Journal:Electronic tools in clinical laboratory diagnostics: Key examples, limitations, and value in laboratory medicine
| Full article title | Electronic tools in clinical laboratory diagnostics: Key examples, limitations, and value in laboratory medicine |
|---|---|
| Journal | Journal of Laboratory Medicine |
| Author(s) | Bohn, Mary K.; Fabiano, Giulia F.; Adeli, Khosrow |
| Author affiliation(s) | The Hospital for Sick Children, University of Toronto |
| Primary contact | khosrow dot adeli at sickkids dot ca |
| Year published | 2021 |
| Volume and issue | 45(6) |
| Page(s) | 319–24 |
| DOI | 10.1515/labmed-2021-0114 |
| ISSN | 2567-9430 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://www.degruyter.com/document/doi/10.1515/labmed-2021-0114/html |
| Download | https://www.degruyter.com/document/doi/10.1515/labmed-2021-0114/pdf (PDF) |
Abstract
Electronic tools in clinical laboratory diagnostics can assist laboratory professionals, clinicians, and patients in medical diagnostic management and laboratory test interpretation. With increasing implementation of electronic health records (EHRs) and laboratory information systems (LIS) worldwide, there is increasing demand for well-designed and evidence-based electronic resources. Both complex data-driven and simple interpretative electronic healthcare tools are currently available to improve the integration of clinical and laboratory information towards a more patient-centered approach to medicine. Several studies have reported positive clinical impact of electronic healthcare tool implementation in clinical laboratory diagnostics, including in the management of neonatal bilirubinemia, cardiac disease, and nutritional status. As patients have increasing access to their medical laboratory data, it is essential that accessible electronic healthcare tools are evidence-based and user-friendly for individuals of varying digital and medical literacy. Indeed, studies suggest electronic healthcare tool development processes significantly lack the involvement of relevant healthcare professionals and often present misinformation, including erroneous calculation algorithms or inappropriate interpretative recommendations.
The current review provides an overview of the utility of available electronic healthcare tools in clinical laboratory diagnostics and critically reviews potential limitations and benefits of their clinical implementation. The Canadian Laboratory Initiative on Pediatric Reference Intervals (CALIPER) online database is also detailed as an example of a pediatric diagnostic tool with widespread global impact.
Keywords: electronic health record, electronic laboratory diagnostic tools, patient-centered medicine
Introduction
Clinical laboratories are responsible for accurately performing diagnostic tests, generating test result reports, making appropriate interpretative recommendations, and communicating them clearly and quickly to clinical decision-makers. Increasing implementation of electronic health records (EHRs) and laboratory information systems (LIS) in clinical institutions, in conjunction with increasing test automation and patient data accessibility, has increased the demand and feasibility of electronic healthcare tool integration to improve clinical service and support clinical decisions. An increasing number of web- and mobile-based applications have been developed in recent years to improve access to laboratory test results, increase accuracy in test result interpretation, and aid in complex medical diagnostic management. These resources can be useful to clinicians, laboratory scientists, and patients advocating in their own health care decisions. While this lends itself to a new patient-centered model of healthcare delivery, it also poses further challenge and complexity in both laboratory-clinician and patient-clinician communication, as well as patient privacy and equity. The current review explores available electronic tools in healthcare and laboratory medicine, comments on their respective benefits and limitations, and discusses the future role of electronic healthcare tools in laboratory medicine.
Complex data driven resources: A clinician's aid
Appropriate medical diagnosis of several pathologic conditions requires integration of clinical information (e.g., family history, symptoms, risk factors, imaging) and laboratory data (e.g., biomarker test results) in combination with medical expertise. Retrieving clinical data scattered throughout medical records, synthesizing for risk classification, and applying guideline recommendations to determine appropriate follow-up/treatment can be unnecessarily time consuming. Recent reports also suggest current EHR structures may lead to information overload, as well as missed alerts and information in patient charts, heightening risk of inappropriate follow-up.[1] There is thus increasing interest in data-driven diagnostic resources wherein guideline recommendations for the management of common clinical conditions can be applied automatically to EHR laboratory data, supporting timely and consistent clinical decision-making.
