Journal:Design of generalized search interfaces for health informatics
| Full article title | Design of generalized search interfaces for health informatics |
|---|---|
| Journal | Information |
| Author(s) | Demelo, Jonathan; Sedig, Kamran |
| Author affiliation(s) | Western University |
| Primary contact | Email: sedig at uwo dot ca |
| Editors | Almada, Marta |
| Year published | 2021 |
| Volume and issue | 12(8) |
| Article # | 317 |
| DOI | 10.3390/info12080317 |
| ISSN | 2078-2489 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://www.mdpi.com/2078-2489/12/8/317/htm |
| Download | https://www.mdpi.com/2078-2489/12/8/317/pdf (PDF) |
|
|
This article should be considered a work in progress and incomplete. Consider this article incomplete until this notice is removed. |
Abstract
In this paper, we investigate ontology-supported interfaces for health informatics search tasks involving large document sets. We begin by providing background on health informatics, machine learning, and ontologies. We review leading research on health informatics search tasks to help formulate high-level design criteria. We then use these criteria to examine traditional design strategies for search interfaces. To demonstrate the utility of the criteria, we apply them to the design of the ONTology-supported Search Interface (ONTSI), a demonstrative, prototype system. ONTSI allows users to plug-and-play document sets and expert-defined domain ontologies through a generalized search interface. ONTSI’s goal is to help align users’ common vocabulary with the domain-specific vocabulary of the plug-and-play document set. We describe the functioning and utility of ONTSI in health informatics search tasks through a workflow and a scenario. We conclude with a summary of ongoing evaluations, limitations, and future research.
Keywords: information search, search tasks, health informatics, interface design, ontologies, machine learning, PubMed
Introduction
Health informatics is concerned with emergent technological systems that improve the quality and availability of care, promote the sharing of knowledge, and support the performance of proactive health and wellness tasks by motivated individuals.[1] Subareas of health informatics may include medical informatics, nursing informatics, consumer informatics, cancer informatics, and pharmacy informatics, to name a few. Simply put, health informatics is concerned with harnessing technology for finding new ways to help stakeholders work with health information to be able to perform health-related tasks more effectively.
Users in the health domain are increasingly taking advantage of computer-based resources in their tasks. For instance, a 2017 Canadian survey found that 32% of respondents within their last month had used at least one mobile application for health-related tasks. Even more, those under the age of 35 are twice as likely to do so.[2] Furthermore, studies have calculated that over 58% of Americans have used tools like Google and other domain-specific tools to support their health informatics search tasks, with search being one of the most important and central tasks in most health informatics activities.[3][4]
Yet, search can be challenging, particularly for health informatics tasks that utilize large and complex document sets. For such tasks, health informatics tools may require the use of domain-specific vocabulary. Aligning with this vocabulary can be a significant challenge within health tasks, as they can involve a lexicon of intricate nomenclature, deeply layered relations, and lengthy descriptions that are misaligned with common vocabulary. For instance, one highly cited medical research paper defines the term “chromosomal instability” as “an elevated rate of chromosome mis-segregation and breakage, results in diverse chromosomal aberrations in tumor cell populations.” In this example, those unfamiliar with the defined term could find parsing its definition just as significant a challenge as the term itself.[5] Thus, when communicating across vocabularies, users may struggle to describe the requirements of their search task in a way that is understandable by health informatics tools.[6][7] To deal with this challenge, ontologies can be a valuable mediating resource in the design of user-facing interfaces of health informatics tools.[8] That is, ontologies can bridge the vocabularies of users with the vocabulary of their task and its tools. Yet, the use of ontologies in user-facing interface design is not well established. Furthermore, health informatics tools that present a generalized interface, one that can support search tasks across any number of domain vocabularies and document sets, can allow users to transfer their experience between tasks, presenting users with information-centric perspectives during their performances rather than technology-centered perspectives.[9][10] For this, there is a need to distill criteria that can guide designers during the creation of ontology-supported interfaces for health informatics search tasks involving large document sets.
The goal of this paper is to investigate the following research questions:
- What are the criteria for the structure and design of generalized ontology-supported interfaces for health informatics search tasks involving large document sets?
- If such criteria can be distilled, can they then be used to help create such interfaces?
In this paper, we examine health informatics, machine learning, and ontologies. We then review leading research on health informatics search tasks. From this analysis, we formulate criteria for the design of ontology-supported interfaces for health informatics search tasks involving large document sets. We then use these criteria to contrast the traditional design strategies for search interfaces. To demonstrate the utility of the criteria in design, we will use them to structure the design of a tool, ONTSI (ONTology-supported Search Interface). ONTSI allows users to plug-and-play their document sets and expert-defined ontology files to perform health informatics search tasks. We describe ONTSI through a functional workflow and an illustrative usage scenario. We conclude with a summary of ongoing evaluation efforts, future research, and our limitations.[11]
Background
In this section, we describe the concepts and terminology used when discussing ontology-supported interfaces for health informatics search tasks involving large document sets. We begin with background on health informatics. Next, we examine machine learning. We conclude with coverage of ontologies and their utility as a mediating resource for both human- and computer-facing use.
