Journal:Ten simple rules for managing laboratory information
| Full article title | Ten simple rules for managing laboratory information |
|---|---|
| Journal | PLoS Computational Biology |
| Author(s) | Berezin, Casey-Tyler; Aguilera, Luis U.; Billerbeck, Sonja; Bourne, Philip E.; Densmore, Douglas; Freemont, Paul; Gorochowski, Thomas E.; Hernandez, Sarah I.; Hillson, Nathan J.; King, Connor R.; Köpke, Michael; Ma, Shuyi; Miller, Katie M.; Moon, Tae Seok; Moore, Jason H.; Munsky, Brian; Myers, Chris J.; Nicholas, Dequina A.; Peccoud, Samuel J.; Zhou, Wen; Peccoud, Jean |
| Author affiliation(s) | Colorado State University, University of Groningen, University of Virginia, Boston University, Imperial College, University of Bristol, Lawrence Berkeley National Laboratory, US Department of Energy Agile BioFoundry, US Department of Energy Joint BioEnergy Institute, LanzaTech, University of Washington Medicine, Washington University in St. Louis, Cedars-Sinai Medical Centet, University of Colorado Boulder, University of California Irvine - Irvine |
| Primary contact | Email: jean dot peccoud at colostate dot edu |
| Year published | 2023 |
| Volume and issue | 19(12) |
| Article # | e1011652 |
| DOI | 10.1371/journal.pcbi.1011652 |
| ISSN | 1553-7358 |
| Distribution license | Creative Commons CC0 1.0 Universal |
| Website | https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1011652 |
| Download | https://journals.plos.org/ploscompbiol/article/file?id=10.1371/journal.pcbi.1011652&type=printable (PDF) |
Abstract
Information is the cornerstone of research, from experimental data/metadata and computational processes to complex inventories of reagents and equipment. These 10 simple rules discuss best practices for leveraging laboratory information management systems (LIMS) to transform this large information load into useful scientific findings.
Keywords: laboratory information management, laboratory management, laboratory information management systems, LIMS, computational biology, mathematical modeling, transdisciplinary research
Introduction
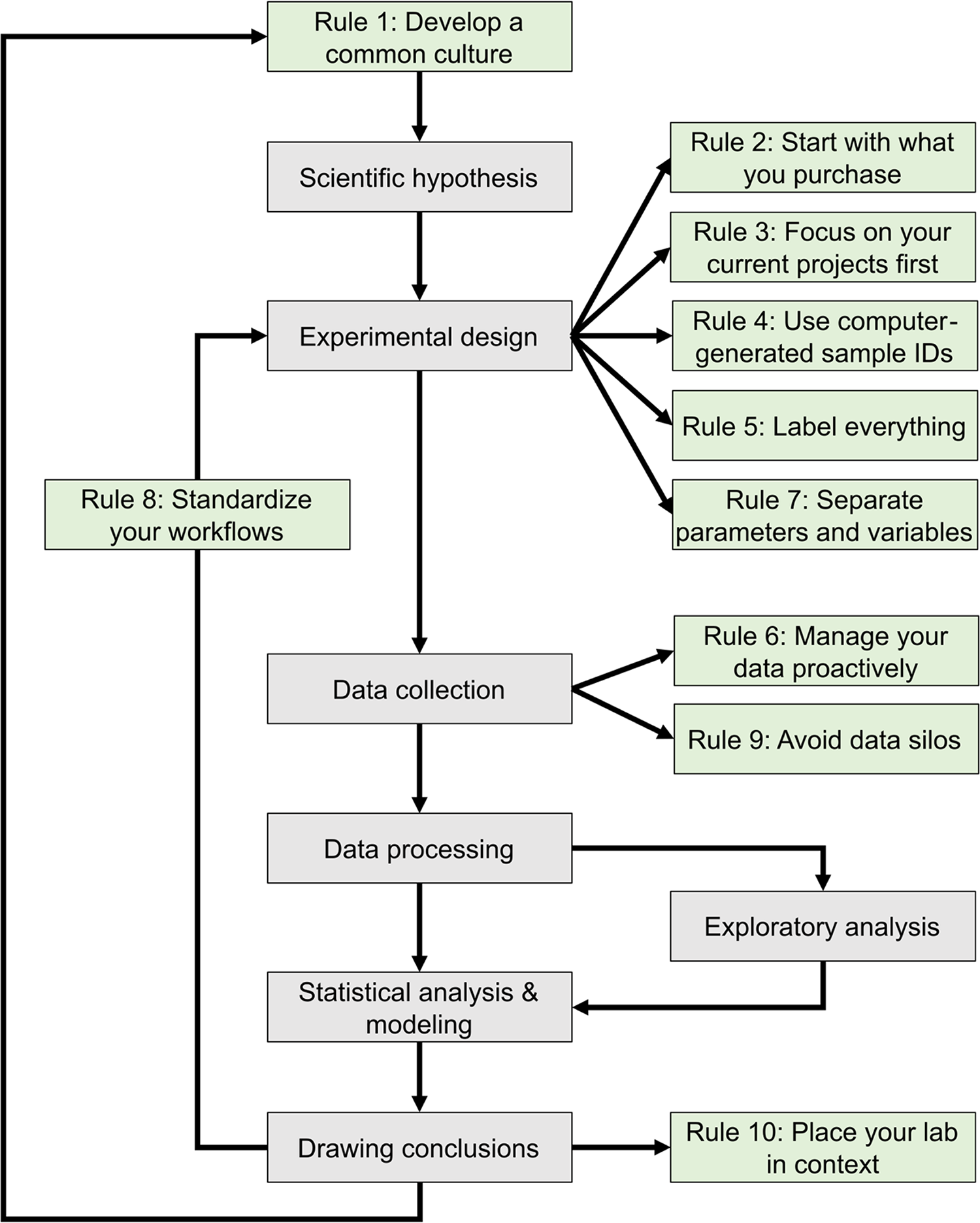
The development of mathematical models that can predict the properties of biological systems is the holy grail of computational biology. [1,2] Such models can be used to test biological hypotheses [3], quantify the risk of developing diseases [3], guide the development of biomanufactured products [4], engineer new systems meeting user-defined specifications, and much more. [4,5] Irrespective of a model’s application and the conceptual framework used to build it, the modeling process proceeds through a common iterative workflow. A model is first evaluated by fitting its parameters such that its behavior matches experimental data. Models that fit previous observations are then further validated by comparing the model predictions with the results of new observations that are outside the scope of the initial dataset (Fig. 1).
