Journal:Development of Biosearch System for biobank management and storage of disease-associated genetic information
| Full article title | Development of Biosearch System for biobank management and storage of disease-associated genetic information |
|---|---|
| Journal | Journal of King Saud University - Science |
| Author(s) | Karim, Sajjad; Al-Kharraz, Mona; Mirza, Zeenat; Noureldin, Hend; Abusamara, Heba; Alganmi, Nofe; Merdad, Adnan; Jastaniah, Saddig; Kumar, Sudhir; Rasool, Mahmood; Abuzenadah, Adel; Al-Qahtani, Mohammed |
| Author affiliation(s) | King Abdulaziz University, Temple University |
| Primary contact | Email: skarim1 at kau dot edu dot sa |
| Year published | 2022 |
| Volume and issue | 34(2) |
| Article # | 101760 |
| DOI | 10.1016/j.jksus.2021.101760 |
| ISSN | 1018-3647 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://www.sciencedirect.com/science/article/pii/S1018364721004225 |
| Download | https://www.sciencedirect.com/science/article/pii/S1018364721004225/pdfft (PDF) |
Abstract
Objective: Databases and software are important to manage modern high-throughput laboratories and store clinical and genomic information for quality assurance. Commercial software is expensive, with proprietary code issues, while academic versions have adaptation issues. Our aim was to develop an adaptable in-house software system that can store specimen- and disease-associated genetic information in biobanks to facilitate translational research.
Methods: A prototype was designed per the research requirements, and computational tools were used to develop the software under three tiers, using Visual Basic and ASP.net for the presentation tier, SQL Server for the data tier, and Ajax and JavaScript for the business tier. We retrieved specimens from the biobank using this software and performed microarray-based transcriptomic analysis to detect differentially expressed genes (DEGs) with p-value <0.05 and fold change (FC) ±2 in triple-negative breast cancer cases. The Ingenuity Pathway Analysis (IPA) tool was used to predict canonical molecular pathways associated with disease. Overall performance and utility of software was evaluated by Apache JMeter software, CRUD function testing, and a set of user feedback questionnaires.
Results: We developed Biosearch System, a web-based laboratory information management system (LIMS) enabling management of biobank samples (e.g., tissue, blood, FTTP slides) and their extracts (e.g., DNA, RNA, and proteins) with clinical and experimental details. The client satisfaction feedback was excellent, with a score of 4.7 out of 5. We identified a total of 1,181 DEGs, including both upregulated (IFI6, LEF1, FANCI, CASC5, PLXNA3, etc.) and down-regulated (ADH1B, LYVE1, ADH1C, ADH1B, ADIPOQ, PLIN1, LYVE1, etc.) genes in triple-negative breast cancer. Pathway analysis of DEGs revealed significant activation of interferon signaling (z-score 2.646) and kinetochore metaphase signaling pathway (z-score 2.138) in cancer.
Conclusion: Biosearch System is a user friendly LIMS for collection, storage, and retrieval of specimen and clinical information. It is secure, efficient, and convenient in sample tracking and data analysis. We illustrated its utility in transcriptomic study of breast cancer. Additionally, it can facilitate and speed up any genomic study and translational research publications.
Keywords: Biosearch System, LIMS, database, biobank, genomics, microarray, bioinformatics
Graphical abstract:
Introduction
Biobanks are established for long-term storage and conservation facility for biological specimens along with their demographic, clinical and experimental information to support scientific investigation using bioinformatics tools.[1] That investigation, tied to high-throughput technological advancements in next-generation sequencing (NGS) and microarrays, has generated significant amounts of data at relatively low costs.[2][3][4][5] However, as the amount of biological specimens grows into the thousands, manual methods fail in the efficient handling of specimens and samples, and the risk of information being lost increases. Quality becomes another major concern with increasing volume of big data. As such, a robust software solution becomes necessary to effectively manage biobank and laboratory information and ensure the quality of results with approved ethical guidelines and personal integrity.[6][7][8][9] Software is helpful in managing the workflow cycle—comprising of collection, storage, analysis, and report generation—facilitating quick and easy retrieval of information and speeding up biomarker and therapeutic discoveries.[10]
The collection of a significant cohort size with crucial factors like number and type of samples, clinical information, pathological finding, follow-up data, etc. is a time-consuming process but is strongly recommended to facilitate comprehensive translational research.[11][12][13][14] Bioinformatics software aiding with these tasks is either available as commercial off-the-shelf (COTS), with proprietary code and high cost, or as academic or open-source, with complexity that is difficult to adopt within other laboratories.[13][15][16][17][18] As such, we developed and implemented a software system to support biobank investigators in accessing all clinical and experimental disease-associated information required for basic and translational research. Herein, we discuss the application of our in-house laboratory information management system (LIMS), Biosearch System, for genomic analysis specifically for breast cancer transcriptomics, with further extensibility for any other diseases.
Materials and methods
Biosearch System was developed using a three-tier architecture model:
- Presentation tier: An interactive web browser for the end user’s computer
- Data tier: A SQL Server Management Studio that manages the storage and DB-server
- Business tier: A bridge between the other two tiers, facilitating the collection of data from the presentation tier and validating it before finally sending it to the data tier, and vice versa
Biosearch System follows standard guidelines like legal requirements, standardized architecture, workflow, and dedicated staff members, as well as technical procedures to record and access clinical data, as described herein.
Legal requirements
The LIMS has been designed in accordance with the Saudi Arabian approved regulation for genomic medicine research. Patients were informed prior and written consents were taken by their doctors for their samples to be sent to the biobank for research. We provided consent forms to patients approved by the ethics committee. Additionally, the software stores data in coded format to hide the identity of patients during any research presentation and publications. In order to ensure bioethical safety, we followed Saudi's National Committee of Biological and Medical Ethics guidelines. (Royal decree No. M/59, dated 14/9/1431H – 24/8/2010)
Software and hardware architecture
We began with a prototype, an early model building process to test a proposed concept to enhance precision by system analysts and users. Database software design started by first designing a low fidelity prototype (paper-based prototype) and then designing a high-fidelity prototype (computer-based prototype). Collected information was categorized into logical groups or entities like "sample," "storage condition," "specimen request," "approval system," "project type," "diagnosis," and "disease type," and an entity-relationship diagram was built to show numerical relationship among different entities.
The LIMS itself was built on servers with following specifications: Windows Server 2012 R2 Enterprise Edition service packs 2, .NET 4.5, IIS 8.5, VS 2012, Microsoft SQL Server 2008 R2, ASP.NET, and AJAX Control Kit 4.5. A web browser was used for the graphical user interface (GUI). Database software was developed to support the high-performance hardware system and is compatible with common web browsers. To develop a robust and reliable LIMS, the following features were included:
- a web-based application for wide access to the database;
- a specimen and sample labelling mechanism before enrolling them into the database;
- security mechanisms that limit role-based access to authorized personnel; and
- a disaster management system to cope with any natural disaster.
System structure
To manage the laboratory's services, the software addresses the (i) control of specimen and sample data (i.e., add, edit, confirm transfer, retrieve, and search); (ii) viewing of the log book; (iii) display of a specimen or sample's status; (iv) viewing of box content; (v) addition of new entities, tests, categories, etc. (e.g., diagnostic tests, extractions, hospitals, sample types, projects); and (vi) request of a specimen or sample. Specimens and samples stored in the biobank are provided to researchers per project requirements and with proper justification and ethical approval. The requested sample is first checked by biobank staff then forwarded to the lab manager, before finally going to the director for final approval. Researchers can track the processing steps while the software updates the decision status by email. After delivery of approved specimens, requisite volume is automatically subtracted from biobank stock.
Database structure
A normal relational database is used for the system. Information is stored in specific tables like the SampleInfo table, which contains the sample's information, the ProductInfo table, which contains the type of sample information (e.g., DNA, RNA), the PatientInfo table, which contains a patient's related information, the SampleStorage table, which contains the storage location information (e.g., refrigerators, shelf, box container), and the UsersInfo table, which contains user-related information. The type of relation is one-to-many (one patient::multi-samples, one sample::multi-products, one refrigerator::multi-samples, etc.).
Workflow and biobank organizational structure
The software can only be used with an authorized username and password, and after successful login on a web-based client PC, users are allowed to add new data, update/edit data, follow up with patients, and retrieve data for analysis and interpretation. Biosearch System also manages the storage, quality, quantity, distribution, and maintenance of specimens (e.g., tissue, blood, serum, etc.) and its derivatives (e.g., DNA, RNA, protein, plasma, etc.). Per the existing CEGMR biobank organizational structure, assigned users have different level of rights for using the software. Researchers are end users of the biobank; they request samples based on their active research projects. Dedicated biobank staff examine the request and update the status, handing it off to the supervisor, who approves or rejects the request with valid reason. However, the final decision is made by the director, who has ultimate authority to reconsider the request or reverse the supervisors' decision. Once approved by the director, the biobank delivers the requested items within a week. Reminders for pending tasks are emailed to relevant staff until the final decision is made. This system enables researchers full access to the biobank information by allowing for the management of specimens and samples, the notification for request approval or rejection, and provision of the overall summary of the biobank’s inventory to researchers.