Health informatics
Health informatics is broadly concerned with emergent technological systems for improving the quality and availability of care, promoting the sharing of knowledge, and supporting the performance of proactive health and wellness practices by motivated individuals.[1] Initially, the need for expanded health and wellness services stemmed from rising population levels combined with the growing complexity of medical sciences. These issues made it challenging to maintain quality care within increasingly stressed medical systems.[12] Thus, a central objective for health informatics is the development of strategies to tackle large-scale problems that harm trained medical professionals’ ability to perform their tasks in a timely and effective manner. For instance, telehealth services allowed doctors to practice remote medicine, providing care to those without local medical services. Another early innovation was standardized electronic health care records (EHRs), where patient records were given standardized encodings to provide an increased ability to track, compare, manage, and share personal health information.[3] Some examples of current research directions are the push for stronger patient privacy, personalized medicine, and the expansion of healthcare into underserved regions and communities.[1][2][3][13][14]
The rising production and availability of health-related data has resulted in a growing number of data-intensive tasks within health. Both private and public entities like health industry companies, government bodies, and everyday citizens are turning to health informatics tools as they manage and activate their health data.[2] A growing number of health-related tasks involve searching document sets. During these tasks, the aim of the user is to use the information described within their document set to increase their understanding of a topic or concept. For example, a search task could be a practitioner searching the EHRs of their patients, a member of the general public using public materials for their general health concerns, or a researcher performing a literature review.[12][15][16] In general, a search task involves the generation of a query based on an information-seeking objective. The computation systems of these tools then use this query to map and extract relevant documents out from the document set.[15] Powerful technologies like machine learning are increasingly being integrated within tools to help perform rapid and automated computation on document sets.[4] Yet, when taking advantage of these technologies, designers must be mindful of human factors when generating the user-facing interfaces of their tools, as a task cannot be performed effectively without direction from an empowered user.[12]
Machine learning
Machine learning techniques are increasingly being utilized to tackle analytic problems once considered too complex to solve in an effective and timely manner.[16] Yet, recent analysis[17][18][19] on the human factors in machine learning environments have found that the current design strategies continually limit users’ ability to take part in the analytic process. More so, it has produced a generation of machine learning-integrated tools that are failing to provide users a complete understanding on how computational systems of their tools arrive at their results. This has significantly reduced users’ control and lowered the ability to achieve task objectives. In response, there is a growing desire to promote the “human-in-the-loop,” bringing the benefits of human reasoning back to the forefront of the design process.[20][21][22]
When considering the interaction loop of a machine learning-integrated tool, Sacha et al.[23] present a five-stage conceptual framework: producing and accessing data, preparing data for tool use, selecting a machine learning model, visualizing computation in the tool interface, and users applying analytic reasoning to validate and direct further use. Assessing this framework, a machine learning-integrated search tool must provide users with a functional workflow where:
- users communicate their task requirements as a query;
- users ask their tool to apply that query as input within its computational system;
- the tool performs its computation, mapping the features against the document set;
- the tool represents the results of the computation in its interface;
- users assess whether they are or are not satisfied with the results; and
- users restart the interaction loop with adjustments or conclude their use of the tool.
Thus, a primary responsibility for users within machine learning environments is the need to assess how well the results of machine learning have aligned with their task objectives. A systematic review by Amershi et al.[24] suggests six considerations for the user’s role in arbitrating machine learning performance:
- Users are people, not oracles (i.e., they should not be expected to repeatedly answer whether a model is right or wrong).
- People tend to give more positive than negative feedback.
- People need a demonstration of how machine learning should behave.
- People naturally want to provide more than just data labels.
- People value transparency.
- Transparency can help people provide better labels.
Ontologies
In search tasks involving large document sets, many challenges can arise that reduce performance quality, harm user satisfaction, and increase the time for task completion.[17][18][19] Often, these challenges result from misalignment between the vocabularies used by the document sets, storage maintainers, interface designers, and users. For instance, Qing et al.[25] outline the difficulties faced when translating between common and domain vocabularies in health tasks. They describe a study that found that up to 50% of health expressions by consumers were not represented by public health vocabularies.[25]
Within the pipeline of a search task, both the human and computational system can only perform optimally if communication is strong.[26] Ontologies are representational artifacts that reflect the entities, relations, and structures of its domain. Ontologies are of three types: a philosophical ontology for describing and structuring reality, a domain ontology for structuring the entities and relations of a knowledge base, and a top-level ontology for interfacing between different domain ontologies.[26] Ontologies provide the flexibility, extensibility, generality, and expressiveness necessary to bridge the gap when mapping domain knowledge for effective computer-facing and human-facing use.[8] For this purpose, ontologies are increasingly being used within tools to help users perform their challenging search tasks.[27][28][29][30][31]
When creating an ontology, experts construct a network of entities and relations, which together yield various structures.[32][33] Ontology entities reflect the objects of the domain, like a phenotype in a medical abnormality ontology, a processor in a computer architecture ontology, or a precedent in a legal ontology.[34] In some ontologies, like the top-level ontology, Basic Formal Ontology, designers go as far as denoting qualities such as materiality, object composition, and spatial qualities in reality.[26] Ontology entities are encoded with information about their role in the vocabulary, definitions, descriptions, and contexts, as well as metadata that can inform the performance of future ontology engineering tasks.
Ontology relations are the links between ontology entities that express the quality of interaction between them and the domain as a whole.[35] When assessing ontology relations, Arp et al.[26] distinguish relations under the categories of universal–universal (dog “is_a” animal), particular–universal (this dog “instance_of” dog), and particular–particular (this dog “continuant_parts” of this dog grouping). Domain ontology relations are realized through unique interoperability between ontology entities. For instance, an animal ontology may have an ontology entity reflecting the concept of a “human,” which may have the ontology relations “domesticates/is domesticated by” between it and the “dog” ontology entity.
After defining the entities, relations, and other features of an ontology, experts record their work in ontology files of standardized data formats like RDF, OWL, and OBO. These ontology files are then distributed amongst users. They can then be integrated into the computational and human-facing systems of tools for use during tasks. Some examples of current ontology use are information extraction on unstructured text, behavior modeling of intellectual agents, and an increasing number of human-facing visualization tasks such as decision support systems within critical care environments.[20][21][22]
Methods
In this section, we describe the methods used for criteria formulation. We begin with a review of literature for health informatics search tasks. Based on the insights gained from this review, we distill a set of criteria. We then use these criteria to contrast traditional design strategies for interfaces of search tasks.
Task review
Here, we review some research on interfaces for health informatics search tasks. We used Google Scholar, IEEE Xplore, and PubMed to conduct an exhaustive search of articles and reviews published between 2015 and 2021. We have divided our findings into three sections. First, we explore research on health data, information management, and information-centric interfaces. This is followed by research discussing the types of search tasks and their use in structuring the design of interfaces for health informatics. Finally, we investigate the requirements for aligning vocabularies for health informatics search tasks.