Through emerging technology, EHRs now allow the integration of add-on apps, whereby third-party app developers can enhance the utility of EHRs for targeted healthcare tasks. Substitutable Medical Applications and Reusable Technologies on Fast Healthcare Interoperability Resources (SMART on FHIR) is the open-access technology enabling EHR add-on apps, focusing on utility, usability, and privacy.[2] However, few groups have demonstrated measurable positive outcomes of SMART on FHIR app implementation in clinical settings. Kawamoto et al. recently reported implementing an EHR add-on app for neonatal bilirubin management, which saved clinicians significant time for bilirubin management tasks and significantly increased the odds of clinically appropriate phototherapy orders during hospitalization by 84%.[3] Twitchell et al. also reported the introduction of a SMART on FHIR app for pediatric blood pressure visualization in a tertiary hospital was associated with increased recognition of abnormal blood pressure.[4] Additional proposed applications of SMART on FHIR app technology for clinical decision support include safe opioid prescription[5], cholesterol and cardiac risk management[6], as well as precision cancer medicine[7] and genome and phenome variant identification[8], among others. Patient-clinician tools for the management of patients with severe hemophilia A and/or B have also been established to assist clinicians in pharmacokinetic modelling to determine appropriate prophylactic factor replacement based on laboratory results. These tools also enable patients to document relevant outcomes, improving disease monitoring.[9] Development and implementation of such technology can improve laboratory data interpretation by enabling the automatic consideration of other covariates (e.g., multiple laboratory test measures, clinical information) to complete the clinical picture, thus harnessing the inherent structure of EHRs to propel healthcare into the future of personalized laboratory medicine.
In addition to integrating add-on EHR apps based on existing clinical guidance or published studies, data contained in EHRs can also be utilized for research purposes to develop AI-powered algorithms that could potentially identify novel pathophysiological clusters of diagnostic importance.[10] Indeed, deep learning methods have already been used in healthcare to develop electronic diagnostic tools.[10] For example, Gulshan et al. developed an algorithm for automated detection of diabetic retinopathy and diabetic macular edema using EHR retinal fundus photographs with high sensitivity and anticipated clinical value.[11] In addition, Yuan et al. harnessed EHR data to assemble a large lung cancer cohort and develop a prognostic survival model based on multiple clinical covariates.[12] Rahmini et al. also developed an ontological approach to accurately identify type 2 diabetes mellitus patients in EHRs to support decision-making, as well as conduct audit and evaluation research in diabetes cases.[13] Further, Tomašev et al. developed a model harnessing a large longitudinal dataset of EHRs to assist in predicting future acute kidney injury and provide confidence assessment for clinical outcome predications, including trajectories of clinically relevant blood tests.[14]
These studies provide proof-of-concept examples of how EHR laboratory and clinical data can be harnessed to develop electronic diagnostic and prognostic tools that may reduce healthcare burden and lead to more personalized and timelier patient management, as well as identify potentially novel disease subsets and risk factors for medical conditions. As the provider of the majority of objective data in patient charts, clinical laboratories should consider how integrating clinical information through visual and algorithmic outputs could improve clinical service and lead to healthcare benefits. However, sources of bias in EHR data and their potential to contribute to healthcare disparities should be considered. For example, EHR data may be biased toward certain patient populations (e.g., more severe) and make inappropriate inferences regarding diagnosis and prognosis.[15] Predictive analytical algorithms in certain populations may also not be able to be applied at all institutions due to population differences such as ethnicity.[15] Regardless, research suggests integrating complex data-driven resources into the standard of care for certain conditions (e.g., diabetes or jaundice management) could be useful in the future of healthcare.
Simple interpretative resources: A patient's aid
In addition to the complex medical diagnostic electronic tools geared to clinician’s use, there are also more simplistic tools available, designed specifically for patients.[16][17] Indeed, widespread use of health information technology has increased patients' access to their health information, facilitating goals of advancing patient-centered care. Electronic tools such as patient portals, artificial intelligence-powered symptom checkers, as well as diet, weight, and fertility monitoring applications have engaged patients in their own medical care.
The number of patient-facing health apps has increased substantially in the last five years, and it is estimated that healthcare apps were downloaded more than 3.8 billion times in 2017, increasing 16% from the previous year.[18] The unexpected transition to telemedicine throughout the COVID-19 pandemic has also accelerated the utilization of electronic healthcare apps, offering educational resources for individuals with varying levels of health and digital literacy.
