Journal:Fueling clinical and translational research in Appalachia: Informatics platform approach
| Full article title | Fueling clinical and translational research in Appalachia: Informatics platform approach |
|---|---|
| Journal | JMIR Medical Informatics |
| Author(s) | Cecchetti, Alfred A.; Bhardwaj, Niharika; Murughiyan, Usha; Kothakapu, Gouthami; Sundaram, Uma |
| Author affiliation(s) | Joan C. Edwards School of Medicine at Marshall University |
| Primary contact | Email: cecchetti at marshall dot edu |
| Editors | Eysenbach, G. |
| Year published | 2020 |
| Volume and issue | 8(10) |
| Article # | e17962 |
| DOI | 10.2196/17962 |
| ISSN | 2291-9694 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://medinform.jmir.org/2020/10/e17962/ |
| Download | https://medinform.jmir.org/2020/10/e17962/PDF (PDF) |
Abstract
Background: The Appalachian population is distinct, not just culturally and geographically but also in its healthcare needs, facing the most health care disparities in the United States. To meet these unique demands, Appalachian medical centers need an arsenal of analytics and data science tools with the foundation of a centralized data warehouse to transform healthcare data into actionable clinical interventions. However, this is an especially challenging task given the fragmented state of medical data within Appalachia and the need for integration of other types of data such as environmental, social, and economic with medical data.
Objective: This paper aims to present the structure and process of the development of an integrated platform at a midlevel Appalachian academic medical center, along with its initial uses.
Methods: The Appalachian Informatics Platform (AIP) was developed by the Appalachian Clinical and Translational Science Institute’s Division of Clinical Informatics and consists of four major components: a centralized clinical data warehouse, modeling (statistical and machine learning), visualization, and model evaluation. Data from different clinical systems, billing systems, and state- or national-level data sets were integrated into a centralized data warehouse. The platform supports research efforts by enabling curation and analysis of data using the different components, as appropriate.
Results: The AIP is functional and has supported several research efforts since its implementation for a variety of purposes, such as increasing knowledge of the pathophysiology of diseases, risk identification, risk prediction, and healthcare resource utilization research and estimation of the economic impact of diseases.
Conclusions: The platform provides an inexpensive yet seamless way to translate clinical and translational research ideas into clinical applications for regions similar to Appalachia that have limited resources and a largely rural population.
Keywords: Appalachian region, medical informatics, health care disparities, electronic health records, data warehousing, data mining, data visualization, machine learning, data science
Introduction
Background: Unique challenges in Appalachia
Appalachia, with its predominantly rural communities, is known to have one of the worst sets of healthcare outcomes in the United States. This is especially true of southern and central rural Appalachia, which face some of the most severe health disparities in the nation.[1] Over the years, the gap in the overall health between Appalachia and the nation as a whole has continued to grow.[2][3] To close this gap, it is critical to identify the cause of these disparities and direct efforts toward developing necessary interventions to address them.
Such an effort necessitates the adoption of modern technologies such as a centralized research data warehouse to house all data necessary to obtain a comprehensive picture of the health of the Appalachian population before analysis to gain actionable insights can be performed. A centralized data warehouse, once considered strictly a business tool, has evolved into an important instrument for cost containment, tracking of patient outcomes, providing clinical decision support at the point of care, improving prognostic accuracy, and facilitating research.[4] Thus, rural academic medical centers have moved toward implementing data warehouse systems that feed analytical systems for research needs.[5] This entails (1) the integration of data from different types of medical settings (i.e., multi-institutional) such as hospitals, clinics, and specialty centers; (2) linkage of financial data with clinical data, a well-established practice proven to be pivotal to high-quality care and great economic outcomes[6][7]; and (3) integration of other determinants of health such as environmental[8], social[9], and spiritual factors[10] to create longitudinal health records across the care continuum.
However, there are challenges in creating a multi-institutional data warehouse.[11] Electronic health records (EHRs) do not easily interact with one another due to the use of nonstandard terminologies and difficulty in understanding the flow of information. Additionally, significant differences exist between rural and urban health systems.[12][13][14][15][16] Unlike their urban counterparts, healthcare data in Appalachia are typically fragmented, existing in silos within dissimilar databases, registries, data collections, and departmental systems. With innovations in medical technology, the list of data sources continues to grow, producing unprecedented amounts of data from all aspects of care, including diagnosis, medication, procedures, laboratory testing, imaging, and patient self-monitoring.[17][18][19][20][21] To complicate matters, the overall health and health behaviors of Appalachians are strongly affected by Appalachia’s unique culture, geography, and health system issues.[22][23][24] Consequently, Appalachian academic medical centers face the complex challenge of collecting, organizing, standardizing, and analyzing these enormous quantities of heterogeneous data originating from a wide variety of sources to address the unmet needs of the population they serve.
Why an informatics platform?
Data integration and interoperability have been shown to be key to unlocking these data for data analytics, enabling the development of novel patient management strategies for rural hospitals[25][26] and translational research that leads to new approaches at the bedside for prevention, diagnosis, and treatment of disease, which are essential to improving the health of a population.[27][28][29] Data analytics, once the domain of the statistician, has now become an equal partner in clinical research and research operations.[30][31] Following the data explosion, data analytics increasingly involves the use of visual analytics tools such as Tableau (Tableau Software Inc.) and Power BI (Microsoft Corp.) to explore data easily and in a self-service fashion and to clearly and effectively communicate complex ideas[32], especially to those members of the medical community who might not have an intimate understanding of the underlying data. Furthermore, machine learning is gaining importance, especially in the area of predictive analytics, to improve the practice of medicine and to infer potentially innovative risk factors.[28][33][34][35]
However, these applications (e.g., data warehouse, data analytics, statistical analysis, machine learning, visual analytics) are generally uncoordinated without any overarching governance. Thus, we developed an informatics platform—that is, a suite of interconnected, coordinated applications hosted within an operational environment[36]—called the Appalachian Informatics Platform (AIP), in West Virginia (the only state located entirely in Appalachia) that facilitates interoperable access to integrated information, data visualization, and data analytics, thereby functioning as an excellent basis for clinical and translational research to improve health care.
The goal of this study is to describe the structure and process of development of the AIP and demonstrate its value in supporting clinical and translational research.
Methods
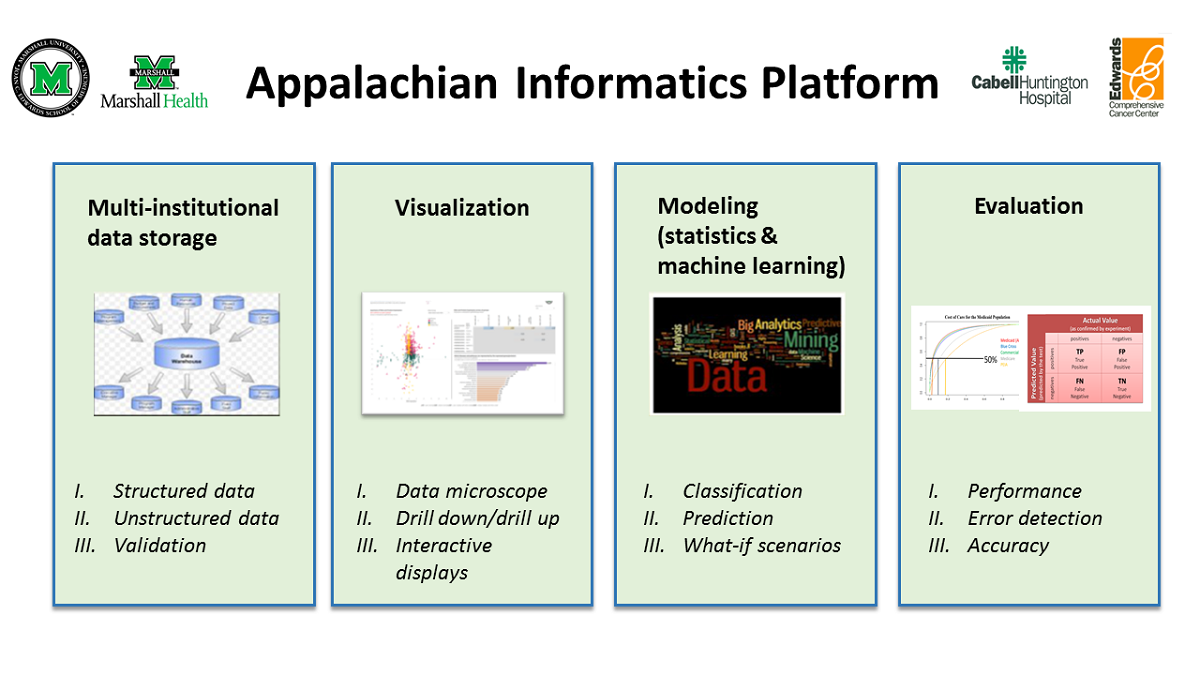
The AIP (Figure 1) is composed of four major components: (1) a multi-institutional data storage or clinical data warehouse (CDW); (2) modeling tools (statistical and machine learning); (3) visualization tools; and (4) evaluation tools. Each of these components is described in detail in separate sections.