Health data, information management, and information-centric interfaces
Health data is constantly generated, highlighted by reports that within just a year the U.S. healthcare system created 150 new exabytes of data.[36] Yet, the information that is expressed by this data, such as personal medical records, research publications, and consumer health media, is not useful unless it can be effectively understood and utilized by users. As such, it is critical to examine the challenges facing users when performing their tasks, and through this establish novel strategies for supporting the activation of health data.
Fang et al.[10] explore the pressing challenges for accessing health data under the four categories: volume, variety, velocity, and veracity. They find that the volume of health care data creates challenges in the management of data sources and stores. They describe that existing strategies are struggling, and that novel designs should be established for scaling data services. They explain that a variety of challenges come with the management of data characteristics, ranging from unstructured datapoints generated from sources like sensors, to structured data entities like research papers and medical documents. For this, they state that designers should concentrate on aligning with the characteristics of the information being encountered. Next, they explore the challenges of velocity, which involves the rate at which users require their data to move from source to activation within their task. They highlight novel research in the networking and data management space. Finally, they explore veracity challenges, such as the assessment and validation of data quality and the quality of information that the data may produce.
Gibson et al.[9] provide a review of the evolving fields of health information management and informatics. They review the topics of data capture, digital e-record systems, aggregate health management, healthcare funding models, data-oriented evidence-based medicine, consumer health applications, health governance, personal health access, and genomic personalized medicine. Similar to Fang et al.[10], they note that the predominant work for health informatics should be concerned with presenting users with information-centric perspectives during their performances rather than technology-centered perspectives. More so, they describe that users in healthcare “must often navigate and understand complex clinical workflows to effectively … capture, store, or exchange information.” In other words, task workflows are already complex; therefore, effective interfaces should promote information encounters that help users perform better, rather than engage in unrelated technical details.
From the above research, we distill the criterion: Designs should maintain an information-centric interface that is flexible with respect to the dynamic requirements of search tasks like veracity of data sources, variety of data types, and evolving needs of users for health informatics.
Search tasks and structuring the design of interfaces for health informatics
Russell-Rose et al.[37] describe professional health workplace tasks. They find that the most prevalent types of search tasks are literature reviews for overviewing a topic, scoping reviews for rapidly inspecting the possible relevance of an information source, rapid evidence reviews for appraising the overall quality of a scoping review, and, finally, systematic reviews for exploring a topic in a robust manner.
During search tasks, users often lack the ability to perceive how their query decisions impact, relate, and interact with the document set. This is an important consideration for users who might want to adjust a query to better align with their information-seeking objectives. A further study by Russell-Rose et al.[38] analyzes search strategies performed by healthcare professionals. They find that a large majority of participants have a general desire to utilize advanced search functionalities when available. This suggests that users are not hesitant to take advantage of resources that they believe help optimize their task performance. Huurdeman [39] outlines that for this, a good course of action is to leverage query corrections, autocomplete, and suggestions. Yet, they find that such additions can be harmful if those features do not provide appropriate domain context. That is, resources must allow users to be contextually aware of how their query aligns with the contents of document sets, as well as the conditions of computational technologies used by interfaces.
In the same research, Huurdeman[39] investigates complex tasks involving information search and information-seeking models when using multistage search systems. In this research, they explore requirements that designers must account for when supporting users. Challenging search tasks require users to learn about the searched domain, understand how their objectives align, and formulate their objectives into a way that can be used by their tool. In other words, query building requires users to be domain cognizant, as they must communicate information-seeking objectives in a way that is understood by the tool, yet also aligns with the information found within the document sets. Thus, a health informatics tool that supports search tasks should provide the opportunity for understanding the domain of the document set being explored.
Zahabi et al.[40] describe a set of nine requirements for designers when considering how to design usable interfaces for health informatics search tasks, summarized as:
- Naturalness: The workflow of the system must present a natural task progression.
- Consistency: The parts of the system should present similar functional language.
- Prevent errors: Be proactive in the prevention of potential errors.
- Minimizing cognitive load: Align cognitive load to the requirements of the task.
- Efficient interaction: Be efficient in the number of steps to complete a task.
- Forgiveness and feedback: Supply proper and prompt feedback opportunities.
- Effective use of language: Promote clear and understandable communication.
- Effective information presentation: Align with information characteristics.
- Customizability/flexibility: The system should remain flexible to the task requirements.
Additional research by Dudley et al.[41] reviews user interface design for machine learning environments. They provide a set of principles that can be used by designers:
- Make task goals and constraints explicit.
- Support user understanding of model uncertainty and confidence.
- Capture intent rather than input.
- Provide effective data representations.
- Exploit interactivity and promote rich interactions.
- Engage the user.
From this research, we can distill two criteria. First, designs should provide interaction loops that promote prompt and effective feedback opportunities for the user. Second, designs should provide representations that are natural and consistent to the requirements of the information source, the user, and the task.
Aligning vocabularies for health informatics search tasks
When considering interfaces for health informatics search tasks, a major challenge for users is the need to overcome problem formulation deficiencies when encountering unfamiliar domains. This is because, according to Harvey et al.[42], users have been found to consistently suffer from four major issues during the performance of search tasks:
- difficulty understanding the domain being searched;
- an inability to apply their domain expertise;
- lacking the capacity to formulate an effective search query within the interface that accurately reflects their information-seeking objective; and
- deficient understanding of how to assess results produced by search, to decide whether the search has or has not satisfied their objective.
Harvey et al.[42] show that in domains with complex vocabularies, such as health and medicine, the disparity of potential users’ prior knowledge is extreme. They find that non-expert users routinely do not possess enough domain knowledge to address their information-seeking needs. This can cause significant issues during query formulation. As a result, non-expert users must first step away from their tool to learn specialized vocabulary before they can begin query building. Both Soldaini et al.[43] and Anderson and Wischgoll[44] describe that this issue can still affect even experts. This is because experts often must make assumptions when attuning to their tool.