Despite the positive outcomes of increased medical data accessibility to patients, there is concern regarding misinformation and safety.[17] In a recent scoping review, Akbar et al. found that electronic healthcare app development processes significantly lacked the involvement of relevant healthcare professionals and often presented misinformation, including erroneous calculation algorithms and incorrect information on expected disease symptoms and treatment options.[18] These concerns are not limited to third-party apps. Indeed, a recent survey assessing the frequency and types of errors identified by patients who read physician notes in their EHRs found one in five patients reported finding an error (e.g., mistakes in diagnoses, medical history, medications, physical examination, test results) and 40% perceived the error as serious.[19]
Specific to laboratory medicine, patients often assume their test results should be the same regardless of which laboratory or institution completed the bloodwork. It is not understood by patients, and sometimes clinicians, that laboratory test results for many markers of health and disease will vary by testing platform/assay due to lack of assay standardization.[20] As patients have increasing access to their medical data, it is critical that the field of laboratory medicine continue efforts in analytical standardization and reference interval harmonization for candidate assays to ensure consistent test result interpretation, especially given the increasingly multi-hospital network nature of patient care complicating longitudinal follow-up. In addition, without a clinician contextualizing the result, it is important that there are processes and educational resources in place to reduce misinformation and ensure accessible options are available when questions or concerns arise. Indeed, a recent study conducted by Zhang et al. reported 35.2% of individuals have difficulty understanding their laboratory medical test results and 67% are unable to comprehend different aspects of their test results, including reference range, medical terminology, and the meaning of the laboratory value.[21] It is therefore important that electronic tools informed by scientific evidence and expertise are created to bridge this gap.
CALIPER electronic tool for pediatric test result interpretation: Our experience