The CDW forms an integral part of the AIP. It also contains embedded data analytics (modeling and evaluation) and interactive visualization tools (e.g., Tableau, Power BI). Together, these enable the analysis of Appalachian health information to speed up the transition of translational research ideas into clinical practice.
The CDW serves as a secure source of quality data for descriptive, diagnostic, predictive, and prescriptive analytics for research and operational needs. The visual analytics tools enable an initial exploratory analysis of the processed data and the interactive presentation of analytical findings for further analysis and review. Depending on the use case, data can be analyzed using statistical modeling via external (e.g., SPSS [IBM Corp], Stata [StataCorp]) or integrated (e.g., R [R Foundation for Statistical Computing], Python [Python Software Foundation] in SQL [Structured Query Language]) applications or machine learning modeling. The performance of the resulting models was evaluated using appropriate metrics. Once trained and evaluated, machine learning models can be deployed and stored in the CDW for future use if needed. Furthermore, the stored machine learning models can be continuously evaluated and improved as more data are generated.
|
The informatics committee governs the access to and utilization of AIP and ensures adherence to security and privacy rules. In addition, team-building activities are also incorporated into our clinical informatics model to foster the development of an effective clinical informatics team.
Multi-institutional data storage: The Appalachian Clinical and Translational Science Institute-Clinical Data Warehouse
The Appalachian Clinical and Translational Science Institute (ACTSI)’s Division of Clinical Informatics solicited buy-in from different entities—namely Cabell-Huntington Hospital (CHH), Edwards Comprehensive Cancer Institute (ECCC), Marshall Health (MH) practice plan, and Marshall University Joan C. Edwards School of Medicine (MU JCESOM)—to build the Appalachian Clinical and Translational Science Institute-Clinical Data Warehouse (ACTSI-CDW) in West Virginia. An agreement was created between these entities that provided access to both financial and clinical data.
The multi-institutional CDW contains more than nine years of billing and clinical data. It comprises relational tables and dimension and fact tables (Online Analytical Processing [OLAP] cube), which enable secure data storage and data access. Designed from the start to facilitate information flow, the CDW can send out a stream of near real-time data that can be used for any authorized research purpose. Documentation includes a data dictionary and flowcharts. Flowcharts follow the patient from admission (or appointment, if outpatient) to discharge (or exit, if outpatient). The data dictionary contains the standardized and source field names, descriptions, and properties, along with the associated metadata for the data contained within the data warehouse. For instance, (1) the entry of a patient into any medical service (admission or appointment) was combined with the single term encounter, and (2) a higher level of precision was introduced by separating patient age into two variables: current age or the age when the procedure was performed.
The CDW process is based on an older data warehouse process developed at the University of Pittsburgh.[37] The process is as follows:
- Data dictionaries are created by recording institutional source field names and field properties and linking them to the standardized CDW names and properties found within the CDW databases. Descriptions of each field (source and CDW) are included.
- Individual institutional flowcharts show the workflow of the data and the location of the people responsible for the quality of the data, which are also used for quality control purposes.
- At present, the CDW contains data from six institutional software packages hosted in various parts of the country (e.g., Cerner data from Kansas City, Missouri; McKesson data from North Druid Hills, Georgia; etc.). The data are exported in a standard format (i.e., ASCII flat file, XML, etc.) and transferred through secure file transfer protocol (e.g., Cerberus [Cerberus, LLC]) to the CDW Development server.
- The data are integrated into the Microsoft SQL databases using Microsoft SQL Server Integration Services (SSIS), a graphical tool that extracts, transforms, and loads (ETL) the data to target schemas that will be used to contain the target data objects: relational tables, dimensions, and cubes. ETL systems enable a smooth migration from one system to another irrespective of the underlying storage system.
- Conformed dimensions were developed, and patient linkages using various methods (e.g., simple heuristics)[38] were also available and made at this time.
- At present, a transactional grain fact table has been developed, but other fact tables will be created as needed.
- The CDW contains internal structured billing and EHR data (i.e., demographics, encounter details, vitals, medications, procedures, diagnoses, orders, immunizations, laboratory and imaging results, date and time, payee, and provider). It also contains unstructured EHR data (e.g., H&P, admission notes, discharge summaries, other clinical notes). These data are received from MH, CHH, and MU JCESOM’s ECCC, as well as from other outside institutions. In addition, non-EHR data are incorporated using REDCap.
- Unstructured data are analyzed using text analytics tools, and classification variables based on text mining are incorporated into the CDW.
- The data structure (OLAP cubes and relational tables), once checked and verified, is transferred from the secure development server to the secure production server for use.
- Various security measures (e.g., IP and password restrictions) are in place to prevent unauthorized use.
- The CDW structure, which stores multi-institutional medical information, can now provide data for both operational and research analytical model development (statistical or machine learning) using very simple de-identified interfaces (e.g., Excel [Microsoft Corp]) or more complex interactive tools (e.g., R, Tableau, Power BI, etc.). Within the CDW, the data can be manipulated, cleaned, and prepared before the analysis as needed.
- Structured and unstructured data currently exist within the CDW. Image and BioSample data will soon be incorporated (like the Pittsburgh model), but the full design has not been finalized yet. An Honest Broker person assumes control of sample shipping and receiving.
- Standard operating procedures (SOPs) have been developed for administrative and technical areas.
- The Health Insurance Portability and Accountability Act (HIPAA) guidelines are followed, and protocol to protect patient information has also been implemented.
The CDW is contained within a Microsoft SQL database that can interact with outside objects using other electronic methods such as SignalR, a software library for Microsoft ASP.NET that allows server code to send asynchronous notifications to client-side web applications, and SqlDependency, an object that represents a query notification dependency between an application and an instance of SQL server. Objects such as these provide the ability for the data warehouse to interact in real time with the outside regional population using the newest technologies such as Microsoft Machine Learning Server with embedded R or Python procedure coding.
Data validation
The information derived from multiple data sources can have inconsistencies and missing values because of their heterogeneous nature, which requires correction.[39][40][41][42] Thus, for each research study, clinical and translational researchers using the data warehouse are required to verify a random sample (calculated on the basis of the size of the study population) of all extracted study data are directly verified at the original data source to ensure data accuracy and validity. Identified errors or omissions are transmitted back to the host systems for correction or inclusion.
Augmenting the CDW using REDCap
For certain studies, data available in the CDW may not be precise enough or include variables needed to perform this study. For such studies, data can be augmented using data capture tools. One such tool is the Research Electronic Data Capture, or REDCap, a workflow methodology and software solution designed for the rapid development and deployment of electronic data capture tools to support clinical and translational research.[43][44][45]
Our institution has deployed and maintains 2 REDCap servers: secure (located under institutional firewall) and global (outside the firewall). The secure REDCap system is used for storing data considered protected health information (PHI) under HIPAA. The global system, on the other hand, is used to store de-identified or non-PHI data. These data are then transferred to and stored within the multi-institutional data warehouse. This method of augmenting the information pulled from the existing source systems provides research-grade data from outside sources that are normally not contained within a data warehouse.
Visualization
Visualization of information is an excellent method of providing knowledge that can be easily understood by any member of the health care discipline. Within the informatics platform, Tableau provides interactive drill-down and drill-up capabilities for specific projects.
Tableau is a visual analytics tool that provides an interactive method of exploring multidimensional data, optimized from the data warehouse and OLAP data sources. Tableau, using either indexed relational tables or a data cube, can perform associated operations such as slice, dice, roll-up, and drill-down on the data, providing detailed interactive visual overlays that range from the lowest grain of the data to high-level representations of the data. Tableau charts, graphs, filters, and maps can provide visualization of the various subgroups of interest using a storyboard approach that presents a specific question followed by an interactive dashboard that explores that question in detail. The use of visual elements such as logos, pictograms, icons, or pictures into the dashboards, in association with the subgroups, provides easy-to-reference image aids that provide clarity and understanding of complex information. The data warehouse provides the drill-down, drill-up and slice and dice capability, whereas the hub design connects both financial and clinical data to provide a full picture.