Technical procedure for linking clinical information to biobank specimens
The LIMS manages all the relevant information for all deposited patients’ specimens and samples for systematic clinical research. We use a medical record number (MRN) as the primary key and a biobank number as the secondary key to connect two sections of the database. Patients’ samples are stored at the biobank with a unique allocated biobank number, which consists of 10 digits formatted as sample type (XX), serial number (XXXX), year (XX), and extraction type (XX). Sample types in our database includes AM (amniotic fluid), BL (blood), BO (bone marrow), CO (cord blood), CS (cervix swab), LN (lymph node), PC (product of conception), PL (paraffin embedded tissue with lymph node), PN (normal paraffin embedded tissue), PT (tumor paraffin embedded tissue), TM (tumor tissue), and TN (normal tissue). Depending on research needs, extraction is done from raw specimens, and the extraction type includes D (DNA), R (RNA), P (protein), etc. For a peripheral blood specimen received with serial number 1252 in year 2014, with a DNA sample then extracted from it, our assigned biobank number will be “BL-1252-14D.” This nomenclature system provides a clue about samples; however, patient confidentiality is protected as per current security regulations. To guarantee quality of samples, the biobank has ascertained standard policies via a quality management system. Specimens and samples are stored in liquid nitrogen, −80 °C, −20 °C, or 4 °C refrigerators depending on its type and requirement. Vials containing specimen or extracted sample are stored in an assigned area so that all aliquots can be retrieved in the LIMS from the defined physical location.
Evaluation of software features and performance efficiency
We evaluated the efficiency of Biosearch System by performance testing using (i) Apache JMeter software, (ii) Create, Read, Update, and Delete (CRUD) function testing, and (ii) and a set of user feedback questionnaires. To verify the efficiency (i.e., speed, scalability, and stability) of the LIMS, a performance test was done through Apache JMeter by running different numbers of users (i.e., 500, 100, 50, 10, 1), with 50 loop and 10 ramp-up periods. CRUD function testing using different SQL statement was also used for performance testing.
We also evaluated the LIMS through feedback provided by questionnaire concerning the LIMS' features and efficiency. Answers allowed for a response using a scale from 1 to 5 to specific questions such as:
- Is it easy to reach the website application?
- Does website application load quickly?
- Are the fonts easy to read on various screen resolutions?
- Is the color scheme appropriate and comfortable to the eye?
- Is the content logically separated and appearing in an appropriate way?
- Is the website application easy to use?
- Is navigation easy?
- Are all buttons (internal and external) valid and active?
- Is the copy and paste feature allowed?
- Is the autocomplete feature allowed?
- Do clickable icons work smoothly on a single click?
- Is the website application free from server-side errors?
- Are you able to search, retrieve, and edit data (e.g., samples description, patient information, project details, etc.) easily?
- Do you accurately receive an alert message for missing data?
- Is data printed in an appropriate tabular format?
- Is the help desk easy to contact for any issue in regards to the web-based application?
Transcriptomic analysis of breast cancer using specimens from Biosearch System
We retrieved breast cancer samples and healthy controls using transcriptomic profiling via the Affymetrix platform. Partek GS v6.7 (Partek, USA) was used for data analysis. Imported data was normalized with a robust multi-array averaging process, and ANOVA was applied to the generated DEG gene list using a p value <0.05 and fold change (FC) >2. Principle component analysis (PCA) was done for high-dimensional visualization. Unsupervised hierarchical clustering was done for significant DEG as a similarity matrix.
Molecular pathway analysis
Pathways for DEGs were analyzed using the Ingenuity Pathway Analysis (IPA) tool (Ingenuity, USA) in triple-negative breast cancer cases. IPA predicts molecular networks, canonical pathways associated with uploaded DEG with p-value and FC cutoffs.
Results
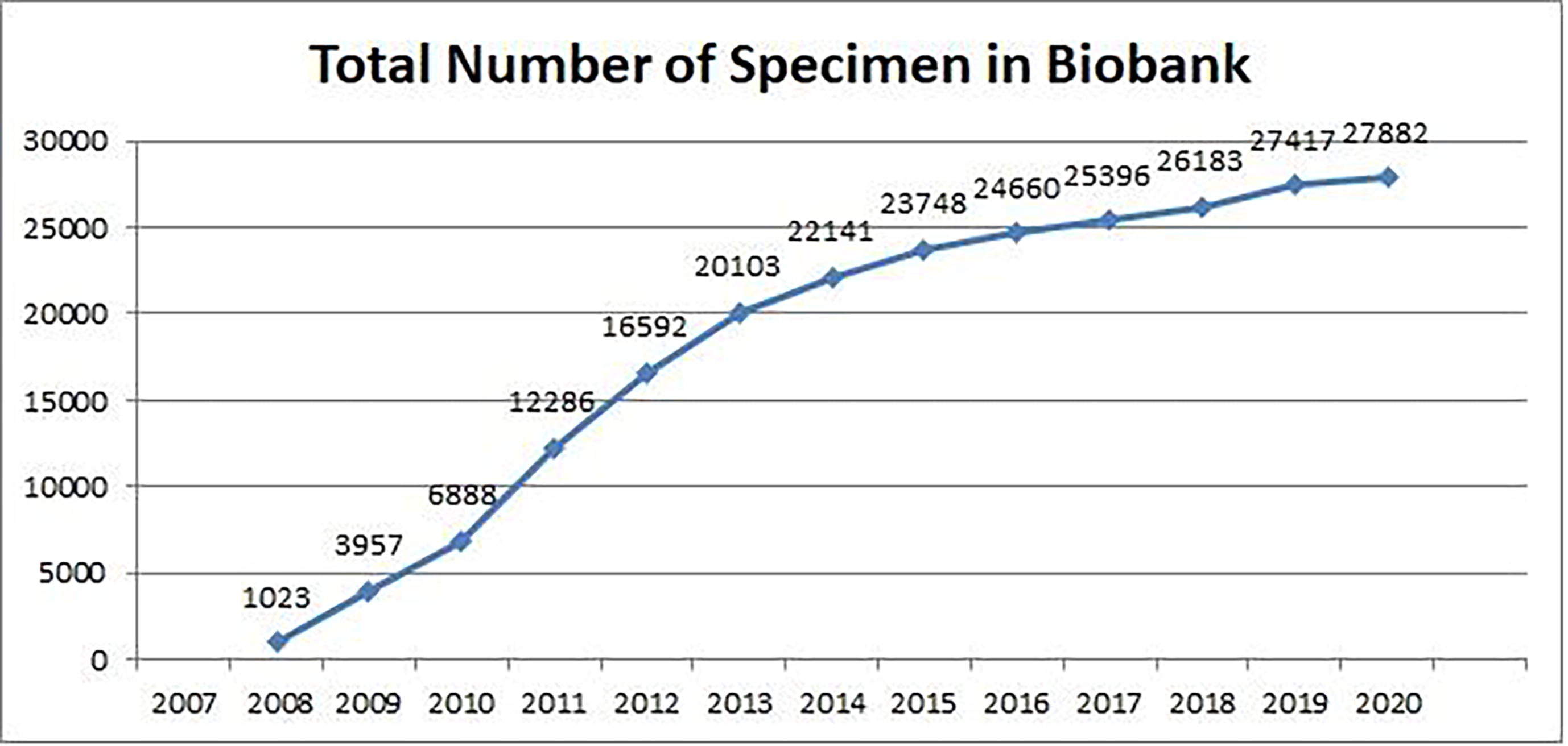
Over the past 13 years (2007–2020), we have collected a significantly high number of specimens (n = 25,396) and their derivatives (n = 27,882), with those numbers growing every year (Fig. 1, Table 1). Given the challenges of managing all that associated data, we established a disease-oriented biobank and developed Biosearch System, an in-house LIMS to better manage biobanks and store disease-associated genomic information.