The Canadian Laboratory Initiative on Pediatric Reference Intervals (CALIPER) is a nationwide initiative aimed to improve blood test result interpretation in children by establishing accurate and robust pediatric reference intervals for important biomarkers of health and disease.[22] For many years, blood test interpretation in children was inappropriately based on adult reference values, significantly increasing the risk of misinformed clinical decisions and treatment/follow-up. Children are not small adults, and the physiological changes occurring throughout growth and development often necessitate pediatric-specific interpretation for many laboratory tests.[23] Since 2008, CALIPER has emerged as a global leader in establishing and publishing comprehensive pediatric reference intervals for over 200 laboratory biomarkers, including biochemical, immunological, hematological, nutritional, endocrine, and fertility markers, as well as many special chemistry evaluations on several analytical platforms based on data from thousands of healthy children and adolescents from birth to 18 years.[22]
In addition to the generation and publication of these rich datasets, it was equally important to the CALIPER team to develop knowledge translation tools to ensure data is accessible to clinicians, laboratory professionals, patients, and families worldwide. Thus, a mobile application (available for download on Google Play and Apple Store) and online database (www.caliperdatabase.org) was developed in 2014 and recently updated. These electronic tools enable patient laboratory results to be interpreted against the most appropriate and up-to-date reference intervals developed by CALIPER on multiple clinical chemistry platforms and assays. Both electronic tools are freely accessible worldwide and updated with new tests as further peer-reviewed studies are completed.
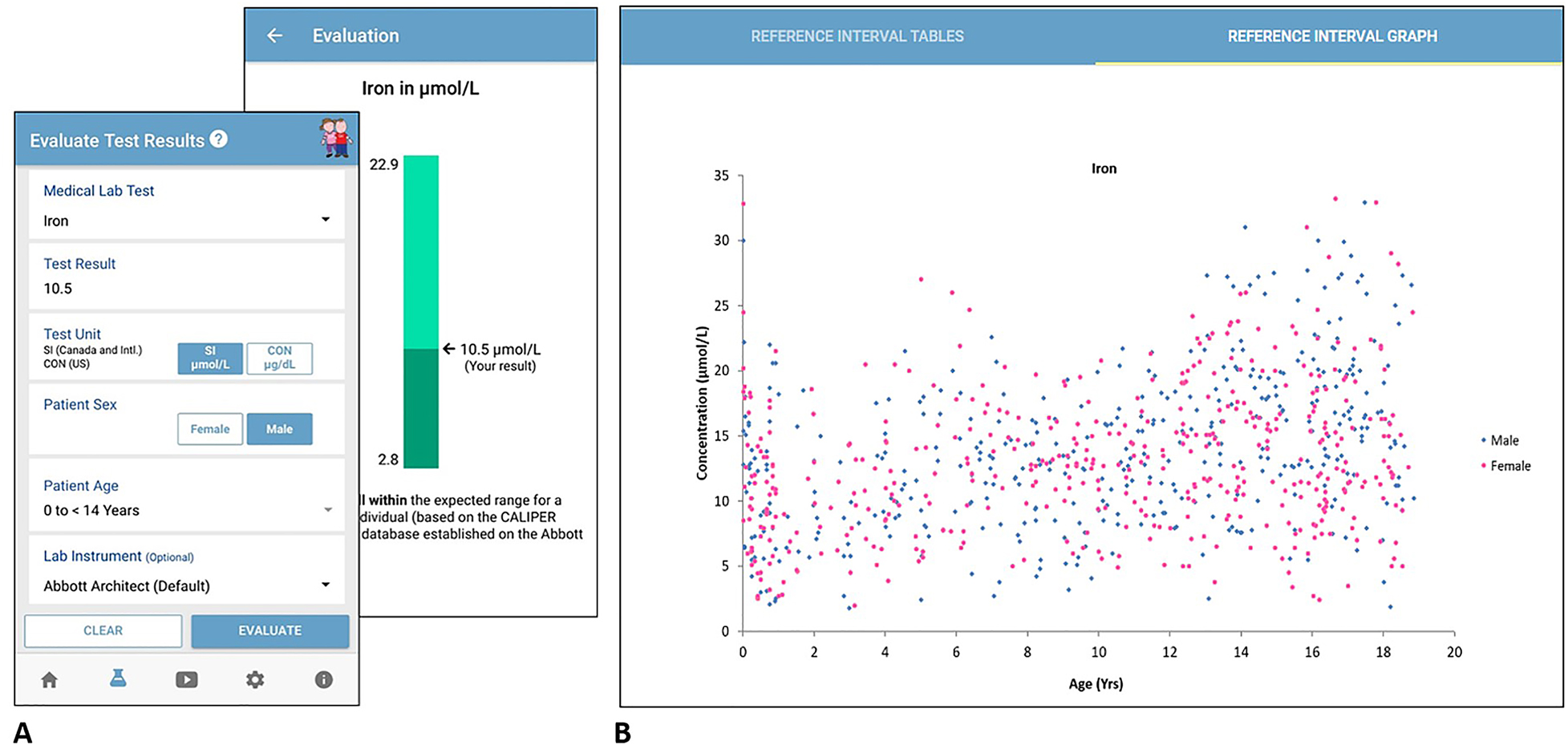
To use the mobile app or online database, one can simply select the medical lab test of interest, enter the child’s test result, and select the child’s sex and age from the drop-down menu. Users have the option to select the lab instrument they want to assess, which include systems manufactured by various IVD companies. To increase usability worldwide, users can also input the unit of measure, including International System of Units (SI) or Conventional (CON) units. After inputting the necessary information, the mobile app assesses the patient result value as compared to CALIPER reference intervals (Figure 1). The online database provides even more in-depth information, including a summary of all pediatric reference intervals for that analyte, as well as age- and sex-specific scatterplots to better visualize and depict dynamic reference value concentrations observed (Figure 1).
|
