Difference between revisions of "Journal:Laboratory information management system for COVID-19 non-clinical efficacy trial data"
Shawndouglas (talk | contribs) (Saving and adding more.) |
Shawndouglas (talk | contribs) (Saving and adding more.) |
||
| Line 37: | Line 37: | ||
The novel [[severe acute respiratory syndrome coronavirus 2]] (SARS-CoV-2) has resulted in the serious respiratory-related [[coronavirus disease 2019]] (COVID-19) [[pandemic]]. It was first reported in Wuhan, the capital of Hubei, China, in late December 2019 and has attracted international attention. [1] After four months, it spread to multiple countries, leading to more than one million confirmed cases worldwide. [2] SARS-CoV-2, a single-stranded RNA-enveloped virus [3], enters the host cells by binding its structural spike (S) protein to cell surface receptors, angiotensin-converting enzyme 2 (ACE2.) [4] After entering the host cells, the virus hijacks the cell to undergo viral replication. [5] During this process, an aberrant host immune condition called a cytokine storm can occur in response to SARS-CoV-2 infection, which is characterized by high concentrations of pro-inflammatory cytokines and chemokines, including tumor necrosis factor-α, interleukin-1, and interleukin-6 [6], resulting in severe COVID-19 associated with excessive inflammation. [7] | The novel [[severe acute respiratory syndrome coronavirus 2]] (SARS-CoV-2) has resulted in the serious respiratory-related [[coronavirus disease 2019]] (COVID-19) [[pandemic]]. It was first reported in Wuhan, the capital of Hubei, China, in late December 2019 and has attracted international attention. [1] After four months, it spread to multiple countries, leading to more than one million confirmed cases worldwide. [2] SARS-CoV-2, a single-stranded RNA-enveloped virus [3], enters the host cells by binding its structural spike (S) protein to cell surface receptors, angiotensin-converting enzyme 2 (ACE2.) [4] After entering the host cells, the virus hijacks the cell to undergo viral replication. [5] During this process, an aberrant host immune condition called a cytokine storm can occur in response to SARS-CoV-2 infection, which is characterized by high concentrations of pro-inflammatory cytokines and chemokines, including tumor necrosis factor-α, interleukin-1, and interleukin-6 [6], resulting in severe COVID-19 associated with excessive inflammation. [7] | ||
Along with understanding the mechanism of transmissibility and pathogenesis of SARS-CoV-2, significant efforts are being devoted to developing novel vaccines and therapeutic agents to prevent or treat COVID-19. In particular, various COVID-19-related research has been conducted with the participation of multiple organizations. Among such studies, animal models of monkeys, cats, ferrets, hamsters, and human angiotensin converting enzyme 2 (hACE2) transgenic mice [8], capable of conducting controlled experiments, have made significant contributions to vaccine development. [9,10,11,12,13] (Note: Though the U.S. [[Food and Drug Administration]] doesn't consider animal studies to be part of "non-clinical laboratory studies," many research organizations with animal lab capabilities still use "non-clinical" to include and describe these sorts of animal modelling studies.