|
Historically, the collection of experimental data and the development of mathematical models were performed by different scientific communities. [6] Computational biologists had little control over the nature and quality of the data they could access. With the emergence of systems biology and synthetic biology, the boundary between experimental and computational biology has become increasingly blurred. [6] Many laboratories and junior scientists now have expertise in both producing and analyzing large volumes of digital data due to high-throughput workflows and an ever-expanding collection of digital instruments. [7] In this context, it is critically important to properly organize the exponentially growing volumes of experimental data to ensure they can support the development of models that can guide the next round of experiments. [8]
We are a group of scientists representing a broad range of scientific specialties, from clinical research to industrial biotechnology. Collectively, we have expertise in experimental biology, data science, and mathematical modeling. Some of us work in academia, while others work in industry. We have all faced the challenges of keeping track of laboratory operations to produce high-quality data suitable for analysis. We have experience using a variety of tools, including spreadsheets, open-source software, homegrown databases, and commercial solutions to manage our data. Irreproducible experiments, projects that failed to meet their goals, datasets we collected but never managed to analyze, and freezers full of unusable samples have taught us the hard way lessons that have led to these 10 simple rules for managing laboratory information.
This journal has published several sets of rules regarding best practices in overall research design [9,10], as well as the computational parts of research workflows, including data management [11–13] and software development practices. [14–16] The purpose of these 10 rules (Fig. 1) is to guide the development and configuration of laboratory information management systems (LIMS). LIMS typically offer electronic laboratory notebook (ELN), inventory, workflow planning, and data management features, allowing users to connect data production and data analysis to ensure that useful information can be extracted from experimental data and increase reproducibility. [17,18] These rules can also be used to develop training programs and lab management policies. Although we all agree that applying these rules increases the value of the data we produce in our laboratories, we also acknowledge that enforcing them is challenging. It relies on the successful integration of effective software tools, training programs, lab management policies, and the will to abide by these policies. Each lab must find the most effective way to adopt these rules to suit their unique environment.