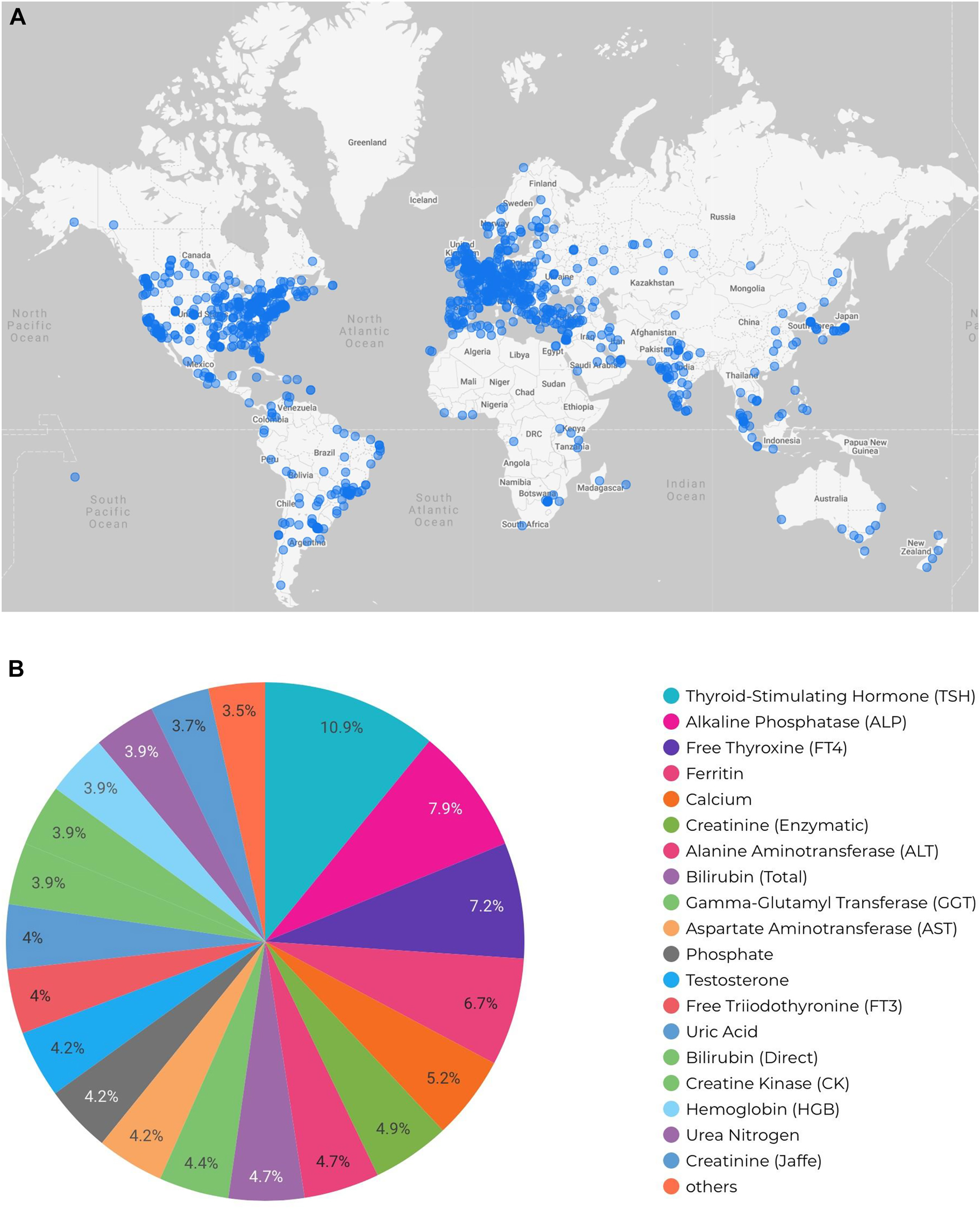
The utility and value of electronic healthcare tools, such as the CALIPER database, is underscored by their global reach with a high number of routine app users. To date, the CALIPER online database has successfully established a strong global presence with over 5,000 registered users from 3,650 institutions spanning over 100 countries in all regions around the world (Figure 2). The demonstrated global use of these electronic tools speaks not only to the value of the data contained within the database but also the value of well-designed electronic tools in laboratory medicine. Analytical outputs from such electronic tools can also inform further research and consumer interest. For example, Figure 2B demonstrates the most commonly searched biomarkers on the CALIPER database. In this report, it is evident that thyroid markers such as thyroid stimulating hormone (TSH) and free thyroxine (FT4) are of significant interest to database users. This is not surprising as thyroid disorders—including hyperthyroidism, hypothyroidism, Grave's disease, and Hashimoto's—are common in children, particularly adolescent females.[24] Other markers of significant interest include liver markers (e.g., alkaline phosphatase, alanine aminotransferase, bilirubin, gamma-glutamyl transferase) and nutritional markers (e.g., ferritin, calcium).
|
With the tremendous global interest CALIPER has generated to help fill the critical gap in establishing pediatric reference intervals, there has been an urgency to create a global resource for all adult and pediatric reference interval studies across the world in an effort to ensure all countries have access to appropriate reference intervals for test interpretation. An International Federation for Clinical Chemistry and Laboratory Medicine (IFCC) Task Force on Global Reference Interval Database (TF-GRID) is currently under development[25], with the aim of creating a centralized global reference interval database by collating data from high-quality pediatric and adult reference interval initiatives. This database will enable users to search for major reference interval studies and view reference interval data. It will also include an interactive component in which the user could input a patient’s test result for comparison to available reference interval data within the database. This centralized resource will improve the accessibility of reference intervals globally, in addition to contributing and promoting global harmonization for assays that demonstrate minimal analytical and population differences across studies.