The developed interactive dashboards are securely shared with users within a department or a team, as needed, through the use of Tableau Server.[46]
Modeling (statistics and machine learning)
The modeling component of the informatics platform supports the construction of tailored regional models (statistical or machine learning) to understand and predict disease and other medical events within this region. EHR is primarily a billing system, research only being a secondary function and, thus, is heterogeneous, incomplete, and noisy[25], leading to unrepresentative samples, selection bias, and misclassification.[47] During the modeling process, these issues are eliminated or minimized.
To assist in modeling, software packages such as SPSS and Stata, as well as embedded open-source machine learning programs (e.g., R, Python) are used. This enables faster and easier development of classification, regression, and clustering algorithms for research use. In addition, we utilize products such as Microsoft’s LINQ to electronically gather information and directly incorporate that information into the CDW.
Evaluation
During the modeling process, evaluation of the data set as it relates to the regional population is carried out. Local experts native to this region are asked to evaluate the model from a clinical as well as a financial standpoint. Poverty is endemic within the Appalachian population, and a model that suggests the use of a very expensive medication or procedure over an older but less expensive medication or procedure is unlikely to be used.[48] Thus, the model must take into account whether the patient has the means and access to the recommended medication or procedure.[49] In addition, the willingness of Appalachian medical institutions and health care providers to follow the model’s suggestions must also be evaluated.
Once developed, the models were tuned and tested. Location, time of treatment, outside temperature, and other contributory factors available within the CDW were employed to fine-tune the models, as applicable. The performance of the models was measured using the R programming environment using measures such as area under curve, sensitivity, specificity, F1 score, precision, recall, etc.
Security, privacy, and the informatics committee
Data access and usage are permitted only as described in the mutual agreement between the three institutions and are subject to internal security and privacy rules. All data requests must follow the standard operating procedure built on the basis of mutual multi-institutional agreement. Foremost, the researcher must have appropriate credentials and authorization to be able to request for data. If the researcher is authorized to make requests, he or she must obtain the IRB approval for his or her proposed study and submit the IRB proposal and supporting documentation for review by the informatics committee. The informatics committee, independent of the IRB, reviews all requests for data from the data warehouse to ensure compliance with the agreement. If the research project is approved, the research team designated members are scheduled for the de-identified data extraction process.
Team building
Integral to the informatics platform is team building that builds upon previous work.[37] To facilitate effective team meetings and inter-professional collaboration (local and global) without the need or expense of constant travel, a permanent clinical informatics conference room with a fixed connected computer, an uninterruptable power supply (UPS), a smart board, a camera, and a speaker system, along with a video conferencing system (Zoom) connectivity, was built. This ensures adequate communication among all those involved (i.e., team members, users, leadership, etc.) and access to resources that would otherwise be unavailable.
Results
Since the implementation of the platform, several studies have been conducted. Each study listed below was approved by the informatics committee, and the de-identified data and platform tools were made available securely to the research team.
To evaluate the functionality and value of this platform, we first analyzed the aggregated data of Medicaid-insured patients across different health systems using the interconnected applications within the platform for population health management. Relevant data were extracted from the CDW, followed by exploratory analysis using a Tableau dashboard. Due to the isolated nature of the study population, regional variables such as distance from the CHH and weather conditions (i.e., temperature) were also included. Errors and missing values were identified using the dashboard, and data were subsequently cleaned and prepared. Using these clean data, the regional population was classified into three spend categories: low cost, acute, and persistent subgroups on the basis of the charges accrued. Next, the Charlson Comorbidity Index (CCI) was incorporated into the CDW to predict mortality risk within one year of hospitalization for patients with comorbid conditions within each spend category (Table 1).[50][51]
| ||||||||||||
Of these categories, the persistent group had the largest percentage of patients with a high risk of mortality, followed by acute and low cost after excluding the deceased patients (persistent: 898/1247, 72.01%; acute: 2074/6946, 29.86%; low cost: 5130/102,814, 4.99%). The CCI was not very sensitive in predicting the risk of mortality but was very specific and accurate (sensitivity: 896/1512, 59.26%; specificity: 102,905/111,007, 92.7%; accuracy: 103,801/112,519, 92.25%). The effect of distance and weather on the CCI needs further investigation that is being conducted. Adjustments are being made to this standard national index to incorporate other Appalachian characteristics that could improve the sensitivity of this risk scoring system.
As such, the platform has been utilized for a variety of purposes such as increasing knowledge of the pathophysiology of diseases, risk identification, risk prediction, health care resource utilization research, and estimation of the economic impact of diseases to enable data-driven clinical decisions, leading to improved clinical outcomes. Table 2 contains a list of studies conducted so far.
|
Five studies utilized the platform for risk identification and risk prediction to improve diagnostic accuracy.[52][53][54][55][56] Sundaram et al.[56] demonstrated the value of ACTSI-CDW as a primary source to improve the diagnosis of metabolic syndrome, a diagnosis highly relevant to the Central Appalachian population. The researchers discovered that utilizing billing codes alone severely underestimated the number of patients with metabolic syndrome by a factor of more than 10, as compared with looking at specific criteria that determine this diagnosis.[56] Another study assessed the relationship between metabolic syndrome and colorectal cancer and found that patients with metabolic syndrome, especially those with insulin resistance, were more likely to have colorectal cancer, indicating the probable need for earlier screening for colorectal cancer in these patients.[55] Elmore et al.[54] examined the role of fever in predicting the development of secondary bacterial pneumonia in children with RSV and other viral illnesses. They found that febrile children were two to eight times (RSV, 47/78 vs 27/100; other bronchiolitis, 54/83 vs 7/88) more likely to have secondary bacterial pneumonia compared with afebrile children and, thus, may need to be aggressively evaluated to enable early diagnosis and treatment.[54] Amro et al.[52] studied the relationship between hypoalbuminemia and peripartum cardiomyopathy and noted that lower albumin levels were significantly associated with peripartum cardiomyopathy (P<.001; odds ratio 0.033, 95% CI 0.034-0.865) and could potentially be used as a risk marker for it. Acosta et al.[53] used data from the ACTSI-CDW to identify risk factors (lower BMI, before CHF, and prior use of aspirin) that predict myocardial injury, detectable troponin, and increase in serial troponin levels in patients with hypertensive crisis.