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
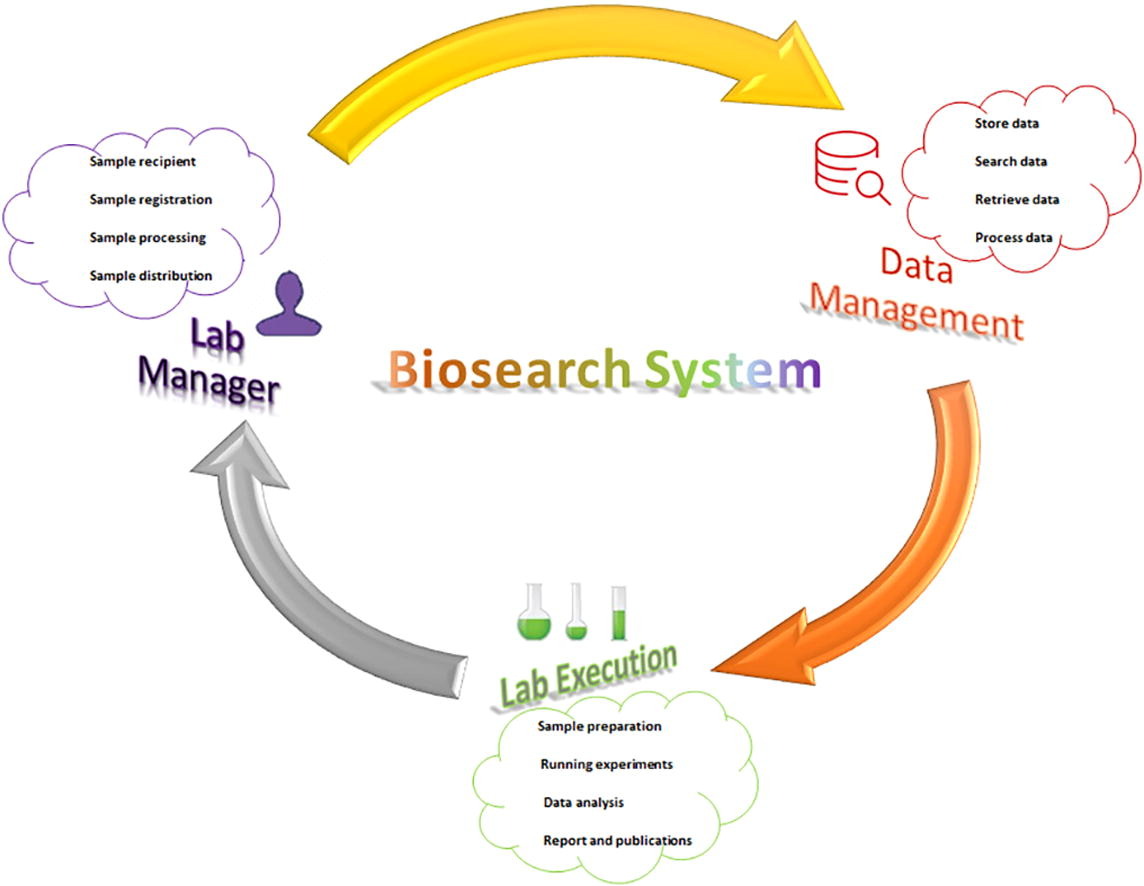
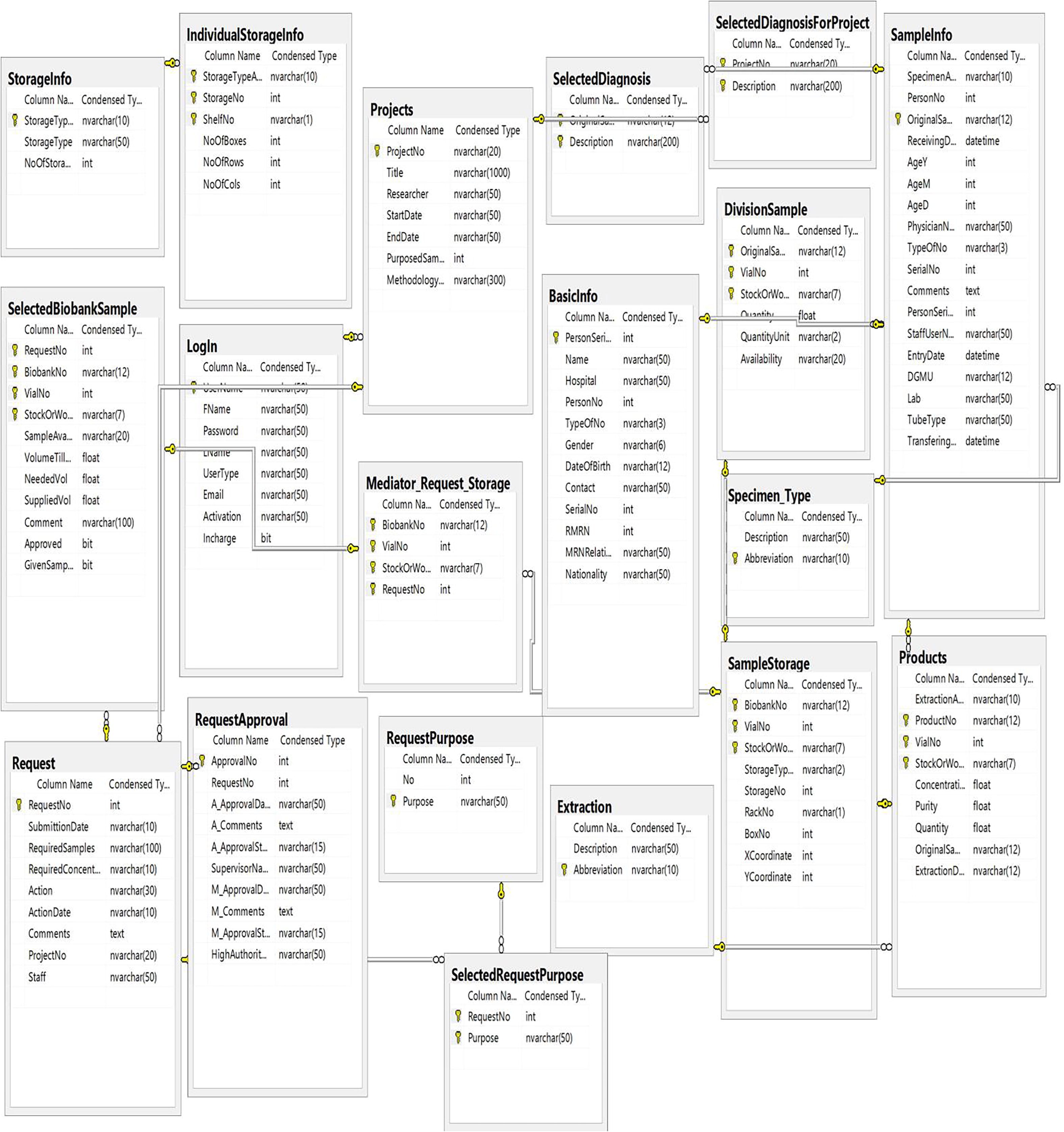
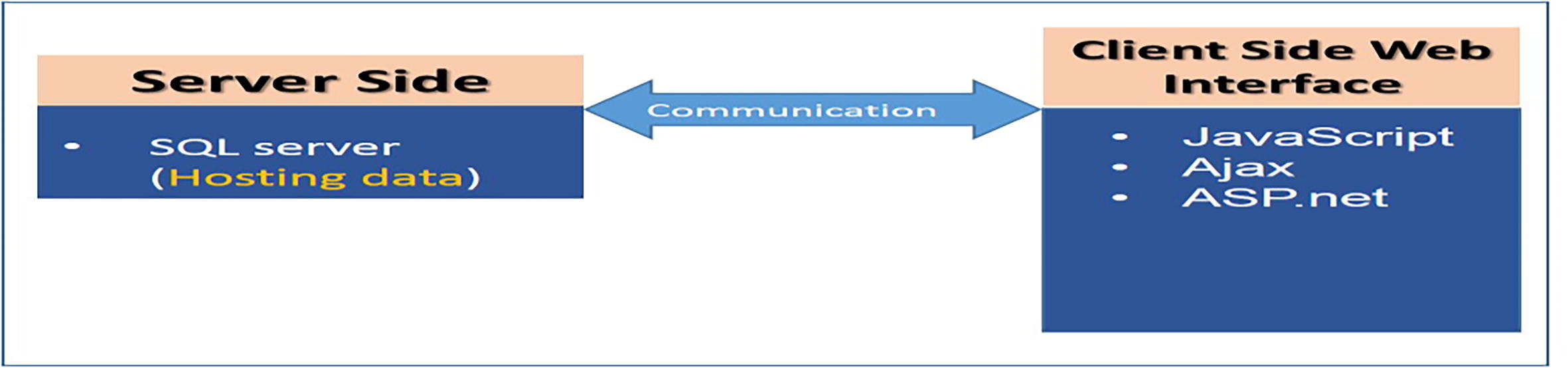
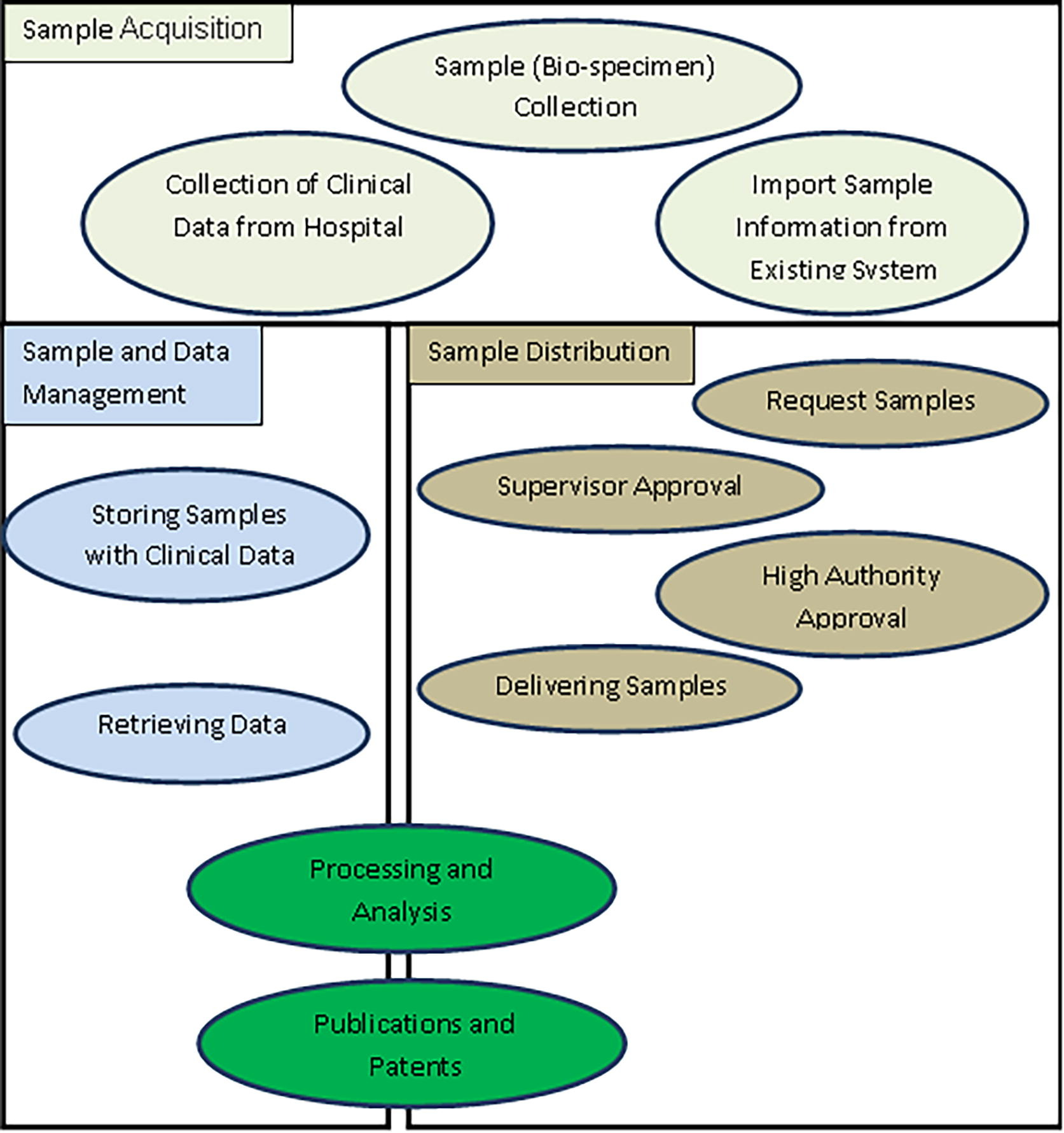
A prototype of an entity-relationship diagram was developed to depict the numerical relationship among different entities such as sample, storage condition, specimen request, approval system, project type, diagnosis, and disease type (Fig. 2). We established communication between the Biosearch System host server using SQL Server Management Studio and employed ASP.NET and JavaScript/Ajax toolkit for end users (Fig. 3). It was divided into three main parts: (i) data and specimen acquisition, (ii) data management, and (iii) specimen distribution/request (Fig. 4). It classifies the user into one of four groups (i.e., biobank technician, researcher, lab manager, and director), and hierarchal permission is given accordingly. The administrator of Biosearch System has the power to perform all functions, including addition of new users and granting permission according to hierarchy (Table 2).
|
|
|
| ||||||||||||
Biosearch System is supported by a data warehouse and query tools interface that can be used efficiently to search and short-list patients as per a particular criterion without disclosing their privacy. Based on evaluation form analysis, we found that it is a robust, secure, flexible, efficient, and user-friendly database. It proves robust as it is platform-independent, compatible with any version of Windows. Scripts are bug-free, so as to avoid hanging up the system. It proves secure since in addition to directing an appropriate permission strategy for different users and password protection, a King Abdulaziz University (KAU) network security strategy was also incorporated to transfer data safely, where each user has their own ID and password to access a specific computer, and the password is stored in a database in an encrypted way. Each machine that has access to the LIMS must have its own firewall and Trend Micro antivirus office scan agent provided by KAU IT deanship. The system performs backups at a regular interval (i.e., monthly) on KAU servers, so any loss of data from the system can be easily restored thereby saving the date from any disasters. The system proves flexible, as new features/modules can easily be added by authorized users to customize the LIMS according to laboratory and researcher requirements. It's also efficient, as it has been managing huge amounts of CEGMR's clinical and experimental data competently since 2014. Finally, the LIMS proves user-friendly, as simple icons, buttons, drop-down lists, etc. make it convenient for users to manage data and easily add or retrieve data using simple icons with multiple other available options.