Summary and outlook
As technology becomes increasingly integrated into healthcare, it is essential that we consider how to harness existent data present in EHRs to better inform clinical decision-making in the transition to more patient-centered medicine. While complex data-driven applications can assist in patient management as well as identify novel findings of potential clinical significance, simpler tools such as intelligent diagnostic apps and databases can be of equal value to both clinicians and patients, creating a new and innovative model of laboratory-clinician-patient communication and healthcare delivery. The CALIPER mobile app and database serve as a concrete example of how freely accessible resources can be utilized worldwide towards improved clinical service. Clinical laboratories should be leading and actively engaged in electronic diagnostic tool innovations that improve laboratory data communication and reduce post-analytical errors in test result reporting and interpretation, which in turn improve patient safety. Inherent biases to data-driven technologies should also be considered as their use increases in healthcare delivery.
Acknowledgements
Author contributions
All authors have accepted responsibility for the entire content of this manuscript and approved its submission.
Research funding
None declared.
Competing interests
Authors state no conflict of interest.
References
- ↑ Singh, Hardeep; Spitzmueller, Christiane; Petersen, Nancy J.; Sawhney, Mona K.; Sittig, Dean F. (22 April 2013). "Information Overload and Missed Test Results in Electronic Health Record–Based Settings" (in en). JAMA Internal Medicine 173 (8): 702. doi:10.1001/2013.jamainternmed.61. ISSN 2168-6106. PMC PMC3822526. PMID 23460235. http://archinte.jamanetwork.com/article.aspx?doi=10.1001/2013.jamainternmed.61.
- ↑ Mandel, Joshua C; Kreda, David A; Mandl, Kenneth D; Kohane, Isaac S; Ramoni, Rachel B (1 September 2016). "SMART on FHIR: a standards-based, interoperable apps platform for electronic health records" (in en). Journal of the American Medical Informatics Association 23 (5): 899–908. doi:10.1093/jamia/ocv189. ISSN 1527-974X. PMC PMC4997036. PMID 26911829. https://academic.oup.com/jamia/article/23/5/899/2379865.
- ↑ Kawamoto, Kensaku; Kukhareva, Polina; Shakib, Julie H.; Kramer, Heidi; Rodriguez, Salvador; Warner, Phillip B.; Shields, David; Weir, Charlene et al. (15 November 2019). "Association of an Electronic Health Record Add-on App for Neonatal Bilirubin Management With Physician Efficiency and Care Quality" (in en). JAMA Network Open 2 (11): e1915343. doi:10.1001/jamanetworkopen.2019.15343. ISSN 2574-3805. PMC PMC6902796. PMID 31730181. https://jamanetwork.com/journals/jamanetworkopen/fullarticle/2755484.
- ↑ Twichell, Sarah A.; Rea, Corinna J.; Melvin, Patrice; Capraro, Andrew J.; Mandel, Joshua C.; Ferguson, Michael A.; Nigrin, Daniel J.; Mandl, Kenneth D. et al. (1 July 2017). "The effect of an electronic health record-based tool on abnormal pediatric blood pressure recognition: TWICHELL et al." (in en). Congenital Heart Disease 12 (4): 484–490. doi:10.1111/chd.12469. PMC PMC5647583. PMID 28493451. http://doi.wiley.com/10.1111/chd.12469.
- ↑ Sinha, Shyamashree; Jensen, Mark; Mullin, Sarah; Elkin, Peter L (8 September 2017). "Safe Opioid Prescripting: A SMART on FHIR Approach to Clinical Decision Support". Online Journal of Public Health Informatics 9 (2). doi:10.5210/ojphi.v9i2.8034. ISSN 1947-2579. PMC PMC5630280. PMID 29026458. https://journals.uic.edu/ojs/index.php/ojphi/article/view/8034.
- ↑ Bloomfield, Richard A.; Polo-Wood, Felipe; Mandel, Joshua C.; Mandl, Kenneth D. (1 March 2017). "Opening the Duke electronic health record to apps: Implementing SMART on FHIR" (in en). International Journal of Medical Informatics 99: 1–10. doi:10.1016/j.ijmedinf.2016.12.005. https://linkinghub.elsevier.com/retrieve/pii/S1386505616302738.
- ↑ Warner, Jeremy L; Rioth, Matthew J; Mandl, Kenneth D; Mandel, Joshua C; Kreda, David A; Kohane, Isaac S; Carbone, Daniel; Oreto, Ross et al. (1 July 2016). "SMART precision cancer medicine: a FHIR-based app to provide genomic information at the point of care" (in en). Journal of the American Medical Informatics Association 23 (4): 701–710. doi:10.1093/jamia/ocw015. ISSN 1527-974X. PMC PMC6080684. PMID 27018265. https://academic.oup.com/jamia/article/23/4/701/2200286.