Rule 1: Develop a common culture
Data-driven research projects generally require contributions from multiple stakeholders with complementary expertise. The success of these projects depends on entire teams developing a common vision of the projects' objectives and the approaches to be used. [19–21] Interdisciplinary teams, in particular, must establish a common language, as well as mutual expectations for experimental and publication timelines. [19] Unless the team develops a common culture, one stakeholder group can drive the project and impose its vision on the other groups. Although interdisciplinary (i.e., wet-lab and computational) training is becoming more common in academia, it is not unusual for experimentalists to regard data analysis as a technique they can acquire simply by hiring a student with computer programming skills. In a corporate environment, research informatics is often part of the information technology group whose mission is to support scientists who drive the research agenda. In both situations, the research agenda is driven by stakeholders who are unlikely to produce the most usable datasets because they lack sufficient understanding of data modeling. [20] Perhaps less frequently, there is also the situation where the research agenda is driven by people with expertise in data analysis. Because they may not appreciate the subtleties of experimental methods, they may find it difficult to engage experimentalists in collaborations aimed at testing their models. [20] Alternatively, their research may be limited to the analysis of disparate sets of previously published datasets. [19] Thus, interdisciplinary collaboration is key to maximizing the insights you gain from your data.
The development of a common culture, within a single laboratory or across interdisciplinary research teams, must begin with a thorough onboarding process for each member regarding general lab procedures, research goals, and individual responsibilities and expectations. [21,22] Implementing a LIMS requires perseverance by users, thus a major determinant of the success of a LIMS is whether end-users are involved in the culture development process. [17,23] When the input and suggestions of end-users are considered, they are more likely to engage with and apply good practices to the LIMS on a daily basis. [23] The long-term success of research endeavors then requires continued training and reevaluation of project goals and success (Fig.1). [19,21]
These 10 simple rules apply to transdisciplinary teams that have developed a common culture allowing experimentalists to gain a basic understanding of the modeling process and modelers to have some familiarity with the experimental processes generating the data they will analyze. [19] Teams that lack a common vision of data-driven research are encouraged to work toward acquiring this common vision through frequent communication and mutual goal setting. [19,20] Discussing these 10 simple rules in group meetings may aid in initiating this process.
Rule 2: Start with what you purchase
All the data produced by your lab are derived from things you have purchased, including supplies (consumables), equipment, and contract manufactured reagents, such as oligonucleotides or synthetic genes. In many cases, data (and metadata) on items in your inventory may be just as important as experimentally derived data, and as such, should be managed according to the FAIR (findable, accessible, interoperable, and reusable) principles. [24] Assembling an inventory of supplies and equipment with their associated locations can be handled in a few weeks by junior personnel without major interruption of laboratory operations, although establishing a thorough inventory may be more difficult and time-consuming for smaller labs with fewer members. Nevertheless, managing your lab inventory provides an immediate return on investment by positively impacting laboratory operations in several ways. People can quickly find the supplies and equipment they need to work, supplies are ordered with appropriate advance notice to minimize work stoppage, and data variation is reduced due to standardized supplies and the ability to track lot numbers easily (Fig. 1). [17,25,26]
Many labs still use Excel to keep track of inventory despite the existence of several more sophisticated databases and LIMS (e.g., Benchling, Quartzy, GenoFAB, LabWare, LabVantage, and TeselaGen). [25] These can facilitate real-time inventory tracking unlike a static document, increasing the findability and accessibility of inventory data. While some systems are specialized for certain types of inventories (e.g., animal colonies or frozen reagents), others are capable of tracking any type of reagent or item imaginable. [25] When considering what items to keep track of, there are three main considerations: expiration, maintenance, and ease of access.
Most labs manage their supplies through periodic cleanups of the lab, during which they sort through freezers, chemical cabinets, and other storage areas; review their contents; and dispose of supplies that are past their expiration date or are no longer useful. By actively tracking expiration dates and reagent use in a LIMS, you can decrease the frequency of such cleanups since the LIMS will alert users when expiration dates are approaching or when supplies are running low. This can prevent costly items from being wasted because they are expired or forgotten, and furthermore, the cost of products can be tracked and used to inform which experiments are performed.
LIMS can also support the use and service of key laboratory equipment. User manuals, service dates, warranties, and other identifying information can be attached directly to the equipment record, which allows for timely service and maintenance of the equipment. Adding equipment to the inventory can also prevent accidental losses in shared spaces where it is easy for people to borrow equipment without returning it. The label attached to the equipment (see Rule 5, later) acts as an indication of ownership that limits the risk of ownership confusion when almost identical pieces of equipment are owned by neighboring laboratories. As the laboratory inventory should focus on larger, more expensive equipment and supplies, inexpensive and easily obtained equipment (i.e., office supplies) may not need to be inventoried. An additional benefit of inventory management in a LIMS is the ability to create a record connecting specific equipment and supplies to specific people and projects, which can be used to detect potential sources of technical bias and variability (see Rules 4 and 5, later).
Rule 3: Focuson your current projects first
References
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added. To more easily differentiate footnotes from references, the original footnotes (which were numbered) were updated to use lowercase letters. Most footnotes referencing web pages were turned into proper citations.