Our measure of the performance and utility of the LIMS evaluates it well with the help of real-time feedback from users, who gave it an overall satisfaction score of 4.7 out of 5. The results of performance analysis of search, insert, update, and delete functions were also satisfactory (Table 3). Biosearch System is efficient in terms of the number of transactions and number of users it manages while maintaining the quality of data and permitting pliability in laboratory workflow.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
differs from other LIMS in its customized approach for CEGMR laboratory while remaining flexible enough to be adjusted for any changes in the future to suit the needs of projects and researchers, including the addition of equipment, procedures, and/or software to complement or improve workflow without changing the core code. For example, essential customization needed for research purposes is to link SMARTGENE software, a tool for the diagnostic lab, with Biosearch System and exchange data, with researchers' consent. It can be used for any laboratory (e.g., QC, R&D, and analytical service) with minor modifications, whereas most commercial software are a bit too rigid. Its flexibility extends to data quality through computerized calculations and statistics, automation of processes, and the minimization of manual lab tasks such as data entry. For instance, the system ensures data entry occurs in a standardized way using pre-defined drop-down lists for most processes.
Biosearch System efficiently maintains the quality of data and permits pliability in the workflow, ultimately synchronizing the biobank system of CEGMR/KAU. The current state of the CEGMR biobank is as follows:
- Infrastructure: The biobank is well equipped with liquid nitrogen cylinders (−196 °C), deep freezers (−80 °C and −20 °C), refrigerators, and UPS systems.
- Personnel: Presently, 14 dedicated staff work to ensure the smooth running of the biobank unit.
- Workflow: Patient specimen/sample management is fully functional and regularly handles the dispatch, collection, processing, and storage of thousands of specimens.