<ref name="PCPreclin20">{{cite web |url=https://premierconsulting.com/resources/blog/preclinical-and-nonclinical-studies-what-is-the-difference-and-where-in-your-program-should-they-fall/ |title=Preclinical and Nonclinical Studies—What Is the Difference, and Where in Your Program Should They Fall? |publisher=Premier Consulting |date=29 July 2020 |accessdate=27 May 2022}}</ref>) Because various parameters such as mortality, clinical signs, body weight, organ weights, viral titer (viral replication and viral RNA), and multi-organ [[histopathology]] should be comprehensively analyzed for a reliable interpretation of the results, there is an increasing need for an integrated system in which multiple institutions can apply a streamlined analysis and input the results simultaneously when conducting large-scale non-clinical efficacy tests of a COVID-19 vaccine. | Along with understanding the mechanism of transmissibility and pathogenesis of SARS-CoV-2, significant efforts are being devoted to developing novel vaccines and therapeutic agents to prevent or treat COVID-19. In particular, various COVID-19-related research has been conducted with the participation of multiple organizations. Among such studies, animal models of monkeys, cats, ferrets, hamsters, and human angiotensin converting enzyme 2 (hACE2) transgenic mice [8], capable of conducting controlled experiments, have made significant contributions to vaccine development. [9,10,11,12,13] (Note: Though the U.S. [[Food and Drug Administration]] doesn't consider animal studies to be part of "non-clinical [[laboratory]] studies," many research organizations with animal lab capabilities still use "non-clinical" to include and describe these sorts of animal modelling studies.<ref name="PCPreclin20">{{cite web |url=https://premierconsulting.com/resources/blog/preclinical-and-nonclinical-studies-what-is-the-difference-and-where-in-your-program-should-they-fall/ |title=Preclinical and Nonclinical Studies—What Is the Difference, and Where in Your Program Should They Fall? |publisher=Premier Consulting |date=29 July 2020 |accessdate=27 May 2022}}</ref>) Because various parameters such as mortality, clinical signs, body weight, organ weights, viral titer (viral replication and viral RNA), and multi-organ [[histopathology]] should be comprehensively analyzed for a reliable interpretation of the results, there is an increasing need for an integrated system in which multiple institutions can apply a streamlined analysis and input the results simultaneously when conducting large-scale non-clinical efficacy tests of a COVID-19 vaccine. | ||
When experimental data are collected from multiple institutions, “organization-level effects” may occur owing to non-standardization in the data coding process and coding errors, which can act as a batch effect in a [[data analysis]]. In large-scale experimental | When experimental data are collected from multiple institutions, “organization-level effects” may occur owing to non-standardization in the data coding process and coding errors, which can act as a batch effect in a [[data analysis]]. In large-scale experimental multi-center research, it is necessary to prevent the loss of reliability in the data analysis caused by batch effects; however, this is a difficult goal to achieve without an integrated system. An effective approach to overcoming the limitations caused by the prior-mentioned batch effects is to use a [[laboratory information management system]] (LIMS). [14] A LIMS systematizes the research process by automating data-related tasks that were conducted manually using information technology, and simultaneously realizes various needs, such as the standardization of experimental data, sharing, problem tracking, and the creation of reports based on document templates. The accumulated data stored in a LIMS can be useful in understanding the current clinical knowledge and making decisions, which is a remarkable advantage in research on rapidly changing epidemics such as COVID-19. | ||