There is growing research targeting the generation and application of mediation resources to help reduce the communication gap while using health informatics tools. Zeng et al.[25] investigate the development of consumer health vocabularies for reducing the discourse gap between lay people and medical information document sets. Furthermore, Soldaini et al.[45] explore the use of novel query computation strategies to improve the quality of medical literature retrieval during search tasks. In their quantitative study, they contrast models generated using combinations of algorithms, vocabularies, and feature weights, assessing the computational performance of different query reformulation techniques. The results of their study suggest “greatly improved retrieval performance” when utilizing combined machine learning and bridged vocabularies. More so, they provide insight regarding the quality of options that can support computational systems for health informatics search tasks.
From the above research, we can distill two criteria. First, designs should provide interactions that allow users to efficiently prepare, perform, assess, and adjust their machine learning to align with information-seeking objectives of search tasks. Second, designs should provide mediation opportunities that assist users in communicating information-seeking objectives into the domain-specific vocabulary of the document set.
High-level criteria
In Table 1, we provide five criteria based on the above review.
| ||||||||||||||
Analysis of traditional interface strategies for health informatics search tasks
We now assess the traditional design strategies for interfaces for health informatics search tasks. Wilson’s comprehensive Search User Interface Design[46] provides a complete survey of the history and current state of search interfaces. Based on their survey, and in particular their discussion of input and control features within the modern search user interfaces, two base strategies and one extension strategy for search interfaces are realized: “structured” interfaces, “unstructured” interfaces, and, in extension, “query expansion” interfaces. Table 2 provides a summary of how the above criteria align with each interface strategy.
| ||||||||||||||||||||||||||||
Structured interface strategy
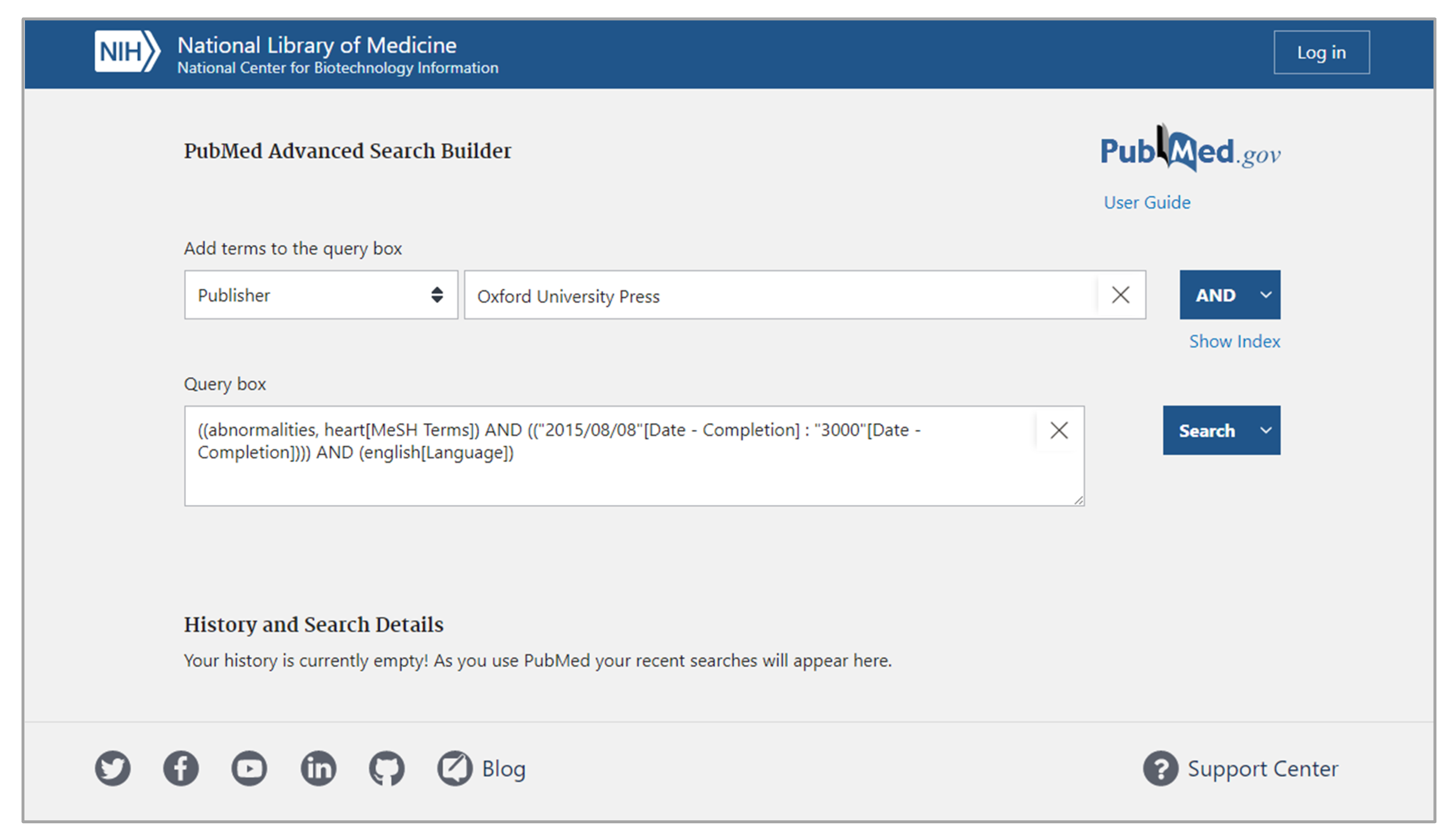
The structured interface strategy creates designs that regulate input during query building. This is achieved by maintaining heavily restricted input control profiles. Designers who implement the structured interface strategy into their interfaces presuppose a search task with specific expectations for input, bounding queries to a limited input profile. One common bounding technique is to constrain query lengths and limiting query content to a controlled set of terms.[47] This restricted scope is considered the sole acceptable input profile, and thus it allows designers to generate interfaces that limit the possible range of inputs and restrict all inputs that fall outside of that range. Designers typically achieve this by using interface elements like dropdowns, checkboxes, and radio buttons instead of elements like text boxes with free typing. For example, Figure 1 depicts the PubMed Advance Search Builder, which implements the structured interface strategy in its design. This interface requires users to select specific query term types from a restricted list, which then guides user input.[48]
|
Since input control is restricted, a strength of the structured interface strategy is that designers can use information characteristics to prescribe the full range of query formulations. This allows for the use of representational and computational designs that optimize for the expected characteristics of the restricted input profiles, per DC2. This strategy provides a designer-friendly environment that is hardened against unwanted queries, which, if effectively communicated in the design of result representations, could allow for alignment with DC3 and DC4. Yet, it can be challenging to designers to use structured interface strategies in a generalized setting. This is because when a document set is swapped, hardened approaches may not align with the information characteristics of the new document set. This negatively affects the flexibility of the interface, and in turn alignment with DC1. A potential weakness of the structured interface strategy is that it requires users to possess expertise on both the controlled vocabulary of the interface as well as the vocabulary of the document set being searched. If this is not known, user experience can suffer, drastically affecting alignment with DC5. Within the context of health informatics, such weaknesses reduce the users’ ability to effectively perform search tasks. This is because the controlled vocabularies within the health and medical domains demand significant expertise and result in numerous points of failure during the query formation process.[49][50]