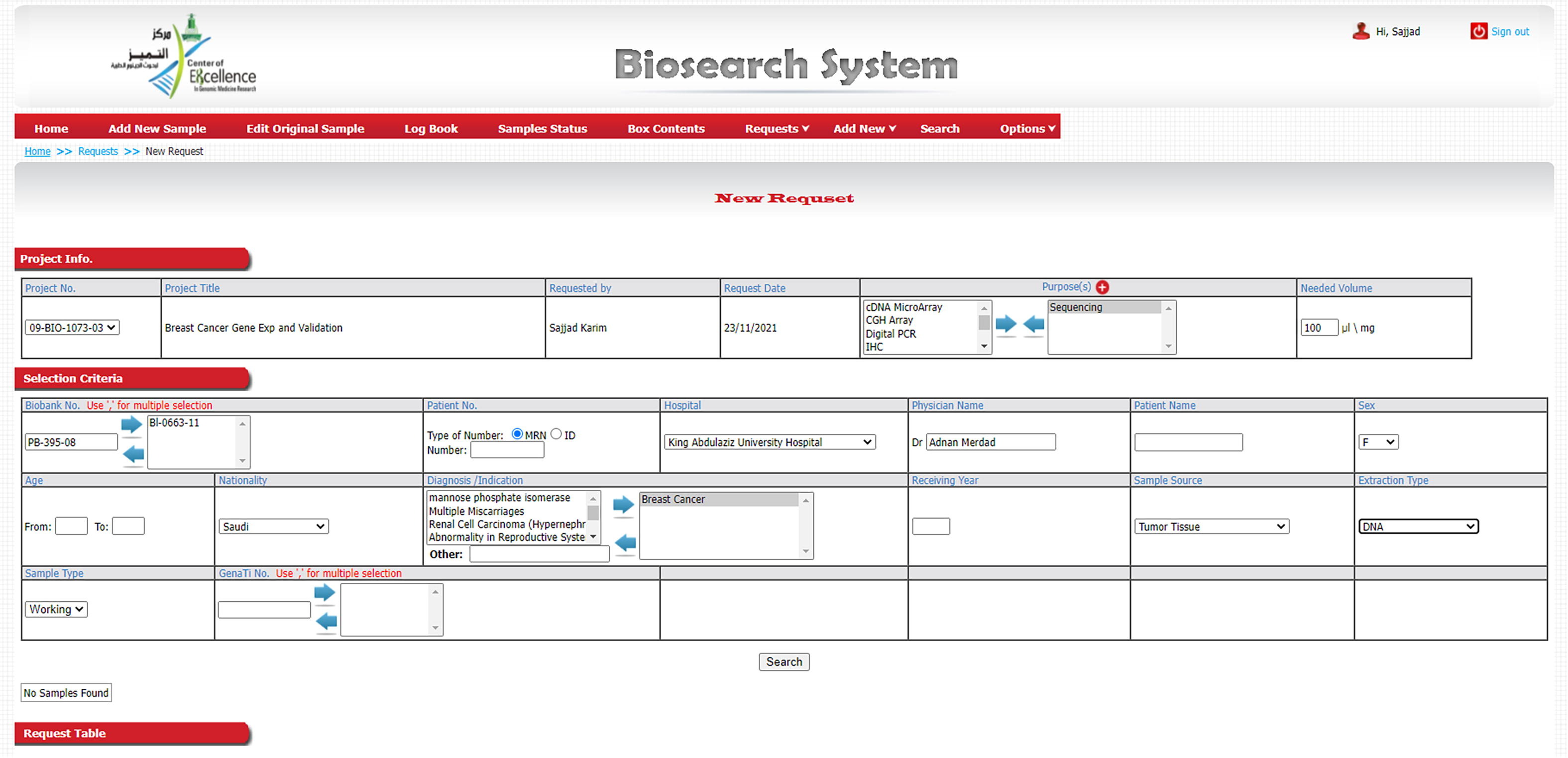
The LIMS can be accessed by authorized personnel using the username only (Fig. 5). Our clinical database contains the following clinico-pathological parameters: CEGMR code for patient, receiving date, hospital MRN number, name, date of birth, age, sex, nationality, disease, date of diagnosis, status, filing date, histology, sites, grade, size, lymph node status, invasion, margin status, immunohistochemical data, family history, medication, follow-up history, etc. Similarly, the LIMS' specimen/sample management functionality manages specimens and their derivative related information, including type of specimen, receiving date, extraction date, storage of specimen and its derivatives, quality and quantity record, handling request of researchers, distribution, and maintenance of specimen.
|
Identification of DEGs for breast cancer
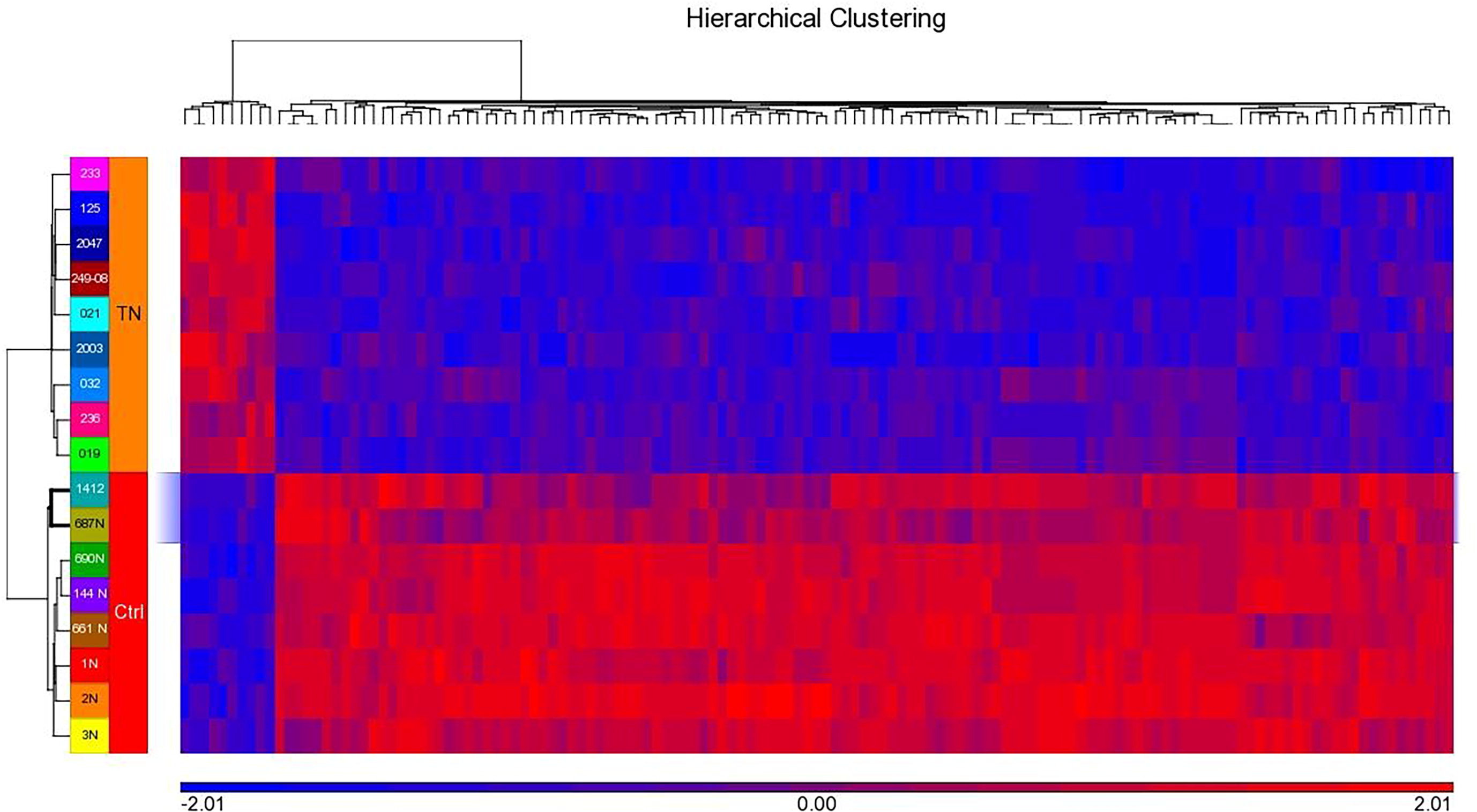
We conducted a genome-wide expression study to understand the molecular phenomenon leading to triple-negative breast cancer and found 1,181 differentially expressed genes, with p-value <0.05 and FC 2. The upregulated genes for TNBC were IFI6, LEF1, CCR8, FANCI, TRIM59, CASC5, and PLXNA3, while downregulated genes were ADH1B, LYVE1, ADH1C, ADH1B, FIGF, ADIPOQ, PLIN1, and LYVE1. Hierarchical clustering of the top 135 DEGs shows distinct pattern for genes in the LIMS' triple-negative and control samples (Fig. 6).