Ferdjallah et al.[59] analyzed the data from the ACTSI-CDW to understand how Alzheimer disease and diabetes mellitus affect serum calcium homeostasis and extracellular fluid volume. They observed that acute changes in serum calcium were significantly correlated with changes in extracellular fluid volume in both disease states.[59]
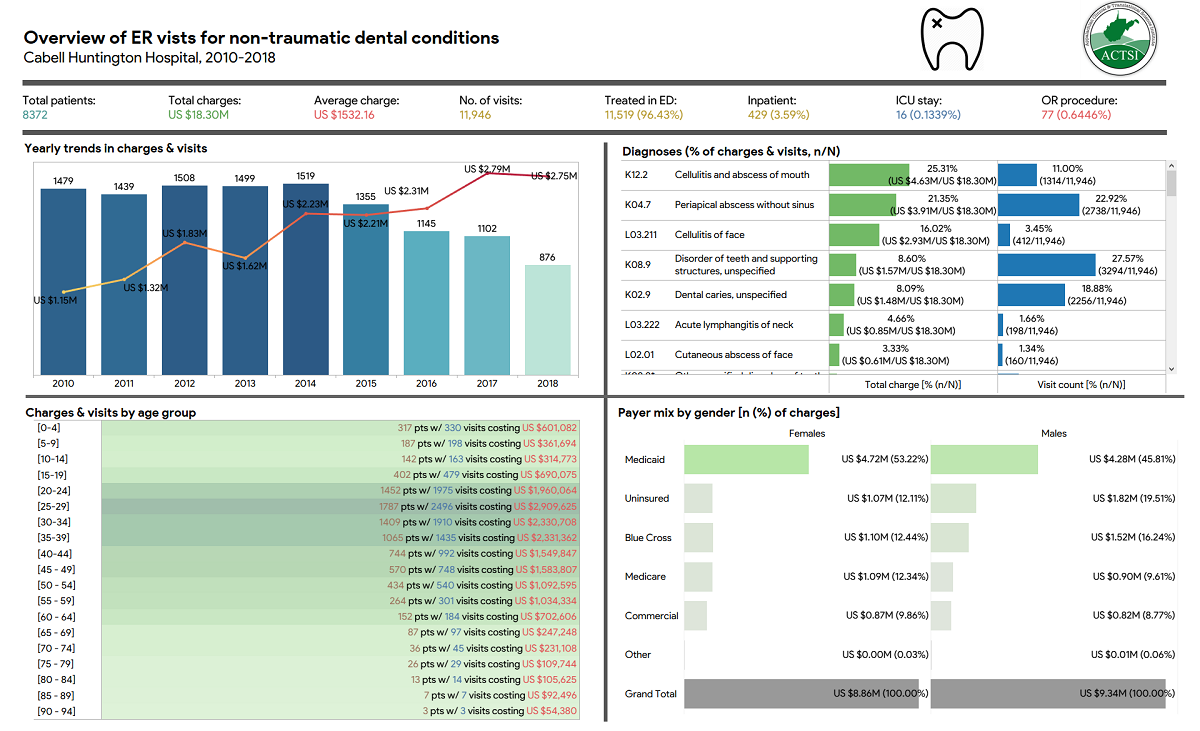
The platform has also been applied in two studies to assess resource utilization (e.g., emergency room, medications, etc.) and the financial impact of the disease. For instance, Bhardwaj et al.[57] utilized the platform to identify the problems associated with benzodiazepine use in geriatric patients within the health system, such as a higher number of emergency room visits and charges in geriatric patients with dementia plus at least one BZD prescription. In another study[58] that aimed to measure the volume and cost of emergency room use for these conditions and identify the factors that predict such use, the researchers built a dashboard (Figure 2) to easily explore and analyze relevant data on non-traumatic dental conditions that led to emergency room visits and to report the key findings of the study. The authors[58] observed that emergency room visits by uninsured patients were four times more likely and those by Medicaid insured two times more likely to be for dental problems than Medicare-insured patients.
|
Discussion
Utility of the AIP
The AIP has supported several research projects involving the use of different components of the platform, depending on project needs. The studies described reported findings that are seldom reported in this region, enhanced our knowledge of pathophysiology and risk factors, and helped estimate and analyze resource utilization and economic burden of certain diseases within Appalachia using minimal resources (i.e., a small IT team and a relatively inexpensive platform).
Before the implementation of the platform, many research studies that followed the patient across multiple care settings or involved analysis of big data were not possible due to the unavailability of technical and economic resources owing to a lack of buy-in from rural health care organizations. As the data existed in silos, there was a lack of standardization and normalization, which resulted in major data inconsistencies. Studies conducted using these disjointed data sets often used unrepresentative small biased samples and had low statistical power and quality.
The introduction of the platform has helped address these issues. It is now easier to pinpoint and correct errors and/or missing values and understand the distribution of data using visual analysis tools. Further, the time needed to conduct these studies from start to finish has been greatly reduced owing to the availability of all applications necessary to complete the study within the platform. This has been specifically useful because many researchers do not have the technical skills needed to perform complex and advanced data analysis, especially on larger data sets.
The paper also revealed that national models do not necessarily perform well when applied to the Appalachian population. The AIP allows for seamless integration of regional variables into the national model, which may improve the performance of these models. For each of the top 10 causes of death in West Virginia in 2017, per the Centers for Disease Control and Prevention[60], a machine learning algorithm was used to predict outcomes on a national level: heart disease[61][62], cancer[63][64], accidents[65][66], respiratory disease[67][68], stroke[69][70], diabetes[71][72], Alzheimer disease[73][74], pneumonia[75][76], kidney disease[77][78], and suicide.[79][80] Each of these cited models could be modified to fit the characteristics of the Appalachian population, especially those characteristics that make this region different in terms of geography, economy, education, and culture from the rest of the United States. The development of these regional models could help rural health general practitioners tackle complex medical conditions without the need for an expensive specialized health care provider nearby.[46]
We hope that this paper will help other rural health care organizations that serve underserved populations, such as ours, realize the value and ease of using an informatics platform to conduct research and improve care for their patients despite limited resources.
Ongoing projects and future directions
At present, a model that utilizes embedded data analytics to monitor the side effects of certain types of cancer by ingesting de-identified statements in the regional variety of English language from patients within this region[81][82] is under development. This model could be used to analyze patient responses at a certain point in time for a cross-sectional study or continuously in real time for a long-term longitudinal study to identify the patients in need of care before their scheduled follow-up visit. The ongoing results from this model would be sent to their health care providers for appropriate actions. In case of an emergency, patient-designated community support networks such as religious or other support groups may be intimated to bring the patient to the emergency department so that the patient can receive timely care.
We plan to expand upon our unified informatics platform to integrate programming applications for the development of state-of-the-art applications targeted specifically toward the unmet health care needs of the Appalachian population.
Conclusions
This paper establishes the value of the Appalachian Informatics Platform (AIP) in enabling seamless and secure data access, model development through an analytics engine to explore novel and unexpected hypotheses, and simple yet effective communication of all findings via interactive visualization.
The relatively inexpensive nature of such a platform coupled with its demonstrated advantages will hopefully encourage small and midsized rural academic centers, which traditionally have fewer resources than their urban counterparts, to adopt a research informatics platform within their institutions using the template described in this paper as a guide.
Abbreviations, acronyms, and initialisms
ACTSI: Appalachian Clinical and Translational Science Institute
AIP: Appalachian Informatics Platform
CCI: Charlson Comorbidity Index
CDW: clinical data warehouse
CHH: Cabell-Huntington Hospital
ECCC: Edwards Comprehensive Cancer Institute
EHR: electronic health record
ETL: extract, transform, and load
HIPAA: Health Insurance Portability and Accountability Act
MH: Marshall Health
MU JCESOM: Marshall University Joan C Edwards School of Medicine
OLAP: Online Analytical Processing
PHI: protected health information
SQL: structured query language
Acknowledgements
Funding
This work was supported by the National Institutes of Health grants DK-67420, DK-108054, and P20GM121299-01A1 and Veteran’s Administration Merit Review grant BX003443-01 to US and UL1TR00011719 to PK.
Conflicts of interest
None declared.
References
- ↑ Krometis, Leigh-Anne; Gohlke, Julia; Kolivras, Korine; Satterwhite, Emily; Marmagas, Susan West; Marr, Linsey C. (26 September 2017). "Environmental health disparities in the Central Appalachian region of the United States". Reviews on Environmental Health 32 (3): 253–66. doi:10.1515/reveh-2017-0012. ISSN 2191-0308. https://www.degruyter.com/document/doi/10.1515/reveh-2017-0012/html.
- ↑ Singh, Gopal K.; Kogan, Michael D.; Slifkin, Rebecca T. (1 August 2017). "Widening Disparities In Infant Mortality And Life Expectancy Between Appalachia And The Rest Of The United States, 1990–2013" (in en). Health Affairs 36 (8): 1423–1432. doi:10.1377/hlthaff.2016.1571. ISSN 0278-2715. http://www.healthaffairs.org/doi/10.1377/hlthaff.2016.1571.
- ↑ Marshall, J.L.; Thomas, L.; Lane, N.M. et al. (August 2017). "Health Disparities in Appalachia" (PDF). Creating a Culture of Health in Appalachia: Disparities and Bright Spots. Appalachian Regional Commission. https://www.arc.gov/wp-content/uploads/2020/06/Health_Disparities_in_Appalachia_August_2017.pdf. Retrieved 25 September 2020.
- ↑ Foran, David J; Chen, Wenjin; Chu, Huiqi; Sadimin, Evita; Loh, Doreen; Riedlinger, Gregory; Goodell, Lauri A; Ganesan, Shridar et al. (1 January 2017). "Roadmap to a Comprehensive Clinical Data Warehouse for Precision Medicine Applications in Oncology" (in en). Cancer Informatics 16: 117693511769434. doi:10.1177/1176935117694349. ISSN 1176-9351. PMC PMC5392017. PMID 28469389. http://journals.sagepub.com/doi/10.1177/1176935117694349.