Unstructured interface strategy
The unstructured interface strategy creates designs that provide limited input regulation. Unlike the structured interface strategy, it provides an open input control that accepts most input profiles during query building. Designers who implement the unstructured interface strategy do so without presupposing particular input, only accounting for general user error. That is, this input can originate from anywhere, such as common vocabulary, rather than from a pre-determined set of terms provided by the designer. Often, this input is directed to a single interface element. Implementations of the unstructured interface strategy typically present a text box that allows users to freely type their own text into the interface. These implementations will perform some input processing prior to use; however, the presentation of this processing to users is usually limited to correcting typographical errors rather than semantic ones. For example, the interface of Google aligns with the unstructured interface strategy, presenting users with an open, text-box input control without domain-specific assumptions or requirements. Of course, Google’s computational systems use extensive processing between receiving input from users and presenting the results of computation back to users.[51] Yet, users themselves are not informed of how their results came to be, even after changing to Google Instant.[52] Another example of an implementation of the unstructured interface strategy is WebMD’s search interface. This interface processes a free-text input with basic sanitization techniques before generating features for its search engine system, as depicted in Figure 2.
References
- ↑ 1.0 1.1 1.2 Wickramasinghe, Nilmini (2019/08). "Essential Considerations for Successful Consumer Health Informatics Solutions" (in en). Yearbook of Medical Informatics 28 (01): 158–164. doi:10.1055/s-0039-1677909. ISSN 0943-4747. PMC PMC6697544. PMID 31419828. http://www.thieme-connect.de/DOI/DOI?10.1055/s-0039-1677909.
- ↑ 2.0 2.1 2.2 Canadian Medical Association (2018). "The Future of Technology in Health and Health Care: A Primer". Canadian Medical Association. Archived from the original on 30 April 2019. https://web.archive.org/web/20190430220959/https://www.cma.ca/sites/default/files/pdf/health-advocacy/activity/2018-08-15-future-technology-health-care-e.pdf.
- ↑ 3.0 3.1 3.2 Demiris, G. (2016). "Consumer Health Informatics: Past, Present, and Future of a Rapidly Evolving Domain" (in en). Yearbook of Medical Informatics 25 (S 01): S42–S47. doi:10.15265/IYS-2016-s005. ISSN 0943-4747. PMC PMC5171509. PMID 27199196. http://www.thieme-connect.de/DOI/DOI?10.15265/IYS-2016-s005.
- ↑ 4.0 4.1 Zuccon, G.; Koopman, B. (2014). Goeuriot, L.; Jones, G.J.F.; Kelly, L. et al.. ed. "Integrating understandability in the evaluation of consumer health search engines". Proceedings of the Medical Information Retrieval Workshop at SIGIR 2014 (MedIR@SIGIR 2014) 1276: 32–35. http://ceur-ws.org/Vol-1276/.
- ↑ Chinese Journal of Cancer (1 December 2017). "The 150 most important questions in cancer research and clinical oncology series: questions 67–75: Edited by Chinese Journal of Cancer" (in en). Chinese Journal of Cancer 36 (1): 86, s40880–017–0254-z. doi:10.1186/s40880-017-0254-z. ISSN 1944-446X. PMC PMC5664810. PMID 29092716. https://cancercommun.biomedcentral.com/articles/10.1186/s40880-017-0254-z.
- ↑ Mehta, N.; Pandit, A. (1 June 2018). "Concurrence of big data analytics and healthcare: A systematic review" (in en). International Journal of Medical Informatics 114: 57–65. doi:10.1016/j.ijmedinf.2018.03.013. ISSN 1386-5056. https://www.sciencedirect.com/science/article/abs/pii/S1386505618302466.
- ↑ Thiébaut, Rodolphe; Cossin, Sébastien; Informatics, Section Editors for the IMIA Yearbook Section on Public Health and Epidemiology (2019/08). "Artificial Intelligence for Surveillance in Public Health" (in en). Yearbook of Medical Informatics 28 (01): 232–234. doi:10.1055/s-0039-1677939. ISSN 0943-4747. PMC PMC6697516. PMID 31419837. http://www.thieme-connect.de/DOI/DOI?10.1055/s-0039-1677939.
- ↑ 8.0 8.1 Saleemi, M.M.; Rodríguez, N.D.; Lilius, J.; Porres, I. (2011). "A Framework for Context-aware Applications for Smart Spaces". In Balandin, Sergey; Koucheryavy, Yevgeni; Hu, Honglin et al.. Smart Spaces and Next Generation Wired/Wireless Networking. Lecture notes in computer science. Heidelberg: Springer. pp. 14–25. ISBN 978-3-642-22874-2. OCLC 844916767. https://www.worldcat.org/title/mediawiki/oclc/844916767.
- ↑ 9.0 9.1 Gibson, C. J.; Dixon, B. E.; Abrams, K. (2015). "Convergent evolution of health information management and health informatics" (in en). Applied Clinical Informatics 06 (01): 163–184. doi:10.4338/ACI-2014-09-RA-0077. ISSN 1869-0327. PMC PMC4377568. PMID 25848421. http://www.thieme-connect.de/DOI/DOI?10.4338/ACI-2014-09-RA-0077.