|
Pathways associated with triple-negative breast cancer
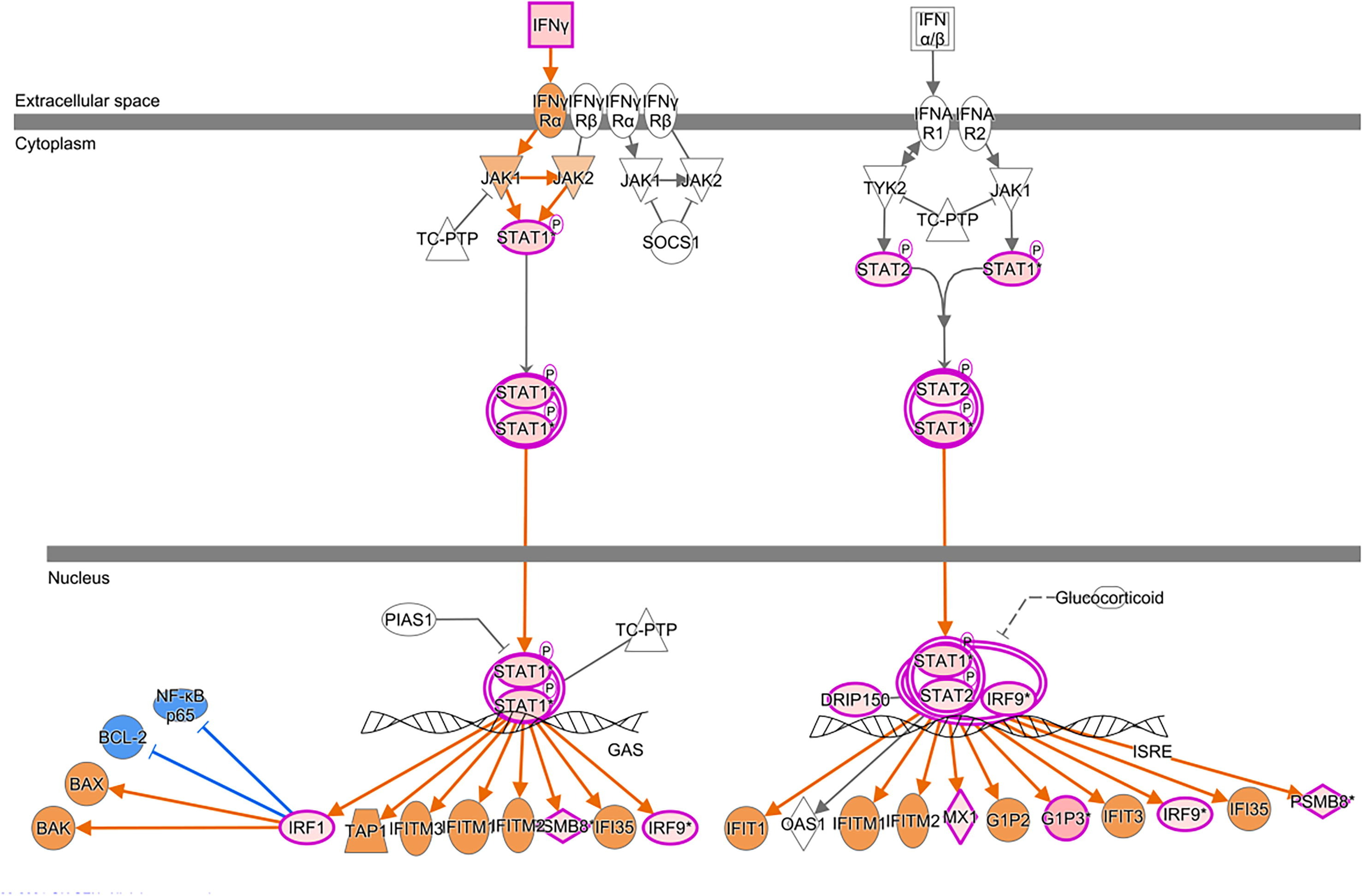
Molecular pathway analysis revealed more than 100 canonical pathways, where most activated pathways were (i) interferon signaling pathways (z-score = 2.646) with participating genes IFI6, FNG, IRF1, IRF9, MED14, MX1, PSMB8, STAT1, STAT2; and (ii) kinetochore metaphase signaling pathways (z-score = 2.138) with participating genes BUB1, BUB1B, CDK1, CENPL, KIF2C, KNL1, KNTC1, NUF2, PLK1, PPP1CA, PPP1R14B, PTTG1, RAD21, REC8. and TTK (Fig. 7).
|
Discussion
Today’s biobanks are much more than just sample repositories. They store a huge amount of clinical and experimental data related to specimens.[19] However, efficient data management is a bottleneck in the genomic medicine research process. It has been observed on many occasions that researchers face difficulties in data collection, maintenance, follow-up studies, and sometime even publishing the work without missing data.[15][16][17] Translational genomic research is based on a secure database and biobank system. A well-designed data management system provides the possibility of finding significant associations among stored information and facilitates diagnostics and therapeutics research.[20][21]
Our LIMS, Biosearch System, is used to collect, store, distribute and maintain specimen data, as well as its relevant clinical and experimental information to support ongoing research that can lead to the discovery of novel cancer biomarkers and beneficial therapeutic targets.[5][22][23][24][25][26][27][28][29][30][31] We used SQL Server Management Studio, Visual Basic, .NET, and the Ajax control kit to develop its interactive database, and ASP.NET for better code management, cleaner code structure, and faster web applications. The first prototype of Biosearch System was ready in 2010, and after feasibility testing and a successful trial run, we released the present final version for use within CEGMR/KAU, with minor modifications. The LIMS is a web-based tool accessible from both on and off campus. A high level of security allows only authorized researchers to access the database. Security threats such as hacking, virus attack, etc. were addressed without any significant compromise.[32][33][34][35] Bioethical safety was another major concern, which demanded proper care in establishing the biobank and developing the database.[6][7][36][37][38]
Biosearch System makes the job easier for everybody involved in research. First, clinicians and pathologists can include the clinico-pathological information of each provided specimen from hospitals/clinics through an online portal. Second, biobank staff can store and maintain the specimen and extracted derivatives easily, and finally, researchers can extract clinical information and request specific specimens/derivatives per the requirements of their ongoing project.[8] Presently our dataset is small, and we are collecting selected cases only, so the frequency of disease should not be used to represent society at large. However, the frequency of different type of cancer in our biobank is similar to our national cancer registry. In the future, the LIMS can pave the way for expanding this database in association with the Saudi cancer registry or any other national-level databases.
The LIMS was tested by CEGMR staff and was found satisfactory. The users were comfortable organizing all specimen/sample-related information and found it user-friendly with simple icons, buttons, drop-down lists, etc. To ensure the safety of data, KAU network security was utilized, and only users with approved permissions and sufficient supporting anti-virus tools were allowed to access the system. A periodic system backup strategy also ensured avoidance of any data loss due to unseen disaster. The LIMS is customized according to the actual activities and workflow in the CEGMR lab, while remaining flexible enough to adopt new modules to add more features in the future.
Presently, Biosearch System is not available as open-source software because of institutional policy, but we encourage interested researchers to request the code on an individual basis.
Conclusion and future work
Biosearch System is a user-friendly LIMS solution for managing biobank specimens and their associated clinical information to facilitate genomic medicine research, leading to the discovery of disease biomarkers and therapeutic targets. It is a flexible system, one that is able to be modified to meet specific workflows. Going forward, new features and modules to be added to the CEGMR/KAU implementation include a barcoding system, quality control system, and reagent purchasing system.
Acknowledgements
We would like to thank the Biobank and IT unit staff of Center of Excellence in Genomic Medicine Research, AZIZ Supercomputing facilities at High-Performance Computing Center, and Deanship of Scientific Research, King Abdulaziz University for their help and technical support.
Author contributions
Sajjad Karim: Conceptualization, Data curation, Fund acquisition, Investigation, Project Administration, Software, Supervision, Writing - original draft. Adnan Merdad: Conceptualization, Fund acquisition, Validation, Writing - review & editing. Saddig Jastaniah: Conceptualization, Fund acquisition, Validation, Writing - review & editing. Sudhir Kumar: Conceptualization, Fund acquisition, Supervision. Adel Abuzenadah: Conceptualization, Project Administration, Resources, Validation, Writing - review & editing. Mohammed Al-Qahtani: Conceptualization, Project Administration, Resources, Validation, Writing - review & editing. Mona Alkharaz: Data curation, Formal analysis, Investigation, Methodology. Zeenat Mirza: Data curation, Investigation, Methodology, Software, Visualization, Writing - original draft. Mahmood Rasool: Data curation, Investigation, Validation, Writing - review & editing. Hend Noureldin: Formal analysis. Heba Abusamra: Formal analysis. Nofe Alganmi: Methodology, Resources, Software, Supervision, Visualization, Writing - review & editing.
Funding
This study was funded by Deanship of Scientific Research, King Abdulaziz University (2-117-1434-HiCi).
Ethics approval and consent to participate
Ethical committee approved this study (Reference Number: 08-CEGMR-02-ETH) of CEGMR, KAU.
Availability of data and materials
Datasets (.CEL file) submitted to NCBI’s GEO, accession number GSE36295.
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Abbreviations, acronyms, and initialisms
CEGMR: Center of Excellence in Genomic Medicine Research
DEG: differentially expressed gene
FC: fold change
GUI: graphical user interface
IPA: Ingenuity Pathway Analysis
KAU: King Abdulaziz University
LIMS: laboratory information management system
MRN: medical record number
NGS: next-generation sequencing
PCA: principle component analysis
References
- ↑ Artene, Stefan-Alexandru; Ciurea, Marius Eugen; Purcaru, Stefana Oana; Tache, Daniela Elise; Tataranu, Ligia Gabriela; Lupu, Mihaela; Dricu, Anica (2013). "Biobanking in a Constantly Developing Medical World" (in en). The Scientific World Journal 2013: 1–5. doi:10.1155/2013/343275. ISSN 1537-744X. PMC PMC3794514. PMID 24174912. http://www.hindawi.com/journals/tswj/2013/343275/.
- ↑ Diamandis, Eleftherios P (1 December 2009). "Next-Generation Sequencing: A New Revolution in Molecular Diagnostics?" (in en). Clinical Chemistry 55 (12): 2088–2092. doi:10.1373/clinchem.2009.133389. ISSN 0009-9147. https://academic.oup.com/clinchem/article/55/12/2088/5631746.
- ↑ Fehniger, Thomas E.; Marko-Varga, György A. (7 January 2011). "Clinical Proteomics Today" (in en). Journal of Proteome Research 10 (1): 3–3. doi:10.1021/pr1012569. ISSN 1535-3893. https://pubs.acs.org/doi/abs/10.1021/pr1012569.