- ↑ Kaufman, Arthur; Rhyne, Robert L.; Anastasoff, Juliana; Ronquillo, Francisco; Nixon, Marnie; Mishra, Shiraz; Poola, Charlene; Page-Reeves, Janet et al. (1 January 2017). "Health Extension and Clinical and Translational Science: An Innovative Strategy for Community Engagement" (in en). The Journal of the American Board of Family Medicine 30 (1): 94–99. doi:10.3122/jabfm.2017.01.160119. ISSN 1557-2625.
- ↑ Roberts, Mark S.; Dreese, Elizabeth M.; Hurley, Noreen; Zullo, Nan; Peterson, Mark (1991). "Blending Administrative and Clinical Needs: The Development of a Referring Physician Database and Automatic Referral Letter". Proceedings of the Annual Symposium on Computer Application in Medical Care: 559–563. ISSN 0195-4210. PMC 2464572. PMID 1807664. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2464572/.
- ↑ Raghupathi, Wullianallur; Raghupathi, Viju (1 December 2014). "Big data analytics in healthcare: promise and potential" (in en). Health Information Science and Systems 2 (1): 3. doi:10.1186/2047-2501-2-3. ISSN 2047-2501. PMC PMC4341817. PMID 25825667. http://link.springer.com/10.1186/2047-2501-2-3.
- ↑ Ahern, Melissa M.; Hendryx, Michael (1 June 2008). "Health Disparities and Environmental Competence: A Case Study of Appalachian Coal Mining" (in en). Environmental Justice 1 (2): 81–86. doi:10.1089/env.2008.0511. ISSN 1939-4071. http://www.liebertpub.com/doi/10.1089/env.2008.0511.
- ↑ McCulloch, B.Jan (1 March 1995). "The relationship of family proximity and social support to the mental health of older rural adults: The Appalachian context" (in en). Journal of Aging Studies 9 (1): 65–81. doi:10.1016/0890-4065(95)90026-8. https://linkinghub.elsevier.com/retrieve/pii/0890406595900268.
- ↑ Simpson, Mary Rado; King, Marilyn Givens; Mary Rado Simpson is a Doctoral Candidate at the College of Nursing and Marilyn Givens King is an Associate Professor at the College of Nursing and the Center for Rural Health at the University of Kentucky, Lexington, Kentucky. (1 February 1999). "“God Brought All These Churches Together”: Issues in Developing Religion‐Health Partnerships in an Appalachian Community" (in en). Public Health Nursing 16 (1): 41–49. doi:10.1046/j.1525-1446.1999.00041.x. ISSN 0737-1209. https://onlinelibrary.wiley.com/doi/abs/10.1046/j.1525-1446.1999.00041.x.
- ↑ Holve, Erin; Segal, Courtney; Hamilton Lopez, Marianne (1 July 2012). "Opportunities and Challenges for Comparative Effectiveness Research (CER) With Electronic Clinical Data: A Perspective From the EDM Forum" (in en). Medical Care 50: S11–S18. doi:10.1097/MLR.0b013e318258530f. ISSN 0025-7079. https://journals.lww.com/00005650-201207001-00006.
- ↑ Rabinowitz, Howard K.; Paynter, Nina P. (2 January 2002). "MSJAMA. The rural vs urban practice decision". JAMA 287 (1): 113. ISSN 0098-7484. PMID 11754723. https://pubmed.ncbi.nlm.nih.gov/11754723.
- ↑ Anderson, Allison E.; Henry, Kevin A.; Samadder, N. Jewel; Merrill, Ray M.; Kinney, Anita Y. (1 May 2013). "Rural vs Urban Residence Affects Risk-Appropriate Colorectal Cancer Screening" (in en). Clinical Gastroenterology and Hepatology 11 (5): 526–533. doi:10.1016/j.cgh.2012.11.025. PMC PMC3615111. PMID 23220166. https://linkinghub.elsevier.com/retrieve/pii/S1542356512014255.
- ↑ Reif, Susan; Whetten, Kathryn; Ostermann, Jan; Raper, James L. (1 September 2006). "Characteristics of HIV-infected adults in the Deep South and their utilization of mental health services: A rural vs. urban comparison" (in en). AIDS Care 18 (sup1): 10–17. doi:10.1080/09540120600838738. ISSN 0954-0121. https://www.tandfonline.com/doi/full/10.1080/09540120600838738.
- ↑ Shubhakaran, K.P.; Khichar, R.J. (27 March 2017). "Stroke management disparity in urban vs rural locations". Neurology. https://n.neurology.org/content/stroke-management-disparity-urban-vs-rural-locations. Retrieved 25 September 2020.
- ↑ Newgard, Craig D.; Fu, Rongwei; Bulger, Eileen; Hedges, Jerris R.; Mann, N. Clay; Wright, Dagan A.; Lehrfeld, David P.; Shields, Carol et al. (1 January 2017). "Evaluation of Rural vs Urban Trauma Patients Served by 9-1-1 Emergency Medical Services" (in en). JAMA Surgery 152 (1): 11–18. doi:10.1001/jamasurg.2016.3329. ISSN 2168-6254. PMC PMC5409522. PMID 27732713. http://archsurg.jamanetwork.com/article.aspx?doi=10.1001/jamasurg.2016.3329.
- ↑ Chen, Min; Mao, Shiwen; Liu, Yunhao (1 April 2014). "Big Data: A Survey" (in en). Mobile Networks and Applications 19 (2): 171–209. doi:10.1007/s11036-013-0489-0. ISSN 1383-469X. http://link.springer.com/10.1007/s11036-013-0489-0.
- ↑ Chen, Min; Hao, Yixue; Hwang, Kai; Wang, Lu; Wang, Lin (2017). "Disease Prediction by Machine Learning Over Big Data From Healthcare Communities". IEEE Access 5: 8869–8879. doi:10.1109/ACCESS.2017.2694446. ISSN 2169-3536. http://ieeexplore.ieee.org/document/7912315/.
- ↑ Jensen, Peter B.; Jensen, Lars J.; Brunak, Søren (1 June 2012). "Mining electronic health records: towards better research applications and clinical care" (in en). Nature Reviews Genetics 13 (6): 395–405. doi:10.1038/nrg3208. ISSN 1471-0056. http://www.nature.com/articles/nrg3208.
- ↑ Wang, Yichuan; Kung, LeeAnn; Byrd, Terry Anthony (1 January 2018). "Big data analytics: Understanding its capabilities and potential benefits for healthcare organizations" (in en). Technological Forecasting and Social Change 126: 3–13. doi:10.1016/j.techfore.2015.12.019. https://linkinghub.elsevier.com/retrieve/pii/S0040162516000500.
- ↑ Bhardwaj, Niharika; Wodajo, Bezawit; Spano, Anthony; Neal, Symaron; Coustasse, Alberto (1 January 2018). "The Impact of Big Data on Chronic Disease Management" (in en). The Health Care Manager 37 (1): 90–98. doi:10.1097/HCM.0000000000000194. ISSN 1550-512X. https://journals.lww.com/00126450-201801000-00014.
- ↑ Elam, C. (2012). "Culture, poverty and education in Appalachian Kentucky". Education & Culture 18 (1): 4. https://docs.lib.purdue.edu/eandc/vol18/iss1/art4/.
- ↑ Coyne, Cathy A; Demian-Popescu, Cristina; Friend, Dana (15 September 2006). "Social and Cultural Factors Influencing Health in Southern West Virginia: A Qualitative Study". Preventing Chronic Disease 3 (4): A124. ISSN 1545-1151. PMC 1779288. PMID 16978499. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1779288/.
- ↑ Behringer, Bruce; Friedell, Gilbert H (15 September 2006). "Appalachia: Where Place Matters in Health". Preventing Chronic Disease 3 (4): A113. ISSN 1545-1151. PMC 1779277. PMID 16978488. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1779277/.
- ↑ 25.0 25.1 Kim, Jungyeon; Ohsfeldt, Robert L.; Gamm, Larry D.; Radcliff, Tiffany A.; Jiang, Luohua (1 June 2017). "Hospital Characteristics are Associated With Readiness to Attain Stage 2 Meaningful Use of Electronic Health Records: Readiness for Stage 2 Meaningful Use" (in en). The Journal of Rural Health 33 (3): 275–283. doi:10.1111/jrh.12193. https://onlinelibrary.wiley.com/doi/10.1111/jrh.12193.