- ↑ 10.0 10.1 10.2 Fang, Ruogu; Pouyanfar, Samira; Yang, Yimin; Chen, Shu-Ching; Iyengar, S. S. (14 June 2016). "Computational Health Informatics in the Big Data Age: A Survey". ACM Computing Surveys 49 (1): 12:1–12:36. doi:10.1145/2932707. ISSN 0360-0300. https://doi.org/10.1145/2932707.
- ↑ Köhler, Sebastian; Carmody, Leigh; Vasilevsky, Nicole; Jacobsen, Julius O B; Danis, Daniel; Gourdine, Jean-Philippe; Gargano, Michael; Harris, Nomi L et al. (8 January 2019). "Expansion of the Human Phenotype Ontology (HPO) knowledge base and resources". Nucleic Acids Research 47 (D1): D1018–D1027. doi:10.1093/nar/gky1105. ISSN 0305-1048. PMC PMC6324074. PMID 30476213. https://doi.org/10.1093/nar/gky1105.
- ↑ 12.0 12.1 12.2 Carayon, Pascale; Hoonakker, Peter (2019/08). "Human Factors and Usability for Health Information Technology: Old and New Challenges" (in en). Yearbook of Medical Informatics 28 (01): 071–077. doi:10.1055/s-0039-1677907. ISSN 0943-4747. PMC PMC6697515. PMID 31419818. http://www.thieme-connect.de/DOI/DOI?10.1055/s-0039-1677907.
- ↑ Gamache, Roland; Kharrazi, Hadi; Weiner, Jonathan P. (2018/08). "Public and Population Health Informatics: The Bridging of Big Data to Benefit Communities" (in en). Yearbook of Medical Informatics 27 (01): 199–206. doi:10.1055/s-0038-1667081. ISSN 0943-4747. PMC PMC6115205. PMID 30157524. http://www.thieme-connect.de/DOI/DOI?10.1055/s-0038-1667081.
- ↑ Brewer, LaPrincess C.; Fortuna, Karen L.; Jones, Clarence; Walker, Robert; Hayes, Sharonne N.; Patten, Christi A.; Cooper, Lisa A. (14 January 2020). "Back to the Future: Achieving Health Equity Through Health Informatics and Digital Health" (in EN). JMIR mHealth and uHealth 8 (1): e14512. doi:10.2196/14512. PMC PMC6996775. PMID 31934874. https://mhealth.jmir.org/2020/1/e14512.
- ↑ 15.0 15.1 Wu, Charley M.; Meder, Björn; Filimon, Flavia; Nelson, Jonathan D. (1 August 2017). "Asking better questions: How presentation formats influence information search." (in en). Journal of Experimental Psychology: Learning, Memory, and Cognition 43 (8): 1274–1297. doi:10.1037/xlm0000374. ISSN 1939-1285. http://doi.apa.org/getdoi.cfm?doi=10.1037/xlm0000374.
- ↑ 16.0 16.1 Talbot, Justin; Lee, Bongshin; Kapoor, Ashish; Tan, Desney S. (4 April 2009). "EnsembleMatrix: Interactive visualization to support machine learning with multiple classifiers" (in en). Proceedings of the SIGCHI Conference on Human Factors in Computing Systems (Boston MA USA: ACM): 1283–1292. doi:10.1145/1518701.1518895. ISBN 978-1-60558-246-7. https://dl.acm.org/doi/10.1145/1518701.1518895.
- ↑ 17.0 17.1 Hohman, Fred; Kahng, Minsuk; Pienta, Robert; Chau, Duen Horng (1 August 2019). "Visual Analytics in Deep Learning: An Interrogative Survey for the Next Frontiers". IEEE Transactions on Visualization and Computer Graphics 25 (8): 2674–2693. doi:10.1109/TVCG.2018.2843369. ISSN 1941-0506. PMC PMC6703958. PMID 29993551. https://ieeexplore.ieee.org/document/8371286/.
- ↑ 18.0 18.1 Yuan, Jun; Chen, Changjian; Yang, Weikai; Liu, Mengchen; Xia, Jiazhi; Liu, Shixia (25 November 2020). "A survey of visual analytics techniques for machine learning" (in en). Computational Visual Media 7 (1): 3–36. doi:10.1007/s41095-020-0191-7. ISSN 2096-0433. https://doi.org/10.1007/s41095-020-0191-7.
- ↑ 19.0 19.1 Endert, A.; Ribarsky, W.; Turkay, C.; Wong, B. L. William; Nabney, I.; Blanco, I. Díaz; Rossi, F. (2017). "The State of the Art in Integrating Machine Learning into Visual Analytics" (in en). Computer Graphics Forum 36 (8): 458–486. doi:10.1111/cgf.13092. ISSN 1467-8659. https://onlinelibrary.wiley.com/doi/abs/10.1111/cgf.13092.
- ↑ 20.0 20.1 Jusoh, S; Awajan, A; Obeid, N (1 May 2020). "The Use of Ontology in Clinical Information Extraction". Journal of Physics: Conference Series 1529: 052083. doi:10.1088/1742-6596/1529/5/052083. ISSN 1742-6588. https://iopscience.iop.org/article/10.1088/1742-6596/1529/5/052083.
- ↑ 21.0 21.1 Lytvyn, Vasyl; Dosyn, Dmytro; Vysotska, Victoria; Hryhorovych, Andrii (1 August 2020). "Method of Ontology Use in OODA". 2020 IEEE Third International Conference on Data Stream Mining & Processing (DSMP) (Lviv, Ukraine: IEEE): 409–413. doi:10.1109/DSMP47368.2020.9204107. ISBN 978-1-7281-3214-3. https://ieeexplore.ieee.org/document/9204107/.
- ↑ 22.0 22.1 Román-Villarán, E.; Pérez-Leon, F. P.; Escobar-Rodriguez, G. A.; Martínez-García, A.; Álvarez-Romero, C.; Parra-Calderón, C. L. (21 August 2019). "An Ontology-Based Personalized Decision Support System for Use in the Complex Chronically Ill Patient". Studies in Health Technology and Informatics 264: 758–762. doi:10.3233/SHTI190325. ISSN 1879-8365. PMID 31438026. https://pubmed.ncbi.nlm.nih.gov/31438026.