- ↑ Glenn, Travis C. (1 September 2011). "Field guide to next-generation DNA sequencers: FIELD GUIDE TO NEXT-GEN SEQUENCERS" (in en). Molecular Ecology Resources 11 (5): 759–769. doi:10.1111/j.1755-0998.2011.03024.x. https://onlinelibrary.wiley.com/doi/10.1111/j.1755-0998.2011.03024.x.
- ↑ 5.0 5.1 Merdad, Adnan; Karim, Sajjad; Schulten, Hans-Juergen; Dallol, Ashraf; Buhmeida, Abdelbaset; Al-Thubaity, Fatima; Gari, Mamdooh A.; Chaudhary, Adeel Ga et al. (1 March 2014). "Expression of matrix metalloproteinases (MMPs) in primary human breast cancer: MMP-9 as a potential biomarker for cancer invasion and metastasis". Anticancer Research 34 (3): 1355–1366. ISSN 1791-7530. PMID 24596383. https://pubmed.ncbi.nlm.nih.gov/24596383.
- ↑ 6.0 6.1 International Society for Biological and Environmental Repositories (ISBER) (1 April 2012). "2012 Best Practices for Repositories Collection, Storage, Retrieval, and Distribution of Biological Materials for Research International Society for Biological and Environmental Repositories" (in en). Biopreservation and Biobanking 10 (2): 79–161. doi:10.1089/bio.2012.1022. ISSN 1947-5535. http://www.liebertpub.com/doi/10.1089/bio.2012.1022.
- ↑ 7.0 7.1 Bredenoord, Annelien L.; Kroes, Hester Y.; Cuppen, Edwin; Parker, Michael; van Delden, Johannes J.M. (1 February 2011). "Disclosure of individual genetic data to research participants: the debate reconsidered" (in en). Trends in Genetics 27 (2): 41–47. doi:10.1016/j.tig.2010.11.004. https://linkinghub.elsevier.com/retrieve/pii/S0168952510002271.
- ↑ 8.0 8.1 Kang, Byunghak; Park, Jaesun; Cho, Sangyun; Lee, Meehee; Kim, Namhee; Min, Haesook; Lee, Sooyoun; Park, Ok et al. (2013). "Current Status, Challenges, Policies, and Bioethics of Biobanks" (in en). Genomics & Informatics 11 (4): 211. doi:10.5808/GI.2013.11.4.211. ISSN 1598-866X. PMC PMC3897848. PMID 24465232. http://genominfo.org/journal/view.php?doi=10.5808/GI.2013.11.4.211.
- ↑ Voegele, C.; Tavtigian, S.V.; de Silva, D.; Cuber, S.; Thomas, A.; Le Calvez-Kelm, F. (15 September 2007). "A Laboratory Information Management System (LIMS) for a high throughput genetic platform aimed at candidate gene mutation screening" (in en). Bioinformatics 23 (18): 2504–2506. doi:10.1093/bioinformatics/btm365. ISSN 1367-4803. https://academic.oup.com/bioinformatics/article-lookup/doi/10.1093/bioinformatics/btm365.
- ↑ Melo, Alexandre; Faria-Campos, Alessandra; DeLaat, Daiane; Keller, Rodrigo; Abreu, Vinícius; Campos, Sérgio (2010). "SIGLa: an adaptable LIMS for multiple laboratories" (in en). BMC Genomics 11 (Suppl 5): S8. doi:10.1186/1471-2164-11-S5-S8. ISSN 1471-2164. PMC PMC3045801. PMID 21210974. http://bmcgenomics.biomedcentral.com/articles/10.1186/1471-2164-11-S5-S8.
- ↑ Betsou, Fay; Rimm, David L.; Watson, Peter H.; Womack, Christopher; Hubel, Allison; Coleman, Robert A.; Horn, Liz; Terry, Sharon F. et al. (1 June 2010). "What Are the Biggest Challenges and Opportunities for Biorepositories in the Next Three to Five Years?" (in en). Biopreservation and Biobanking 8 (2): 81–88. doi:10.1089/bio.2010.8210. ISSN 1947-5535. http://www.liebertpub.com/doi/10.1089/bio.2010.8210.
- ↑ Hewitt, Robert E (1 January 2011). "Biobanking: the foundation of personalized medicine" (in en). Current Opinion in Oncology 23 (1): 112–119. doi:10.1097/CCO.0b013e32834161b8. ISSN 1040-8746. https://journals.lww.com/00001622-201101000-00019.
- ↑ 13.0 13.1 Huang, Y. Wayne; Arkin, Adam P.; Chandonia, John-Marc (1 February 2011). "WIST: toolkit for rapid, customized LIMS development" (in en). Bioinformatics 27 (3): 437–438. doi:10.1093/bioinformatics/btq676. ISSN 1460-2059. PMC PMC3031042. PMID 21258060. https://academic.oup.com/bioinformatics/article-lookup/doi/10.1093/bioinformatics/btq676.
- ↑ Riegman, Peter H.J.; Morente, Manuel M.; Betsou, Fay; de Blasio, Pasquale; Geary, Peter (1 October 2008). "Biobanking for better healthcare" (in en). Molecular Oncology 2 (3): 213–222. doi:10.1016/j.molonc.2008.07.004. PMC PMC5527804. PMID 19383342. http://doi.wiley.com/10.1016/j.molonc.2008.07.004.
- ↑ 15.0 15.1 Greely, Henry T. (1 September 2007). "The Uneasy Ethical and Legal Underpinnings of Large-Scale Genomic Biobanks" (in en). Annual Review of Genomics and Human Genetics 8 (1): 343–364. doi:10.1146/annurev.genom.7.080505.115721. ISSN 1527-8204. https://www.annualreviews.org/doi/10.1146/annurev.genom.7.080505.115721.
- ↑ 16.0 16.1 Kauffmann, Francine; Cambon-Thomsen, Anne (21 May 2008). "Tracing Biological Collections: Between Books and Clinical Trials" (in en). JAMA 299 (19): 2316. doi:10.1001/jama.299.19.2316. ISSN 0098-7484. http://jama.jamanetwork.com/article.aspx?doi=10.1001/jama.299.19.2316.
- ↑ 17.0 17.1 Minamikumo, M. (2012). "Current status and future of biobanks". Policy Inst. News 36: 15–21.
- ↑ Prilusky, Jaime; Oueillet, Eric; Ulryck, Nathalie; Pajon, Anne; Bernauer, Julie; Krimm, Isabelle; Quevillon-Cheruel, Sophie; Leulliot, Nicolas et al. (1 June 2005). "HalX : an open-source LIMS (Laboratory Information Management System) for small- to large-scale laboratories". Acta Crystallographica Section D Biological Crystallography 61 (6): 671–678. doi:10.1107/S0907444905001290. ISSN 0907-4449. http://scripts.iucr.org/cgi-bin/paper?S0907444905001290.
- ↑ Calleros, L.; Cortés, M.A.; Luengo, A. et al. (2012). "Puesta en marcha de una plataforma de proceso, almacenamiento y gestión de muestras clínicas: organización y desarrollo del Biobanco de REDinREN". Nefrología 32 (1): 28–34. doi:10.3265/Nefrologia.pre2011.Oct.11121. ISSN 2013-2514. PMID 22294002.
- ↑ P. Lemmon, Vance; Jia, Yuanyuan; Shi, Yan; Douglas Holbrook, S.; L. Bixby, John; Buchser, William (1 November 2011). "Challenges in Small Screening Laboratories: Implementing an On-Demand Laboratory Information Management System" (in en). Combinatorial Chemistry & High Throughput Screening 14 (9): 742–748. doi:10.2174/138620711796957161. PMC PMC3289069. PMID 21631415. http://www.eurekaselect.com/openurl/content.php?genre=article&issn=1386-2073&volume=14&issue=9&spage=742.
- ↑ Zerhouni, Elias A. (21 September 2005). "US Biomedical Research: Basic, Translational, and Clinical Sciences" (in en). JAMA 294 (11): 1352. doi:10.1001/jama.294.11.1352. ISSN 0098-7484. http://jama.jamanetwork.com/article.aspx?doi=10.1001/jama.294.11.1352.
- ↑ Merdad, Adnan; Karim, Sajjad; Schulten, Hans-Juergen; Jayapal, Manikandan; Dallol, Ashraf; Buhmeida, Abdelbaset; Al-Thubaity, Fatima; GariI, Mamdooh A et al. (1 December 2015). "Transcriptomics profiling study of breast cancer from Kingdom of Saudi Arabia revealed altered expression of Adiponectin and Fatty Acid Binding Protein4: Is lipid metabolism associated with breast cancer?" (in en). BMC Genomics 16 (S1): S11. doi:10.1186/1471-2164-16-S1-S11. ISSN 1471-2164. PMC PMC4315151. PMID 25923423. https://bmcgenomics.biomedcentral.com/articles/10.1186/1471-2164-16-S1-S11.