- ↑ Mason, Patricia; Mayer, Roger; Chien, Wen-Wen; Monestime, Judith (12 November 2017). "Overcoming Barriers to Implementing Electronic Health Records in Rural Primary Care Clinics" (in en). The Qualitative Report 22 (11): 2943–2955. doi:10.46743/2160-3715/2017.2515. ISSN 2160-3715. https://nsuworks.nova.edu/tqr/vol22/iss11/7/.
- ↑ Woolf, Steven H. (9 January 2008). "The Meaning of Translational Research and Why It Matters" (in en). JAMA 299 (2): 211–13. doi:10.1001/jama.2007.26. ISSN 0098-7484. http://jama.jamanetwork.com/article.aspx?doi=10.1001/jama.2007.26.
- ↑ 28.0 28.1 For members of the Jerusalem Trauma Outreach and Prevention Study (J-TOPS) group; Karstoft, Karen-Inge; Galatzer-Levy, Isaac R; Statnikov, Alexander; Li, Zhiguo; Shalev, Arieh Y (1 December 2015). "Bridging a translational gap: using machine learning to improve the prediction of PTSD" (in en). BMC Psychiatry 15 (1): 30. doi:10.1186/s12888-015-0399-8. ISSN 1471-244X. PMC PMC4360940. PMID 25886446. http://bmcpsychiatry.biomedcentral.com/articles/10.1186/s12888-015-0399-8.
- ↑ Ethier, J. -F.; Curcin, V.; Barton, A.; McGilchrist, M. M.; Bastiaens, H.; Andreasson, A.; Rossiter, J.; Zhao, L. et al. (2015). "Clinical Data Integration Model: Core Interoperability Ontology for Research Using Primary Care Data" (in en). Methods of Information in Medicine 54 (01): 16–23. doi:10.3414/ME13-02-0024. ISSN 0026-1270. http://www.thieme-connect.de/DOI/DOI?10.3414/ME13-02-0024.
- ↑ Bates, David W.; Saria, Suchi; Ohno-Machado, Lucila; Shah, Anand; Escobar, Gabriel (1 July 2014). "Big Data In Health Care: Using Analytics To Identify And Manage High-Risk And High-Cost Patients" (in en). Health Affairs 33 (7): 1123–1131. doi:10.1377/hlthaff.2014.0041. ISSN 0278-2715. http://www.healthaffairs.org/doi/10.1377/hlthaff.2014.0041.
- ↑ Handelsman, D. (2012). "Applying Business Analytics to Optimize Clinical Research Operations". SAS Global Forum 2012. https://support.sas.com/resources/papers/proceedings12/171-2012.pdf. Retrieved 28 September 2020.
- ↑ Simpao, Allan F.; Ahumada, Luis M.; Gálvez, Jorge A.; Rehman, Mohamed A. (1 April 2014). "A Review of Analytics and Clinical Informatics in Health Care" (in en). Journal of Medical Systems 38 (4): 45. doi:10.1007/s10916-014-0045-x. ISSN 0148-5598. http://link.springer.com/10.1007/s10916-014-0045-x.
- ↑ Iwabuchi, Sarina J.; Liddle, Peter F.; Palaniyappan, Lena (2013). "Clinical Utility of Machine-Learning Approaches in Schizophrenia: Improving Diagnostic Confidence for Translational Neuroimaging". Frontiers in Psychiatry 4: 95. doi:10.3389/fpsyt.2013.00095. ISSN 1664-0640. PMC PMC3756305. PMID 24009589. http://journal.frontiersin.org/article/10.3389/fpsyt.2013.00095/abstract.
- ↑ Ainali, C. (28 September 2012). "Machine Learning for Translational Medicine" (PDF). King's College London. https://kclpure.kcl.ac.uk/portal/files/31802684/2013_Ainali_Chrysanthi_0829730_ethesis.pdf. Retrieved 28 September 2020.
- ↑ Jiang, Min; Chen, Yukun; Liu, Mei; Rosenbloom, S Trent; Mani, Subramani; Denny, Joshua C; Xu, Hua (1 September 2011). "A study of machine-learning-based approaches to extract clinical entities and their assertions from discharge summaries" (in en). Journal of the American Medical Informatics Association 18 (5): 601–606. doi:10.1136/amiajnl-2011-000163. ISSN 1067-5027. PMC PMC3168315. PMID 21508414. https://academic.oup.com/jamia/article-lookup/doi/10.1136/amiajnl-2011-000163.
- ↑ Sittig, Dean F.; Hazlehurst, Brian L.; Brown, Jeffrey; Murphy, Shawn; Rosenman, Marc; Tarczy-Hornoch, Peter; Wilcox, Adam B. (1 July 2012). "A Survey of Informatics Platforms That Enable Distributed Comparative Effectiveness Research Using Multi-institutional Heterogenous Clinical Data" (in en). Medical Care 50: S49–S59. doi:10.1097/MLR.0b013e318259c02b. ISSN 0025-7079. PMC PMC3415281. PMID 22692259. https://journals.lww.com/00005650-201207001-00012.
- ↑ 37.0 37.1 Cecchetti, Alfred A.; Parmanto, Bambang; Vecchio, Marcella L.; Ahmad, Sjarif; Buch, Shama; Zgheib, Nathalie K; Groark Jr., Stephen J.; Vemuganti, Anupama et al. (1 December 2009). "Team Building: Electronic Management-Clinical Translational Research (eM-CTR) Systems" (in en). Clinical and Translational Science 2 (6): 449–455. doi:10.1111/j.1752-8062.2009.00157.x. PMC PMC3687802. PMID 20443940. https://onlinelibrary.wiley.com/doi/10.1111/j.1752-8062.2009.00157.x.
- ↑ Weber, S. C.; Lowe, H.; Das, A.; Ferris, T. (1 June 2012). "A simple heuristic for blindfolded record linkage" (in en). Journal of the American Medical Informatics Association 19 (e1): e157–e161. doi:10.1136/amiajnl-2011-000329. ISSN 1067-5027. PMC PMC3392854. PMID 22298567. https://academic.oup.com/jamia/article-lookup/doi/10.1136/amiajnl-2011-000329.
- ↑ Palma, G.; (17 September 2015). "Electronic Health Records: The Good, the Bad and the Ugly". Becker's Health IT. https://www.beckershospitalreview.com/healthcare-information-technology/electronic-health-records-the-good-the-bad-and-the-ugly.html. Retrieved 25 September 2020.
- ↑ Kaufman, Kenneth R.; Hyler, Steven E. (1 May 2005). "Problems with the Electronic Medical Record in Clinical Psychiatry: A Hidden Cost:" (in en). Journal of Psychiatric Practice 11 (3): 200–204. doi:10.1097/00131746-200505000-00008. ISSN 1538-1145. http://journals.lww.com/00131746-200505000-00008.
- ↑ Shin, Eric Y.; Ochuko, Patricia; Bhatt, Kunal; Howard, Brian; McGorisk, Gerard; Delaney, Linda; Langdon, Kristan; Khosravanipour, Marjan et al. (17 April 2018). "Errors in Electronic Health Record–Based Data Query of Statin Prescriptions in Patients With Coronary Artery Disease in a Large, Academic, Multispecialty Clinic Practice" (in en). Journal of the American Heart Association 7 (8): e007762. doi:10.1161/JAHA.117.007762. ISSN 2047-9980. PMC PMC6015411. PMID 29650707. https://www.ahajournals.org/doi/10.1161/JAHA.117.007762.
- ↑ Goodloe, Robert; Farber-Eger, Eric; Boston, Jonathan; Crawford, Dana C.; Bush, William S. (26 July 2017). "Reducing Clinical Noise for Body Mass Index Measures Due to Unit and Transcription Errors in the Electronic Health Record". AMIA Summits on Translational Science Proceedings 2017: 102–111. ISSN 2153-4063. PMC 5543370. PMID 28815116. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5543370/.
- ↑ Harris, Paul A.; Taylor, Robert; Thielke, Robert; Payne, Jonathon; Gonzalez, Nathaniel; Conde, Jose G. (1 April 2009). "Research electronic data capture (REDCap)—A metadata-driven methodology and workflow process for providing translational research informatics support" (in en). Journal of Biomedical Informatics 42 (2): 377–381. doi:10.1016/j.jbi.2008.08.010. PMC PMC2700030. PMID 18929686. https://linkinghub.elsevier.com/retrieve/pii/S1532046408001226.