- ↑ Sacha, D.; Sedlmair, M.; Zhang L.et al. (2016). "Human-centered Machine Learning Through Interactive Visualization: Review and Open Challenges". In Verleysen, Michel. 24th European Symposium on Artificial Neural Networks, Computational Intelligence and Machine Learning: ESANN 2016. ESSAN, Université catholique de Louvain, Katholieke Universiteit Leuven. Louvain-la-Neuve, Belgique: Ciaco - i6doc.com. pp. 641–646. ISBN 978-2-87587-027-8. OCLC 964654436. https://www.worldcat.org/title/mediawiki/oclc/964654436.
- ↑ Amershi, Saleema; Cakmak, Maya; Knox, William Bradley; Kulesza, Todd (22 December 2014). "Power to the People: The Role of Humans in Interactive Machine Learning" (in en). AI Magazine 35 (4): 105–120. doi:10.1609/aimag.v35i4.2513. ISSN 2371-9621. https://ojs.aaai.org/index.php/aimagazine/article/view/2513.
- ↑ 25.0 25.1 25.2 Zeng, Qing T.; Tse, Tony (1 January 2006). "Exploring and Developing Consumer Health Vocabularies". Journal of the American Medical Informatics Association 13 (1): 24–29. doi:10.1197/jamia.M1761. ISSN 1067-5027. PMC PMC1380193. PMID 16221948. https://doi.org/10.1197/jamia.M1761.
- ↑ 26.0 26.1 26.2 26.3 Arp, Robert; Smith, Barry; Spear, Andrew D. (2015). Building ontologies with Basic Formal Ontology. Cambridge, Massachusetts: Massachusetts Institute of Technology. ISBN 978-0-262-52781-1.
- ↑ Bikakis, N.; Sellis, T. (2016). "Exploration and Visualization in the Web of Big Linked Data: A Survey of the State of the Art". arXiv. arXiv:1601.08059. https://arxiv.org/abs/1601.08059.
- ↑ Carpendale, Sheelagh; Chen, Min; Evanko, Daniel; Gehlenborg, Nils; Görg, Carsten; Hunter, Larry; Rowland, Francis; Storey, Margaret-Anne et al. (1 March 2014). "Ontologies in Biological Data Visualization". IEEE Computer Graphics and Applications 34 (2): 8–15. doi:10.1109/MCG.2014.33. ISSN 1558-1756. https://ieeexplore.ieee.org/document/6777435/.
- ↑ Dou, Dejing; Wang, Hao; Liu, Haishan (1 February 2015). "Semantic data mining: A survey of ontology-based approaches". Proceedings of the 2015 IEEE 9th International Conference on Semantic Computing (IEEE ICSC 2015) (Anaheim, CA, USA: IEEE): 244–251. doi:10.1109/ICOSC.2015.7050814. ISBN 978-1-4799-7935-6. http://ieeexplore.ieee.org/document/7050814/.
- ↑ Livingston, Kevin M.; Bada, Michael; Baumgartner, William A.; Hunter, Lawrence E. (23 April 2015). "KaBOB: ontology-based semantic integration of biomedical databases". BMC Bioinformatics 16 (1): 126. doi:10.1186/s12859-015-0559-3. ISSN 1471-2105. PMC PMC4448321. PMID 25903923. https://doi.org/10.1186/s12859-015-0559-3.
- ↑ Salkic, S.; Softic, S.; Taraghi, B. Ebner, M. (2015). "Linked data driven visual analytics for tracking learners in a PLE". In Pongratz, Hans; Gesellschaft für Informatik. DeLFI 2015 - die 13. E-Learning Fachtagung Informatik der Gesellschaft für Informatik e.V: 1.-4. September 2015 München, Deutschland. GI-Edition Lecture Notes in Informatics Proceedings. Bonn: Ges. für Informatik. pp. 329–331. ISBN 978-3-88579-641-1. OCLC 927803453. https://www.worldcat.org/title/mediawiki/oclc/927803453.
- ↑ Jakus, Grega; Milutinović, Veljko; Omerović, Sanida; Tomažič, Sašo (2013). Concepts, ontologies, and knowledge representation. SpringerBriefs in computer science. New York: Springer. ISBN 978-1-4614-7821-8. OCLC 841495258. https://www.worldcat.org/title/mediawiki/oclc/841495258.
- ↑ Rector, A.; Schulz, S.; Rodrigues, J.M. et al. (1 January 2019). "On beyond Gruber: “Ontologies” in today’s biomedical information systems and the limits of OWL" (in en). Journal of Biomedical Informatics 100 Suppl.. doi:10.1016/j.yjbinx.2019.100002. ISSN 1532-0464. https://www.sciencedirect.com/science/article/pii/S2590177X19300010.
- ↑ Rodríguez, Natalia Díaz; Cuéllar, M. P.; Lilius, Johan; Calvo-Flores, Miguel Delgado (1 April 2014). "A survey on ontologies for human behavior recognition" (in en). ACM Computing Surveys 46 (4): 1–33. doi:10.1145/2523819. ISSN 0360-0300. https://dl.acm.org/doi/10.1145/2523819.
- ↑ Katifori, Akrivi; Torou, Elena; Vassilakis, Costas; Lepouras, Georgios; Halatsis, Constantin (1 June 2008). "Selected results of a comparative study of four ontology visualization methods for information retrieval tasks". 2008 Second International Conference on Research Challenges in Information Science (Marrakech: IEEE): 133–140. doi:10.1109/RCIS.2008.4632101. ISBN 978-1-4244-1677-6. http://ieeexplore.ieee.org/document/4632101/.
- ↑ Raghupathi, Wullianallur; Raghupathi, Viju (7 February 2014). "Big data analytics in healthcare: promise and potential" (in en). Health Information Science and Systems 2 (1). doi:10.1186/2047-2501-2-3. ISSN 2047-2501. PMC PMC4341817. PMID 25825667. https://doi.org/10.1186/2047-2501-2-3.
