 3.1 García Rojo, M.; Castro, A.M.; Gonçalves, L. (2011). "COST action “EuroTelepath”: digital pathology integration in electronic health record, including primary care centres". Diagnostic Pathology 6 (Suppl 1): S6. doi:10.1186/1746-1596-6-S1-S6. PMC PMC3073224. PMID 21489201. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3073224.
- ↑ Ameisen, D. (2013). "Intégration des lames virtuelles dans le dossier patient électronique". PhD thesis. Univ Paris Diderot-Paris 7.
- ↑ Collan, Y.; Torkkeli, T.; Personen, E.; Jantunen, E.; Kosma, V.M. (1987). "Application of morphometry in tumor pathology". Analytical and Quantitative Cytology and Histology 9 (2): 79–88. PMID 3300687.
- ↑ Wolfe, P.; Murphy, J.; McGinley, J.; Zhu, Z.; Jiang, W.; Gottschall, E.; Thompson, H. (2004). "Using nuclear morphometry to discriminate the tumorigenic potential of cells: A comparison of statistical methods". Cancer Epidemiology, Biomarkers & Prevention 13 (6): 976–988. PMID 15184254.
- ↑ Gürcan, M.N.; Boucheron, L.E.; Can, A.; Madabhushi, A.; Rajpoot, N.M.; Yener, B. (2009). "Histopathological image analysis: a review". IEEE Reviews in Biomedical Engineering 2009 (2): 147–171. doi:10.1109/RBME.2009.2034865. PMC PMC2910932. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2910932.
- ↑ Gerin, C.; Pallud, J.; Deroulers, C.; Varlet, P.; Oppenheim, C.; Roux, F.X.; Chrétien, F.; Thomas, S.R.; Grammaticos, B.; Badoual, M. (2013). "Quantitative characterization of the imaging limits of diffuse low-grade oligodendrogliomas". Neuro-Oncology 15 (10): 1379-88. doi:10.1093/neuonc/not072. PMC PMC3779035. PMID 23771168. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3779035.
- ↑ Wienert, S.; Heim, D.; Kotani, M.; Lindequist, B.; Stenzinger, A.; Ishii, M.; Hufnagl, P.; Beil, M.; Dietel, M.; Denkert, C.; Klauschen, F. (2013). "CognitionMaster: an object-based image analysis framework". Diagnostic Pathology 8: 34. doi:10.1186/1746-1596-8-34. PMC PMC3626931. PMID 23445542. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3626931.
- ↑ Gunduz, C.; Yener, B.; Gultekin, S.H. (2004). "The cell graphs of cancer". Bioinformatics 20 (Suppl 1): i145-i151. doi:10.1093/bioinformatics/bth933. PMID 15262793.
- ↑ Gunduz, C.; Gultekin, S.H.; Yener, B. (2005). "Augmented cell-graphs for automated cancer diagnosis". Bioinformatics 21 (Suppl 2): ii7-ii12. doi:10.1093/bioinformatics/bti1100. PMID 16204128.
- ↑ West, N.P.; Dattani, M.; McShane, P.; Hutchins, G.; Grabsch, J.; Mueller, W.; Treanor, D.; Quirke, P.; Grabsch, H. (2010). "The proportion of tumour cells is an independent predictor for survival in colorectal cancer patients". British Journal of Cancer 102 (10): 1519–1523. doi:10.1038/sj.bjc.6605674. PMC PMC2869173. PMID 20407439. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2869173.
- ↑ Chang, H.; Han, J.; Borowsky, A.; Loss, L.; Gray, J.W.; Spellman, P.T.; Parvin, B. (2013). "Invariant delineation of nuclear architecture in Glioblastoma multiforme for clinical and molecular association". IEEE Transactions on Medical Imaging 32 (4): 670–682. doi:10.1109/TMI.2012.2231420. PMC PMC3728287. PMID 23221815. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3728287.
- ↑ Kayser, K.; Radziszowski, D.; Bzdyl, P.; Sommer, R.; Kayser, G. (2006). "Towards an automated virtual slide screening: theoretical considerations and practical experiences of automated tissue-based virtual diagnosis to be implemented in the internet". Diagnostic Pathology 1: 10. doi:10.1186/1746-1596-1-10. PMC PMC1524814. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1524814.
- ↑ 15.0 15.1 PLGA Foundation (2012). "Meta analysis low grade glioma database project". Archived from the original on 11 July 2013. http://www.webcitation.org/query.php?url=http://www.fightplga.org/research/PLGA-Sponsored_Projects/MetaAnalysis&refdoi=10.1186/1746-1596-8-92.
- ↑ 16.0 16.1 García Rojo, M.; Bueno, G.; Slodkowska, J. (2009). "Review of imaging solutions for integrated quantitative immunohistochemistry in the Pathology daily practice". Folia Histochemica et Cytobiologica 47 (3): 349–354. doi:10.2478/v10042-008-0114-4. PMID 20164017.