- ↑ Harris, Paul A.; Taylor, Robert; Minor, Brenda L.; Elliott, Veida; Fernandez, Michelle; O'Neal, Lindsay; McLeod, Laura; Delacqua, Giovanni et al. (1 July 2019). "The REDCap consortium: Building an international community of software platform partners" (in en). Journal of Biomedical Informatics 95: 103208. doi:10.1016/j.jbi.2019.103208. PMC PMC7254481. PMID 31078660. https://linkinghub.elsevier.com/retrieve/pii/S1532046419301261.
- ↑ Harris, Paul A (1 July 2012). "Research Electronic Data Capture (REDCap) - planning, collecting and managing data for clinical and translational research" (in en). BMC Bioinformatics 13 (S12): A15, 1471–2105–13-S12-A15. doi:10.1186/1471-2105-13-S12-A15. ISSN 1471-2105. PMC PMC3409058. https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-13-S12-A15.
- ↑ 46.0 46.1 Marshall University; Cecchetti, Alfred A. (1 April 2018). "Why Introduce Machine Learning To Rural Health Care?". Marshall Journal of Medicine 4 (2). doi:10.18590/mjm.2018.vol4.iss2.2. http://mds.marshall.edu/mjm/vol4/iss2/2/.
- ↑ McDonald, Helen I.; Shaw, Catriona; Thomas, Sara L.; Mansfield, Kathryn E.; Tomlinson, Laurie A.; Nitsch, Dorothea (1 November 2016). "Methodological challenges when carrying out research on CKD and AKI using routine electronic health records" (in en). Kidney International 90 (5): 943–949. doi:10.1016/j.kint.2016.04.010. https://linkinghub.elsevier.com/retrieve/pii/S0085253816301661.
- ↑ Pierce, Carolyn; Scherra, Elizabeth (1 December 2004). "The Challenges of Data Collection in Rural Dwelling Samples". Online Journal of Rural Nursing and Health Care 4 (2): 25–30. doi:10.14574/ojrnhc.v4i2.197. https://rnojournal.binghamton.edu/index.php/RNO/article/view/197.
- ↑ Verby, J E (1 November 1989). "Patientsʼ and physiciansʼ views of health in a rural area:" (in en). Academic Medicine 64 (11): 665–6. doi:10.1097/00001888-198911000-00009. ISSN 1040-2446. http://journals.lww.com/00001888-198911000-00009.
- ↑ Hendryx, Michael; Ahern, Melissa M.; Nurkiewicz, Timothy R. (13 November 2007). "Hospitalization Patterns Associated with Appalachian Coal Mining" (in en). Journal of Toxicology and Environmental Health, Part A 70 (24): 2064–2070. doi:10.1080/15287390701601236. ISSN 1528-7394. http://www.tandfonline.com/doi/abs/10.1080/15287390701601236.
- ↑ Ortmeyer, Carl E.; Costello, Joseph; Morgan, William Keith C.; Sweeker, Steve; Peterson, Martin (1 August 1974). "The Mortality of Appalachian Coal Miners, 1963 to 1971" (in en). Archives of Environmental Health: An International Journal 29 (2): 67–72. doi:10.1080/00039896.1974.10666535. ISSN 0003-9896. http://www.tandfonline.com/doi/abs/10.1080/00039896.1974.10666535.
- ↑ 52.0 52.1 52.2 Amro, Ahmed; Baez, Giancarlo Acosta; Koromia, George Augustine; Bhardwaj, Niharika; Aguilar, Rodrigo; El-Hamdani, Mehiar; Irfan, Affan (1 March 2019). "ALBUMIN LEVEL AS A RISK MARKER AND PREDICTOR OF PERIPARTUM CARDIOMYOPATHY" (in en). Journal of the American College of Cardiology 73 (9): 835. doi:10.1016/S0735-1097(19)31442-1. https://linkinghub.elsevier.com/retrieve/pii/S0735109719314421.
- ↑ 53.0 53.1 53.2 Acosta, Giancarlo; Amro, Ahmed; Aguilar, Rodrigo; Abusnina, Waiel; Bhardwaj, Niharika; Koromia, George Augustine; Studeny, Mark; Irfan, Affan (27 January 2020). "Clinical Determinants of Myocardial Injury, Detectable and Serial Troponin Levels among Patients with Hypertensive Crisis" (in en). Cureus 12 (1): e6787. doi:10.7759/cureus.6787. ISSN 2168-8184. PMC PMC7045977. PMID 32140347. https://www.cureus.com/articles/26546-clinical-determinants-of-myocardial-injury-detectable-and-serial-troponin-levels-among-patients-with-hypertensive-crisis.
- ↑ 54.0 54.1 54.2 54.3 Elmore, Dominique; Yaslam, Balfaqih; Putty, Krista; Magrane, Thomas; Abadir, Anthony; Bhatt, Saloni; Frazier, Marie; Flesher, Susan (1 January 2019). "Is Fever a Red Flag for Bacterial Pneumonia in Children With Viral Bronchiolitis?" (in en). Global Pediatric Health 6: 2333794X1986866. doi:10.1177/2333794X19868660. ISSN 2333-794X. PMC PMC6686317. PMID 31431903. http://journals.sagepub.com/doi/10.1177/2333794X19868660.
- ↑ 55.0 55.1 55.2 Bhardwaj, Niharika; Sundaram, Shanmuga; Carter, Larry E.; Cecchetti, Alfred A.; Sundaram, Uma (1 May 2020). "Tu1801 METABOLIC SYNDROME: ARE CURRENT COLON CANCER SCREENING GUIDELINES ENOUGH IN A RURAL POPULATION?" (in en). Gastroenterology 158 (6): S–1167. doi:10.1016/S0016-5085(20)33589-7. https://linkinghub.elsevier.com/retrieve/pii/S0016508520335897.
- ↑ 56.0 56.1 56.2 56.3 Sundaram, S.; Bhardwaj, N.; Schafer, C. et al. (30 March 2017). "Utilization of Appalachian Clinical and Translational Science Institute Data Warehouse to more accurately predict disease processes important for central Appalachia". 12th Annual CCTS Spring Conference. https://www.ccts.uky.edu/media/913.
- ↑ 57.0 57.1 Bhardwaj, Niharika; Cecchetti, Alfred A; Murughiyan, Usha; Neitch, Shirley (4 August 2020). "Analysis of Benzodiazepine Prescription Practices in Elderly Appalachians with Dementia via the Appalachian Informatics Platform: Longitudinal Study" (in en). JMIR Medical Informatics 8 (8): e18389. doi:10.2196/18389. ISSN 2291-9694. PMC PMC7435704. PMID 32749226. https://medinform.jmir.org/2020/8/e18389.
- ↑ 58.0 58.1 58.2 Khanna, R.; Gress, T.; Cecchetti, A.. "Hospital Emergency Department Visits For Non-Traumatic Oral Health Conditions". 2018 National Oral Health Conference. https://www.nationaloralhealthconference.com/pdfs/2018-poster-abstracts.pdf. Retrieved 29 September 2020.
- ↑ 59.0 59.1 59.2 Ferdjallah, M.; Driscoll, H. (6 March 2020). "Serum Calcium Homeostasis and Volume Dynamics in Alzheimer’s Disease and Diabetes Mellitus-2" (PDF). 32nd Annual Research Day at Marshall University. Marshall University. https://jcesom.marshall.edu/media/58548/9110_researchsyllabus_2020.pdf. Retrieved 29 September 2020.
- ↑ Centers for Disease Control and Prevention (9 April 2018). "Stats of the State of West Virginia". National Center for Health Statistics. https://www.cdc.gov/nchs/pressroom/states/westvirginia/westvirginia.htm. Retrieved 25 September 2020.
- ↑ Mueller, B. (24 June 2016). "Predicting_heart_disease_UCI/heartdisease_UCI.Rmd". GitHub. https://github.com/mbbrigitte/Predicting_heart_disease_UCI/blob/master/heartdisease_UCI.Rmd. Retrieved 25 September 2020.
