Difference between revisions of "Template:Article of the week"
Shawndouglas (talk | contribs) (Updated article of the week text) |
Shawndouglas (talk | contribs) (Updated article of the week text) |
||
| Line 1: | Line 1: | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File: | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Nieminen GigaScience2023 12.jpeg|240px]]</div> | ||
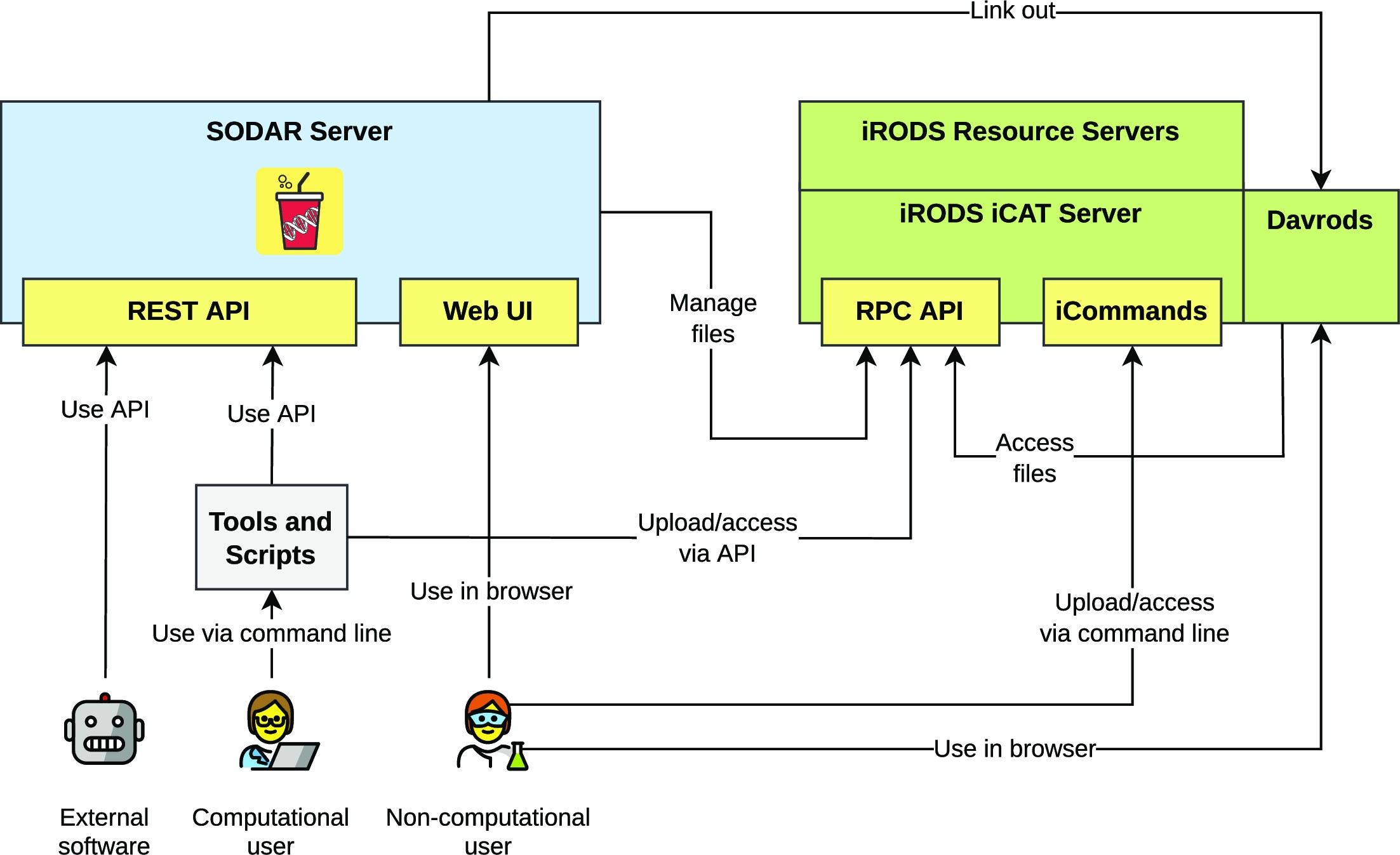
'''"[[Journal: | '''"[[Journal:SODAR: Managing multiomics study data and metadata|SODAR: Managing multiomics study data and metadata]]"''' | ||
Scientists employing omics in [[Life sciences industry|life science]] studies face challenges such as the modeling of multiassay studies, recording of all relevant parameters, and managing many [[Sample (material)|samples]] with their [[metadata]]. They must manage many large files that are the results of the assays or subsequent computation. Users with diverse backgrounds, ranging from computational scientists to wet-lab scientists, have dissimilar needs when it comes to data access, with programmatic interfaces being favored by the former and graphical ones by the latter. We introduce SODAR, the system for [[omics]] data access and retrieval. SODAR is a software package that addresses these challenges by providing a web-based graphical user interface (GUI) for managing multiassay studies and describing them using the ISA (Investigation, Study, Assay) data model and the ISA-Tab file format ... ('''[[Journal:SODAR: Managing multiomics study data and metadata|Full article...]]''')<br /> | |||
''Recently featured'': | ''Recently featured'': | ||
{{flowlist | | {{flowlist | | ||
* [[Journal:Benefits of information technology in healthcare: Artificial intelligence, internet of things, and personal health records|Benefits of information technology in healthcare: Artificial intelligence, internet of things, and personal health records]] | |||
* [[Journal:A quality assurance discrimination tool for the evaluation of satellite laboratory practice excellence in the context of European regulatory meat inspection for Trichinella spp.|A quality assurance discrimination tool for the evaluation of satellite laboratory practice excellence in the context of European regulatory meat inspection for ''Trichinella spp.'']] | * [[Journal:A quality assurance discrimination tool for the evaluation of satellite laboratory practice excellence in the context of European regulatory meat inspection for Trichinella spp.|A quality assurance discrimination tool for the evaluation of satellite laboratory practice excellence in the context of European regulatory meat inspection for ''Trichinella spp.'']] | ||
* [[Journal:Developing a framework for open and FAIR data management practices for next generation risk- and benefit assessment of fish and seafood|Developing a framework for open and FAIR data management practices for next generation risk- and benefit assessment of fish and seafood]] | * [[Journal:Developing a framework for open and FAIR data management practices for next generation risk- and benefit assessment of fish and seafood|Developing a framework for open and FAIR data management practices for next generation risk- and benefit assessment of fish and seafood]] | ||
}} | }} | ||
Revision as of 15:34, 18 March 2024
"SODAR: Managing multiomics study data and metadata"
Scientists employing omics in life science studies face challenges such as the modeling of multiassay studies, recording of all relevant parameters, and managing many samples with their metadata. They must manage many large files that are the results of the assays or subsequent computation. Users with diverse backgrounds, ranging from computational scientists to wet-lab scientists, have dissimilar needs when it comes to data access, with programmatic interfaces being favored by the former and graphical ones by the latter. We introduce SODAR, the system for omics data access and retrieval. SODAR is a software package that addresses these challenges by providing a web-based graphical user interface (GUI) for managing multiassay studies and describing them using the ISA (Investigation, Study, Assay) data model and the ISA-Tab file format ... (Full article...)
Recently featured:
- Benefits of information technology in healthcare: Artificial intelligence, internet of things, and personal health records
- A quality assurance discrimination tool for the evaluation of satellite laboratory practice excellence in the context of European regulatory meat inspection for Trichinella spp.
- Developing a framework for open and FAIR data management practices for next generation risk- and benefit assessment of fish and seafood