- ↑ 17.0 17.1 17.2 Rasband, W.S. (2012). "ImageJ". http://imagej.nih.gov/ij/.
- ↑ ImageMagick Studio, LLC (2013). "ImageMagick". http://www.imagemagick.org/.
- ↑ GraphicsMagick Group (2013). "GraphicsMagick". http://www.graphicsmagick.org/.
- ↑ Kong, J.; Cooper, L.A.D.; Wang, F.; Chisolm, C.; Moreno, C.S.; Kurc, T.M.; Widener, P.M.; Brat, D.J.; Saltz, J.H. (2011). "A comprehensive framework for classification of nuclei in digital microscopy imaging: An application to diffuse gliomas". IEEE International Symposium on Biomedical Imaging 2011 Mar 30: 2128–2131. doi:10.1109/ISBI.2011.5872833. PMC PMC3256584. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3256584.
- ↑ Kayser, K.; Görtler, J.; Borkenfeld, S.; Kayser, G. (2011). "Grid computing in image analysis". Diagnostic Pathology 6 (Suppl 1): S12. PMC PMC3073205. PMID 21516880. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3073205.
- ↑ Granier, A.; Olivier, M.; Laborie, S.; Vaudescal, S.; Baecker, V.; Tran-Aupiais, C. (2013). "WIDE (Web Images and Data Environment)". Archived from the original on 11 July 2013. http://www.webcitation.org/query.php?url=http://www.mri.cnrs.fr/index.php?m=81&refdoi=10.1186/1746-1596-8-92.
- ↑ Kayser, K. (2012). "Introduction of virtual microscopy in routine surgical pathology — a hypothesis and personal view from Europe". Diagnostic Pathology 7: 48. doi:10.1186/1746-1596-7-48. PMC PMC3441330. PMID 22546238. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3441330.
- ↑ 24.0 24.1 Goode, A.; Satyanarayanan, M. (2008). "A vendor-neutral library and viewer for whole-slide images" (PDF). Technical Report CMU-CS-08-136. School of Computer Science, Carnegie Mellon University. Archived from the original on 11 July 2013. http://www.webcitation.org/query.php?url=http://reports-archive.adm.cs.cmu.edu/anon/2008/CMU-CS-08-136.pdf&refdoi=10.1186/1746-1596-8-92.
- ↑ 25.0 25.1 Linkert, M.; Rueden, C.T.; Allan, C.; Burel, J.M.; Moore, W.; Patterson, A.; Loranger, B.; Moore, J.; Neves, C.; MacDonald, D.; Tarkowska, A.; Sticco, C.; Hill, E.; Rossner, M.; Eliceiri, K.W.; Swedlow, J.R. (2010). "Metadata matters: access to image data in the real world". Journal of Cell Biology 198 (5): 777-782. doi:10.1083/jcb.201004104. PMC PMC2878938. PMID 20513764. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2878938.
- ↑ Khushi, M.; Edwards, G.; de Marcos, D.A.; Carpenter, J.E.; Graham, J.D.; Clarke, C.L. (2013). "Open source tools for management and archiving of digital microscopy data to allow integration with patient pathology and treatment information". Diagnostic Pathology 8: 22. doi:10.1186/1746-1596-8-22. PMC PMC3575263. PMID 23402499. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3575263.
- ↑ Sam Leffler, S.; the authors of LibTIFF (2012). "LibTIFF – TIFF Library and Utilities". http://www.remotesensing.org/libtiff/.
- ↑ 28.0 28.1 Lane, T.G.; Vollbeding, G. (2013). "The Independent JPEG Group’s JPEG software". http://www.ijg.org/.
- ↑ Lane, T.G.; Vollbeding, G.; the authors of the libjpeg-turbo software (2012). "libjpeg-turbo". http://libjpeg-turbo.virtualgl.org/.
- ↑ Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. (2012). "NIH Image to ImageJ: 25 years of image analysis". Nature Methods 9: 671-675. doi:10.1038/nmeth.2089. PMID 22930834.
- ↑ Sacha, J. (2004). "Image IO Plugin Bundle". http://ij-plugins.sourceforge.net/plugins/imageio/.
- ↑ Sun Microsystems, Inc (2006). "Java Advanced Library 1.1.3". Archived from the original on 11 July 2013. http://www.webcitation.org/query.php?url=http://www.oracle.com/technetwork/java/current-142188.html&refdoi=10.1186/1746-1596-8-92.
- ↑ "BigTIFF Design". 2012. http://www.remotesensing.org/libtiff/bigtiffdesign.html.
- ↑ "The BigTIFF File Format Proposal". 2008. http://www.awaresystems.be/imaging/tiff/bigtiff.html.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In most of the article's references DOIs and PubMed IDs were not given; they've been added to make the references more useful. In some cases important information was missing from the references, and that information was added.