- ↑ Sreejith, S.; Rahul, S.; Jisha, R. C. (2016), Thampi, Sabu M.; Bandyopadhyay, Sanghamitra; Krishnan, Sri et al.., eds., "A Real Time Patient Monitoring System for Heart Disease Prediction Using Random Forest Algorithm", Advances in Signal Processing and Intelligent Recognition Systems (Cham: Springer International Publishing) 425: 485–500, doi:10.1007/978-3-319-28658-7_41, ISBN 978-3-319-28656-3, http://link.springer.com/10.1007/978-3-319-28658-7_41. Retrieved 2022-06-07
- ↑ Tanaka, Tomohiro; Voigt, Michael D. (1 March 2018). "Decision tree analysis to stratify risk of de novo non-melanoma skin cancer following liver transplantation" (in en). Journal of Cancer Research and Clinical Oncology 144 (3): 607–615. doi:10.1007/s00432-018-2589-5. ISSN 0171-5216. http://link.springer.com/10.1007/s00432-018-2589-5.
- ↑ Paxton, Raheem J.; Zhang, Lingfeng; Wei, Changshuai; Price, Daniel; Zhang, Fan; Courneya, Kerry S.; Kakadiaris, Ioannis A. (1 October 2019). "An exploratory decision tree analysis to predict physical activity compliance rates in breast cancer survivors.". Ethnicity & health 24 (7): 754–766. doi:10.1080/13557858.2017.1378805. ISSN 1355-7858. PMC 6581628. PMID 28922931. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6581628/.
- ↑ Moreno Hoyo, A. (January 2017). "Predicting car accidents in Barcelona using a Random Forest model". Universitat Politècnica de Catalunya. http://hdl.handle.net/2117/100298. Retrieved 25 September 2020.
- ↑ Xiaohui, J. (2016). "Forecast model of road traffic accidents based on LS-SVM with grey correlation analysis". Applied Research in Computing. China National Knowledge Infrastructure. http://en.cnki.com.cn/Article_en/CJFDTotal-JSYJ201603038.htm.
- ↑ Khatri, Krishan L.; Tamil, Lakshman S. (1 January 2018). "Early Detection of Peak Demand Days of Chronic Respiratory Diseases Emergency Department Visits Using Artificial Neural Networks". IEEE Journal of Biomedical and Health Informatics 22 (1): 285–290. doi:10.1109/JBHI.2017.2698418. ISSN 2168-2194. http://ieeexplore.ieee.org/document/7912386/.
- ↑ Oijuela-Canon, Alvaro D.; Gomez-Cajas, Diego F.; Sepulveda-Sepulveda, Alexander (1 September 2015). "Respiratory Diseases discrimination based on acoustic lung signals and neural networks". 2015 20th Symposium on Signal Processing, Images and Computer Vision (STSIVA) (Bogota, Colombia: IEEE): 1–6. doi:10.1109/STSIVA.2015.7330461. ISBN 978-1-4673-9461-1. https://ieeexplore.ieee.org/document/7330461.
- ↑ McKinley, Richard; Häni, Levin; Gralla, Jan; El-Koussy, M; Bauer, S; Arnold, M; Fischer, U; Jung, S et al. (1 August 2017). "Fully automated stroke tissue estimation using random forest classifiers (FASTER)" (in en). Journal of Cerebral Blood Flow & Metabolism 37 (8): 2728–2741. doi:10.1177/0271678X16674221. ISSN 0271-678X. PMC PMC5536784. PMID 27798267. http://journals.sagepub.com/doi/10.1177/0271678X16674221.
- ↑ Chen, L.; Bentley, P.; Rueckert, D. (2015). "A novel framework for sub-acute stroke lesion segmentation based on random forest". Proceedings of Ischemic Stroke Lesion Segmentation. pp. 9–12. http://www.isles-challenge.org/ISLES2015/pdf/20150930_ISLES2015_Proceedings.pdf#page=17.
- ↑ Xu, Weifeng; Zhang, Jianxin; Zhang, Qiang; Wei, Xiaopeng (1 February 2017). "Risk prediction of type II diabetes based on random forest model". 2017 Third International Conference on Advances in Electrical, Electronics, Information, Communication and Bio-Informatics (AEEICB) (Chennai, India: IEEE): 382–386. doi:10.1109/AEEICB.2017.7972337. ISBN 978-1-5090-5434-3. http://ieeexplore.ieee.org/document/7972337/.
- ↑ Shukla, N.; Arora, M. (2016). "Prediction of Diabetes Using Neural Network & Random Forest Tree". International Journal of Computer Sciences and Engineering 4 (7): 101–04. https://www.ijcseonline.org/full_paper_view.php?paper_id=1008.
- ↑ Basaia, Silvia; Agosta, Federica; Wagner, Luca; Canu, Elisa; Magnani, Giuseppe; Santangelo, Roberto; Filippi, Massimo (2019). "Automated classification of Alzheimer's disease and mild cognitive impairment using a single MRI and deep neural networks" (in en). NeuroImage: Clinical 21: 101645. doi:10.1016/j.nicl.2018.101645. PMC PMC6413333. PMID 30584016. https://linkinghub.elsevier.com/retrieve/pii/S2213158218303930.
- ↑ Alzheimer’s Disease Neuroimaging Initiative; Lu, Donghuan; Popuri, Karteek; Ding, Gavin Weiguang; Balachandar, Rakesh; Beg, Mirza Faisal (1 December 2018). "Multimodal and Multiscale Deep Neural Networks for the Early Diagnosis of Alzheimer’s Disease using structural MR and FDG-PET images" (in en). Scientific Reports 8 (1): 5697. doi:10.1038/s41598-018-22871-z. ISSN 2045-2322. PMC PMC5890270. PMID 29632364. http://www.nature.com/articles/s41598-018-22871-z.
- ↑ Wiemken, Timothy L.; Furmanek, Stephen P.; Mattingly, William A.; Guinn, Brian E.; Cavallazzi, Rodrigo; Fernandez-Botran, Rafael; Wolf, Leslie A.; English, Connor L. et al. (2017). "Predicting 30-day mortality in hospitalized patients with community-acquired pneumonia using statistical and machine learning approaches.". Journal of Respiratory Infections 1 (3). doi:10.18297/jri/vol1/iss3/10/. http://ir.library.louisville.edu/jri/vol1/iss3/10/.
- ↑ Menéndez Villanueva, R. (1 August 1995). "[The diagnostic evaluation of rapid sputum technics for Pneumococcus in community-acquired pneumonia. The usefulness of Bayes theorem for clinical application"]. Archivos De Bronconeumologia 31 (7): 317–322. ISSN 0300-2896. PMID 8777525. https://pubmed.ncbi.nlm.nih.gov/8777525.
- ↑ B.V, Ravindra; Sriraam, N; Geetha, M (31 December 2017). "Classification of non-chronic and chronic kidney disease using SVM neural networks". International Journal of Engineering & Technology 7 (1.3): 191–94. doi:10.14419/ijet.v7i1.3.10669. ISSN 2227-524X. https://www.sciencepubco.com/index.php/ijet/article/view/10669.
- ↑ Annapoorani, J.C.B.; Gnanaselvam, C.R. (2018). "Enhancing Prediction Accuracy of Chronic Kidney Disease using Neural Networks". Automation and Autonomous System 10 (1): 10–15. doi:10.36039/AA012018003. http://www.ciitresearch.org/dl/index.php/aa/article/view/AA012018003..
- ↑ Ayat, Saeed; Farahani, Hojjat A.; Aghamohamadi, Mehdi; Alian, Mahmood; Aghamohamadi, Somayeh; Kazemi, Zeynab (1 October 2013). "A comparison of artificial neural networks learning algorithms in predicting tendency for suicide" (in en). Neural Computing and Applications 23 (5): 1381–1386. doi:10.1007/s00521-012-1086-z. ISSN 0941-0643. http://link.springer.com/10.1007/s00521-012-1086-z.
- ↑ Bhat, H.S.; Goldman-Mellor, Sidra J. (2017). "Predicting Adolescent Suicide Attempts with Neural Networks". arXiv. arXiv:1711.10057v2. https://arxiv.org/abs/1711.10057v2.
- ↑ Wolfram, W.; Christian, D. (1976). "Appalachian Speech" (PDF). Center for Applied Linguistics. https://files.eric.ed.gov/fulltext/ED130511.pdf. Retrieved 25 September 2020.
- ↑ Luhman, Reid (1 September 1990). "Appalachian English stereotypes: Language attitudes in Kentucky" (in en). Language in Society 19 (3): 331–348. doi:10.1017/S0047404500014548. ISSN 0047-4045. https://www.cambridge.org/core/product/identifier/S0047404500014548/type/journal_article.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation, grammar, and punctuation. In some cases important information was missing from the references, and that information was added.