- ↑ Alterovitz, Gil; Warner, Jeremy; Zhang, Peijin; Chen, Yishen; Ullman-Cullere, Mollie; Kreda, David; Kohane, Isaac S. (1 November 2015). "SMART on FHIR Genomics: facilitating standardized clinico-genomic apps" (in en). Journal of the American Medical Informatics Association 22 (6): 1173–1178. doi:10.1093/jamia/ocv045. ISSN 1527-974X. https://academic.oup.com/jamia/article/22/6/1173/2357593.
- ↑ McEneny‐King, Alanna; Yeung, Cindy HT; Edginton, Andrea N.; Iorio, Alfonso; Croteau, Stacy E. (19 November 2019). "Clinical application of Web Accessible Population Pharmacokinetic Service—Hemophilia (WAPPS‐Hemo): Patterns of blood sampling and patient characteristics among clinician users". Haemophilia 26 (1): 56–63. doi:10.1111/hae.13882. ISSN 1351-8216. https://doi.org/10.1111/hae.13882.
- ↑ 10.0 10.1 Mandl, Kenneth D.; Bourgeois, Florence T. (21 November 2017). "The Evolution of Patient Diagnosis: From Art to Digital Data-Driven Science" (in en). JAMA 318 (19): 1859. doi:10.1001/jama.2017.15028. ISSN 0098-7484. http://jama.jamanetwork.com/article.aspx?doi=10.1001/jama.2017.15028.
- ↑ Gulshan, Varun; Peng, Lily; Coram, Marc; Stumpe, Martin C.; Wu, Derek; Narayanaswamy, Arunachalam; Venugopalan, Subhashini; Widner, Kasumi et al. (13 December 2016). "Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs" (in en). JAMA 316 (22): 2402. doi:10.1001/jama.2016.17216. ISSN 0098-7484. http://jama.jamanetwork.com/article.aspx?doi=10.1001/jama.2016.17216.
- ↑ Yuan, Qianyu; Cai, Tianrun; Hong, Chuan; Du, Mulong; Johnson, Bruce E.; Lanuti, Michael; Cai, Tianxi; Christiani, David C. (7 July 2021). "Performance of a Machine Learning Algorithm Using Electronic Health Record Data to Identify and Estimate Survival in a Longitudinal Cohort of Patients With Lung Cancer" (in en). JAMA Network Open 4 (7): e2114723. doi:10.1001/jamanetworkopen.2021.14723. ISSN 2574-3805. PMC PMC8264641. PMID 34232304. https://jamanetwork.com/journals/jamanetworkopen/fullarticle/2781685.
- ↑ Rahimi, Alireza; Liaw, Siaw-Teng; Taggart, Jane; Ray, Pradeep; Yu, Hairong (1 October 2014). "Validating an ontology-based algorithm to identify patients with Type 2 Diabetes Mellitus in Electronic Health Records" (in en). International Journal of Medical Informatics 83 (10): 768–778. doi:10.1016/j.ijmedinf.2014.06.002. https://linkinghub.elsevier.com/retrieve/pii/S1386505614001038.
- ↑ Tomašev, Nenad; Glorot, Xavier; Rae, Jack W.; Zielinski, Michal; Askham, Harry; Saraiva, Andre; Mottram, Anne; Meyer, Clemens et al. (1 August 2019). "A clinically applicable approach to continuous prediction of future acute kidney injury" (in en). Nature 572 (7767): 116–119. doi:10.1038/s41586-019-1390-1. ISSN 0028-0836. PMC PMC6722431. PMID 31367026. http://www.nature.com/articles/s41586-019-1390-1.
- ↑ 15.0 15.1 Gianfrancesco, Milena A.; Tamang, Suzanne; Yazdany, Jinoos; Schmajuk, Gabriela (1 November 2018). "Potential Biases in Machine Learning Algorithms Using Electronic Health Record Data". JAMA Internal Medicine 178 (11): 1544. doi:10.1001/jamainternmed.2018.3763. ISSN 2168-6106. PMC PMC6347576. PMID 30128552. https://doi.org/10.1001/jamainternmed.2018.3763.