Despite the advantages of a LIMS, no systems allowing multiple institutions to participate simultaneously in research related to COVID-19 have yet been proposed among developed | Despite the advantages of a LIMS, no systems allowing multiple institutions to participate simultaneously in research related to COVID-19 have yet been proposed among developed open-source LIMSs. Given that, we aimed to clearly understand the standard procedures of non-clinical trials for COVID-19 therapeutics and vaccines and to design and implement a LIMS suitable for these research purposes. | ||
==Results== | ==Results== | ||
| Line 150: | Line 150: | ||
|} | |} | ||
Functions can be classified into four different categories: access management, trial, audit, and comparison. The access management function forms the basis of the system and manages the user group. The trial function controls the entire project process, including initiation, progression, and finalization. The [[Audit trail|audit function]] records all activities that occur in the system and implements data entry, cancelation, and restoration. The comparison function provides several plots according to the investigational product (new drug/repositioning, vaccine/therapeutic), sponsor, experimental animals (hamster model/hACE2 mouse model), research objective, and study site. It performs access control and visualization by classifying all data accumulated in the system. | Functions can be classified into four different categories: access management, trial, audit, and comparison. The access management function forms the basis of the system and manages the user group. The trial function controls the entire project process, including initiation, progression, and finalization. The [[Audit trail|audit function]] records all activities that occur in the system and implements data entry, cancelation, and restoration. The comparison function provides several plots according to the investigational product (new drug/repositioning, vaccine/therapeutic), sponsor, experimental animals (hamster model/hACE2 mouse model), research objective, and study site. It performs access control and [[Data visualization|visualization]] by classifying all data accumulated in the system. | ||
===Data security and audit=== | |||
The proposed system provides several functions for data security and audits. For security, an access management system based on the access control list (ACL) [15] was incorporated to protect data entry. Unintended data leakage was minimized by providing limited access points to users according to the user access levels. For each running instance, data security at the terminal level was obtained through secure HTTPS-based communication. For audits, the proposed system records, monitors, and manages all user activities, including data entry, by generating a separate database. Furthermore, it provides a history-tracking function to maintain data integrity. The chief director can identify the path of a data contamination and restore it to its original state, which minimizes the problem related to data entry or damage to the raw data. | |||
===Process management=== | |||
Data entry and monitoring are possible for each institution, which makes it possible for the local administrator of each institution to check the data status in real time. It is also composed of a system in which the chief director supervises the overall project progress of each organization in real time. In addition, by systematically managing various types of non-clinical data and enabling collaboration between multiple organizations, researchers are expected to be able to quickly derive the desired results. Figure 2 shows the workflow of the COVID-19 non-clinical trial LIMS for vaccine and therapeutics treatments. | |||