- ↑ Russell-Rose, T.; Chamberlain, J.; Azzopardi, L. (2018). "Information retrieval in the workplace: A comparison of professional search practices" (in en). Information Processing & Management 54 (6): 1042–1057. doi:10.1016/j.ipm.2018.07.003. ISSN 0306-4573. https://www.sciencedirect.com/science/article/abs/pii/S0306457318300220.
- ↑ Russell-Rose, Tony; Chamberlain, Jon (2 October 2017). "Expert Search Strategies: The Information Retrieval Practices of Healthcare Information Professionals" (in EN). JMIR Medical Informatics 5 (4): e7680. doi:10.2196/medinform.7680. PMC PMC5643841. PMID 28970190. https://medinform.jmir.org/2017/4/e33.
- ↑ Huurdeman, H.C. (2017). "Dynamic Compositions: Recombining Search User Interface Features for Supporting Complex Work Tasks". Proceedings of the Second Workshop on Supporting Complex Search Tasks 1798: 22–25. http://ceur-ws.org/Vol-1798/.
- ↑ Zahabi, Maryam; Kaber, David B.; Swangnetr, Manida (1 August 2015). "Usability and Safety in Electronic Medical Records Interface Design: A Review of Recent Literature and Guideline Formulation" (in en). Human Factors 57 (5): 805–834. doi:10.1177/0018720815576827. ISSN 0018-7208. https://doi.org/10.1177/0018720815576827.
- ↑ Dudley, John J.; Kristensson, Per Ola (13 June 2018). "A Review of User Interface Design for Interactive Machine Learning". ACM Transactions on Interactive Intelligent Systems 8 (2): 8:1–8:37. doi:10.1145/3185517. ISSN 2160-6455. https://doi.org/10.1145/3185517.
- ↑ 42.0 42.1 Harvey, Morgan; Hauff, Claudia; Elsweiler, David (9 August 2015). "Learning by Example: Training Users with High-quality Query Suggestions" (in en). Proceedings of the 38th International ACM SIGIR Conference on Research and Development in Information Retrieval (Santiago Chile: ACM): 133–142. doi:10.1145/2766462.2767731. ISBN 978-1-4503-3621-5. https://dl.acm.org/doi/10.1145/2766462.2767731.
- ↑ Soldaini, Luca; Yates, Andrew; Yom-Tov, Elad; Frieder, Ophir; Goharian, Nazli (16 July 2015). "Enhancing web search in the medical domain via query clarification" (in en). Information Retrieval Journal 19 (1-2): 149–173. doi:10.1007/s10791-015-9258-y. ISSN 1386-4564. https://doi.org/10.1007/s10791-015-9258-y.
- ↑ Anderson, James D; Wischgoll, Thomas (26 January 2020). "Visualization of Search Results of Large Document Sets". Electronic Imaging 2020 (1): 388–1–388-7. doi:10.2352/ISSN.2470-1173.2020.1.VDA-388. https://www.ingentaconnect.com/content/ist/ei/2020/00002020/00000001/art00006;jsessionid=1ui77rulkmwwg.x-ic-live-03.
- ↑ Soldaini, Luca; Cohan, Arman; Yates, Andrew; Goharian, Nazli; Frieder, Ophir (2015), Hanbury, Allan; Kazai, Gabriella; Rauber, Andreas et al.., eds., "Retrieving Medical Literature for Clinical Decision Support" (in en), Advances in Information Retrieval (Cham: Springer International Publishing) 9022: 538–549, doi:10.1007/978-3-319-16354-3_59, ISBN 978-3-319-16353-6, http://link.springer.com/10.1007/978-3-319-16354-3_59
- ↑ Wilson, Max L. (2012). Search user interface design. Synthesis lectures on information concepts, retrieval, and services. San Rafael, Calif.: Morgan & Claypool. ISBN 978-1-60845-689-5. OCLC 780340844. https://www.worldcat.org/title/mediawiki/oclc/780340844.
- ↑ Zielstorff, R.D. (2003). "Controlled vocabularies for consumer health" (in en). Journal of Biomedical Informatics 36 (4-5): 326–333. doi:10.1016/j.jbi.2003.09.015. ISSN 1532-0464. https://www.sciencedirect.com/science/article/pii/S1532046403000960.
- ↑ National Center for Biotechnology Information. "PubMed.gov". National Institutes of Health. https://pubmed.ncbi.nlm.nih.gov/. Retrieved 18 January 2021.
- ↑ McCray, Alexa T.; Tse, Tony (2003). "Understanding Search Failures in Consumer Health Information Systems". AMIA Annual Symposium Proceedings 2003: 430–434. ISSN 1942-597X. PMC 1479930. PMID 14728209. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1479930/.
- ↑ Keselman, Alla; Browne, Allen C.; Kaufman, David R. (1 July 2008). "Consumer Health Information Seeking as Hypothesis Testing". Journal of the American Medical Informatics Association 15 (4): 484–495. doi:10.1197/jamia.M2449. ISSN 1067-5027. PMC PMC2442260. PMID 18436912. https://doi.org/10.1197/jamia.M2449.
- ↑ Luo, Jake; Wu, Min; Gopukumar, Deepika; Zhao, Yiqing (1 January 2016). "Big Data Application in Biomedical Research and Health Care: A Literature Review" (in en). Biomedical Informatics Insights 8: BII.S31559. doi:10.4137/BII.S31559. ISSN 1178-2226. PMC PMC4720168. PMID 26843812. https://doi.org/10.4137/BII.S31559.
- ↑ Qvarfordt, Pernilla; Golovchinsky, Gene; Dunnigan, Tony; Agapie, Elena (28 July 2013). "Looking ahead: query preview in exploratory search" (in en). Proceedings of the 36th international ACM SIGIR conference on Research and development in information retrieval (Dublin Ireland: ACM): 243–252. doi:10.1145/2484028.2484084. ISBN 978-1-4503-2034-4. https://dl.acm.org/doi/10.1145/2484028.2484084.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. Some grammar and punctuation was cleaned up to improve readability. In some cases important information was missing from the references, and that information was added.