- ↑ Baldwin, Jessica L.; Singh, Hardeep; Sittig, Dean F.; Giardina, Traber Davis (1 September 2017). "Patient portals and health apps: Pitfalls, promises, and what one might learn from the other" (in en). Healthcare 5 (3): 81–85. doi:10.1016/j.hjdsi.2016.08.004. PMC PMC8386919. PMID 27720139. https://linkinghub.elsevier.com/retrieve/pii/S2213076416300124.
- ↑ 17.0 17.1 Jovičić, Snežana; Siodmiak, Joanna; Watson, Ian D. (25 February 2019). "Quality evaluation of smartphone applications for laboratory medicine" (in en). Clinical Chemistry and Laboratory Medicine (CCLM) 57 (3): 388–397. doi:10.1515/cclm-2018-0710. ISSN 1437-4331. https://www.degruyter.com/document/doi/10.1515/cclm-2018-0710/html.
- ↑ 18.0 18.1 Akbar, Saba; Coiera,, Enrico; Magrabi, Farah (1 February 2020). "Safety concerns with consumer-facing mobile health applications and their consequences: a scoping review" (in en). Journal of the American Medical Informatics Association 27 (2): 330–340. doi:10.1093/jamia/ocz175. ISSN 1527-974X. PMC PMC7025360. PMID 31599936. https://academic.oup.com/jamia/article/27/2/330/5585394.
- ↑ Bell, Sigall K.; Delbanco, Tom; Elmore, Joann G.; Fitzgerald, Patricia S.; Fossa, Alan; Harcourt, Kendall; Leveille, Suzanne G.; Payne, Thomas H. et al. (9 June 2020). "Frequency and Types of Patient-Reported Errors in Electronic Health Record Ambulatory Care Notes" (in en). JAMA Network Open 3 (6): e205867. doi:10.1001/jamanetworkopen.2020.5867. ISSN 2574-3805. PMC PMC7284300. PMID 32515797. https://jamanetwork.com/journals/jamanetworkopen/fullarticle/2766834.
- ↑ Plebani, Mario (1 April 2013). "Harmonization in laboratory medicine: the complete picture" (in en). Clinical Chemistry and Laboratory Medicine (CCLM) 51 (4): 741–751. doi:10.1515/cclm-2013-0075. ISSN 1437-4331. https://www.degruyter.com/document/doi/10.1515/cclm-2013-0075/html.
- ↑ Zhang, Zhan; Citardi, Daniel; Xing, Aiwen; Luo, Xiao; Lu, Yu; He, Zhe (7 December 2020). "Patient Challenges and Needs in Comprehending Laboratory Test Results: Mixed Methods Study" (in en). Journal of Medical Internet Research 22 (12): e18725. doi:10.2196/18725. ISSN 1438-8871. PMC PMC7752528. PMID 33284117. http://www.jmir.org/2020/12/e18725/.
- ↑ 22.0 22.1 Adeli, Khosrow; Higgins, Victoria; Trajcevski, Karin; White-Al Habeeb, Nicole (18 August 2017). "The Canadian laboratory initiative on pediatric reference intervals: A CALIPER white paper" (in en). Critical Reviews in Clinical Laboratory Sciences 54 (6): 358–413. doi:10.1080/10408363.2017.1379945. ISSN 1040-8363. https://www.tandfonline.com/doi/full/10.1080/10408363.2017.1379945.
- ↑ Adeli, Khosrow; Higgins, Victoria; Trajcevski, Karin; Palmert, Mark R (24 May 2018). "Important considerations for interpreting biochemical tests in children" (in en). BMJ: k1950. doi:10.1136/bmj.k1950. ISSN 0959-8138. https://www.bmj.com/lookup/doi/10.1136/bmj.k1950.
- ↑ Hanley, Patrick; Lord, Katherine; Bauer, Andrew J. (1 October 2016). "Thyroid Disorders in Children and Adolescents: A Review" (in en). JAMA Pediatrics 170 (10): 1008. doi:10.1001/jamapediatrics.2016.0486. ISSN 2168-6203. http://archpedi.jamanetwork.com/article.aspx?doi=10.1001/jamapediatrics.2016.0486.
- ↑ "Task Force on Global Reference Interval Database (TF-GRID)". International Federation for Clinical Chemistry and Laboratory Medicine. 2021. https://www.ifcc.org/executive-board-and-council/eb-task-forces/task-force-on-global-reference-interval-database-tf-grid/.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation, grammar, and spelling. In some cases important information was missing from the references, and that information was added. A reference for TF-GRID was added for this version.