- ↑ Karim, Sajjad; Al-Maghrabi, Jaudah A.; Farsi, Hasan M. A.; Al-Sayyad, Ahmad J.; Schulten, Hans-Juergen; Buhmeida, Abdelbaset; Mirza, Zeenat; Al-boogmi, Alaa A. et al. (1 September 2016). "Cyclin D1 as a therapeutic target of renal cell carcinoma- a combined transcriptomics, tissue microarray and molecular docking study from the Kingdom of Saudi Arabia" (in en). BMC Cancer 16 (S2): 741. doi:10.1186/s12885-016-2775-2. ISSN 1471-2407. PMC PMC5073805. PMID 27766950. http://bmccancer.biomedcentral.com/articles/10.1186/s12885-016-2775-2.
- ↑ Karim, Sajjad; Malik, Imran Riaz; Nazeer, Quratulain; Zaheer, Ahmad; Farooq, Muhammad; Mahmood, Nasir; Malik, Arif; Asif, Muhammad et al. (1 September 2019). "Molecular analysis of V617F mutation in Janus kinase 2 gene of breast cancer patients" (in en). Saudi Journal of Biological Sciences 26 (6): 1123–1128. doi:10.1016/j.sjbs.2019.08.002. PMC PMC6733781. PMID 31516339. https://linkinghub.elsevier.com/retrieve/pii/S1319562X19301342.
- ↑ Mirza, Zeenat; Schulten, Hans-Juergen; Farsi, Hasan Ma; Al-Maghrabi, Jaudah A.; Gari, Mamdooh A.; Chaudhary, Adeel Ga; Abuzenadah, Adel M.; Al-Qahtani, Mohammed H. et al. (1 April 2014). "Impact of S100A8 expression on kidney cancer progression and molecular docking studies for kidney cancer therapeutics". Anticancer Research 34 (4): 1873–1884. ISSN 1791-7530. PMID 24692722. https://pubmed.ncbi.nlm.nih.gov/24692722.
- ↑ Mirza, Zeenat; Schulten, Hans-Juergen; Farsi, Hasan Ma; Al-Maghrabi, Jaudah A.; Gari, Mamdooh A.; Chaudhary, Adeel Ga; Abuzenadah, Adel M.; Al-Qahtani, Mohammed H. et al. (19 March 2015). Vadgama, Jaydutt V.. ed. "Molecular Interaction of a Kinase Inhibitor Midostaurin with Anticancer Drug Targets, S100A8 and EGFR: Transcriptional Profiling and Molecular Docking Study for Kidney Cancer Therapeutics" (in en). PLOS ONE 10 (3): e0119765. doi:10.1371/journal.pone.0119765. ISSN 1932-6203. PMC PMC4366044. PMID 25789858. https://dx.plos.org/10.1371/journal.pone.0119765.
- ↑ Rasool, Mahmood; Pushparaj, Peter Natesan; Mirza, Zeenat; Imran Naseer, Muhammad; Abusamra, Heba; Alquaiti, Maha; Shaabad, Manal; Sibiany, Abdulrahman Mohamed Saeed et al. (1 January 2020). "Array comparative genomic hybridization based identification of key genetic alterations at 2p21-p16.3 (MSH2, MSH6, EPCAM), 3p23-p14.2 (MLH1), 7p22.1 (PMS2) and 1p34.1-p33 (MUTYH) regions in hereditary non polyposis colorectal cancer (Lynch syndrome) in the Kingdom of Saudi Arabia" (in en). Saudi Journal of Biological Sciences 27 (1): 157–162. doi:10.1016/j.sjbs.2019.06.012. PMC PMC6933242. PMID 31889830. https://linkinghub.elsevier.com/retrieve/pii/S1319562X19301093.
- ↑ Rasool, Mahmood; Natesan Pushparaj, Peter; Buhmeida, Abdelbaset; Karim, Sajjad (1 October 2021). "Mutational spectrum of BRAF gene in colorectal cancer patients in Saudi Arabia" (in en). Saudi Journal of Biological Sciences 28 (10): 5906–5912. doi:10.1016/j.sjbs.2021.06.048. PMC PMC8459112. PMID 34588906. https://linkinghub.elsevier.com/retrieve/pii/S1319562X21005179.
- ↑ Schulten, Hans-Juergen; Hussein, Deema; Al-Adwani, Fatima; Karim, Sajjad; Al-Maghrabi, Jaudah; Al-Sharif, Mona; Jamal, Awatif; Bakhashab, Sherin et al. (1 October 2016). "Microarray expression profiling identifies genes, including cytokines, and biofunctions, as diapedesis, associated with a brain metastasis from a papillary thyroid carcinoma". American Journal of Cancer Research 6 (10): 2140–2161. ISSN 2156-6976. PMC 5088282. PMID 27822408. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5088282/.
- ↑ Subhi, Ohoud; Schulten, Hans-Juergen; Bagatian, Nadia; Al-Dayini, Roa'a; Karim, Sajjad; Bakhashab, Sherin; Alotibi, Reem; Al-Ahmadi, Alaa et al. (30 June 2020). Soares, Paula. ed. "Genetic relationship between Hashimoto`s thyroiditis and papillary thyroid carcinoma with coexisting Hashimoto`s thyroiditis" (in en). PLOS ONE 15 (6): e0234566. doi:10.1371/journal.pone.0234566. ISSN 1932-6203. PMC PMC7326236. PMID 32603365. https://dx.plos.org/10.1371/journal.pone.0234566.
- ↑ Sultan, Samar; Ahmed, Farid; Bajouh, Osama; Schulten, Hans-Juergen; Bagatian, Nadia; Al-Dayini, Roaa; Subhi, Ohoud; Karim, Sajjad et al. (2021). "Alterations of transcriptome expression, cell cycle, and mitochondrial superoxide reveal foetal endothelial dysfunction in Saudi women with gestational diabetes mellitus" (in en). Endocrine Journal 68 (9): 1067–1079. doi:10.1507/endocrj.EJ21-0189. ISSN 0918-8959. https://www.jstage.jst.go.jp/article/endocrj/68/9/68_EJ21-0189/_article.
- ↑ Bjugn, Roger; Hansen, Jarle (1 August 2013). "Learning by Erring: Fire!" (in en). Biopreservation and Biobanking 11 (4): 202–205. doi:10.1089/bio.2013.0020. ISSN 1947-5535. http://www.liebertpub.com/doi/10.1089/bio.2013.0020.
- ↑ Mintzer, Joseph L.; Kronenthal, Courtney J.; Kelly, Victoria; Seneca, Michael; Butler, Gary; Fecenko-Tacka, Karen; Altamuro, Donna; Madore, Steven J. (1 August 2013). "Preparedness for a Natural Disaster: How Coriell Planned for Hurricane Sandy" (in en). Biopreservation and Biobanking 11 (4): 216–220. doi:10.1089/bio.2013.0035. ISSN 1947-5535. http://www.liebertpub.com/doi/10.1089/bio.2013.0035.
- ↑ Rogers, J.; Carolin, T.; Vaught, J.; Compton, C. (1 June 2011). "Biobankonomics: A Taxonomy for Evaluating the Economic Benefits of Standardized Centralized Human Biobanking for Translational Research" (in en). JNCI Monographs 2011 (42): 32–38. doi:10.1093/jncimonographs/lgr010. ISSN 1052-6773. https://academic.oup.com/jncimono/article-lookup/doi/10.1093/jncimonographs/lgr010.
- ↑ Simeon-Dubach, Daniel; Zaayenga, Andy; Henderson, Marianne K. (1 June 2013). "Disaster and Recovery: The Importance of Risk Assessment and Contingency Planning for Biobanks" (in en). Biopreservation and Biobanking 11 (3): 133–134. doi:10.1089/bio.2013.1131. ISSN 1947-5535. http://www.liebertpub.com/doi/10.1089/bio.2013.1131.
- ↑ Hansson, M G (1 January 2009). "Ethics and biobanks" (in en). British Journal of Cancer 100 (1): 8–12. doi:10.1038/sj.bjc.6604795. ISSN 0007-0920. PMC PMC2634684. PMID 19034276. http://www.nature.com/articles/6604795.
- ↑ McGuire, Amy L.; Caulfield, Timothy; Cho, Mildred K. (1 February 2008). "Research ethics and the challenge of whole-genome sequencing" (in en). Nature Reviews Genetics 9 (2): 152–156. doi:10.1038/nrg2302. ISSN 1471-0056. PMC PMC2225443. PMID 18087293. http://www.nature.com/articles/nrg2302.
- ↑ Rotimi, Charles N; Marshall, Patricia A (2010). "Tailoring the process of informed consent in genetic and genomic research" (in en). Genome Medicine 2 (3): 20. doi:10.1186/gm141. ISSN 1756-994X. PMC PMC2873798. PMID 20346094. http://genomemedicine.biomedcentral.com/articles/10.1186/gm141.
Notes
This presentation is faithful to the original, with minor changes to presentation; grammar and spelling required more cleanup for improved readability. In some cases important information was missing from the references, and that information was added. The original article listed references in alphabetical order; this version lists them in order of appearance, by design.