Difference between revisions of "Main Page/Featured article of the week/2020"
Shawndouglas (talk | contribs) (Created as needed.) |
Shawndouglas (talk | contribs) (Added last week's article of the week) |
||
| Line 17: | Line 17: | ||
<!-- Below this line begin pasting previous news --> | <!-- Below this line begin pasting previous news --> | ||
<h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: | <h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: January 2–12:</h2> | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Bhattacharya FrontInOnc2019 9.jpg|240px]]</div> | ||
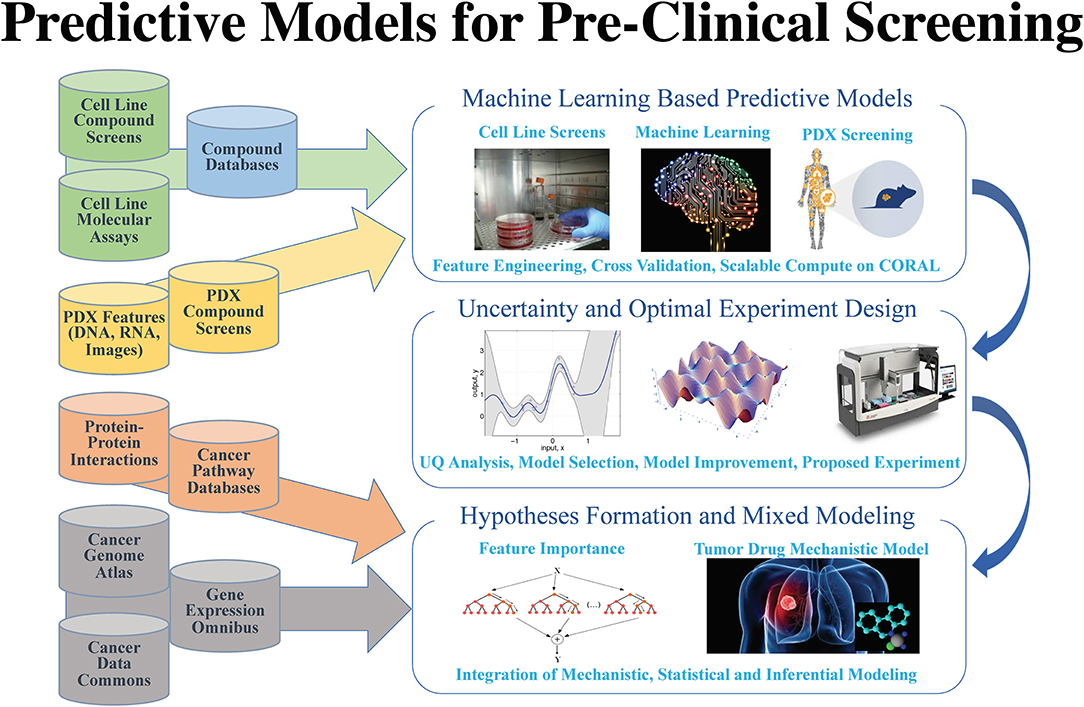
'''"[[Journal: | '''"[[Journal:AI meets exascale computing: Advancing cancer research with large-scale high-performance computing|AI meets exascale computing: Advancing cancer research with large-scale high-performance computing]]"''' | ||
The application of data science in [[cancer]] research has been boosted by major advances in three primary areas: (1) data: diversity, amount, and availability of biomedical data; (2) advances in [[artificial intelligence]] (AI) and machine learning (ML) algorithms that enable learning from complex, large-scale data; and (3) advances in computer architectures allowing unprecedented acceleration of simulation and machine learning algorithms. These advances help build ''in silico'' ML models that can provide transformative insights from data, including molecular dynamics simulations, [[Sequencing|next-generation sequencing]], omics, [[Molecular imaging|imaging]], and unstructured clinical text documents. Unique challenges persist, however, in building ML models related to cancer, including: (1) access, sharing, labeling, and integration of multimodal and multi-institutional data across different cancer types; (2) developing AI models for cancer research capable of scaling on next-generation high-performance computers; and (3) assessing robustness and reliability in the AI models. ('''[[Journal:AI meets exascale computing: Advancing cancer research with large-scale high-performance computing|Full article...]]''')<br /> | |||
|- | |- | ||
|} | |} | ||
Revision as of 17:34, 14 January 2020
|
|
If you're looking for other "Article of the Week" archives: 2014 - 2015 - 2016 - 2017 - 2018 - 2019 - 2020 |
Featured article of the week archive - 2020
Welcome to the LIMSwiki 2020 archive for the Featured Article of the Week.
Featured article of the week: January 2–12:"AI meets exascale computing: Advancing cancer research with large-scale high-performance computing" The application of data science in cancer research has been boosted by major advances in three primary areas: (1) data: diversity, amount, and availability of biomedical data; (2) advances in artificial intelligence (AI) and machine learning (ML) algorithms that enable learning from complex, large-scale data; and (3) advances in computer architectures allowing unprecedented acceleration of simulation and machine learning algorithms. These advances help build in silico ML models that can provide transformative insights from data, including molecular dynamics simulations, next-generation sequencing, omics, imaging, and unstructured clinical text documents. Unique challenges persist, however, in building ML models related to cancer, including: (1) access, sharing, labeling, and integration of multimodal and multi-institutional data across different cancer types; (2) developing AI models for cancer research capable of scaling on next-generation high-performance computers; and (3) assessing robustness and reliability in the AI models. (Full article...) |