[[File:Fig2 Yoon LabAniRes22 38.png|700px]] | |||
{{clear}} | |||
{| | |||
| style="vertical-align:top;" | | |||
{| border="0" cellpadding="5" cellspacing="0" width="700px" | |||
|- | |||
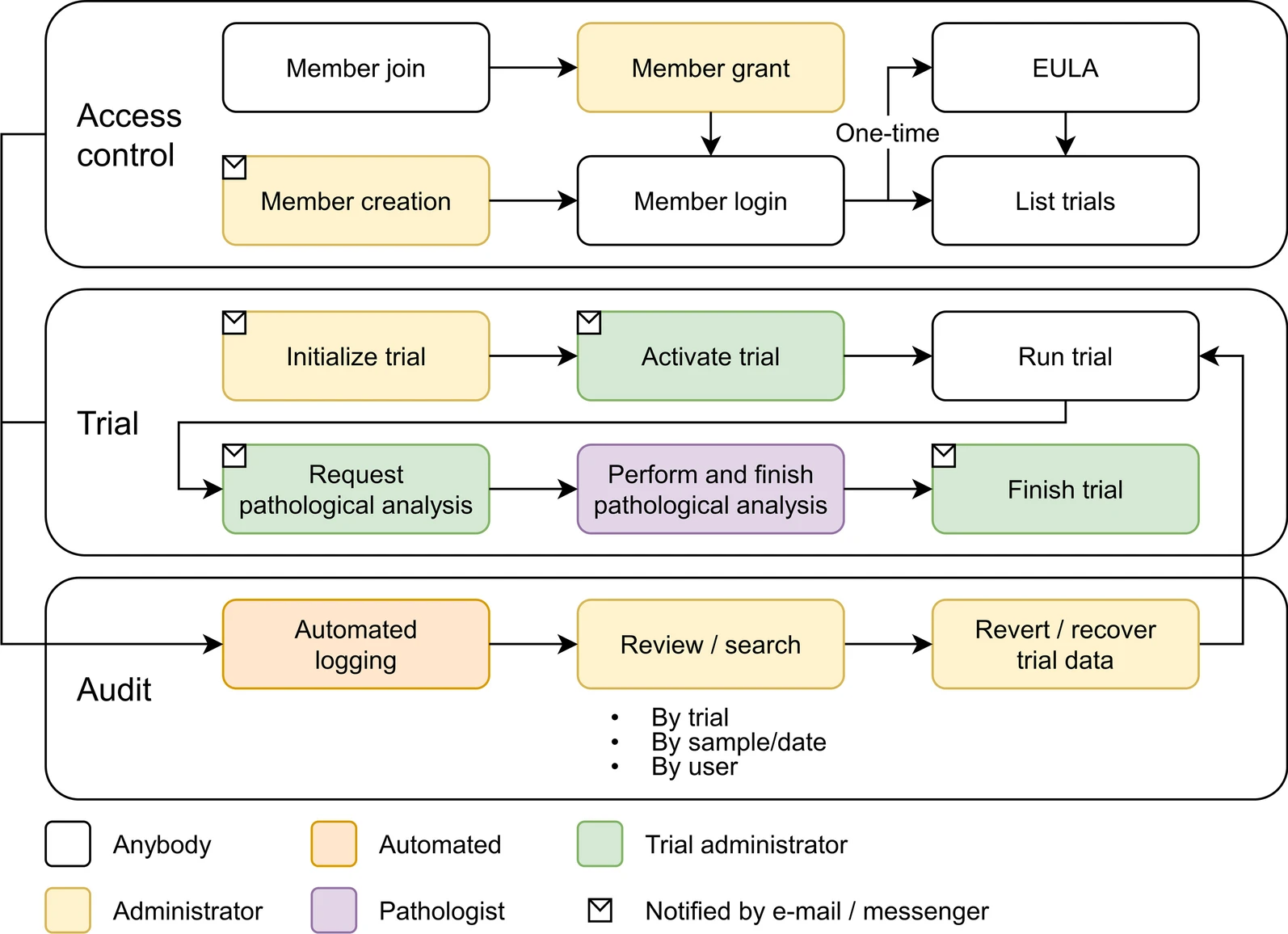
| style="background-color:white; padding-left:10px; padding-right:10px;" |<blockquote>'''Figure 2.''' Recording workflow in LIMS. The proposed LIMS audits activities and controls access throughout the study. All access to the trials is systematically consented and controlled with complete auditing. Therefore, the trial data can be tracked and managed using the audit record.</blockquote> | |||
|- | |||
|} | |||
|} | |||
===Graphics and analysis functions=== | |||
The COVID-19 non-clinical trial LIMS provides various visualizations that can review the data characteristics at each stage for each institution, as well as optimized visualizations such as line diagrams, bar diagrams, and survival curves according to the data type for awareness of current situations and early detection of unusual situations. The proposed system generates suitable plots for a correspondence analysis according to the data types. In particular, a comparison function was developed to facilitate mutual comparisons according to user-selected conditions, and the system can also generate easy-to-read study-specific graphs. | |||
===Data downloading and result reporting=== | |||
After obtaining approval from the chief director and administrators of each organization, the stored data can be downloaded as an Excel or Word file. In particular, it is possible to create an integrated Word file that includes a data description table, summary plot, and raw data, and all currently entered data can be downloaded as a multi-sheet Excel file with a single click for convenient data management. In addition, the proposed system can be used to monitor the data entry status of each participating organization and simultaneously create reports with a function for automatically summarizing the analysis results. | |||
===Notification=== | |||
In applying important commands such as research initiation and termination, a notification system through a mail and messenger (limited to South Korea) notification service was established for relevant individuals such as chief directors and administrators. When a major change occurs in the research, the manager can quickly identify and respond to any unintended situations. | |||
===Data management and recording=== | |||
Data entry control can be achieved according to the characteristics of each data type, and the user input is automatically controlled according to the condition of each stage of the study. Human error is minimized by limiting the range of unobservable values for each data parameter, and entry omission is prevented by notifying the data that need to be input according to the investigation schedule, current date, and data entry status (Fig. 3). In addition, all data entry activities were recorded, and an audit function was reviewed for each dataset. If necessary, an unsuitable data entry can be canceled to prevent data errors. On the data entry screen, users can check all data during the full research period (Fig. 4A) or partial data by filtering the data based on date (Fig. 4B). | |||
[[File:Fig3 Yoon LabAniRes22 38.png|800px]] | |||
{{clear}} | |||
{| | |||
| style="vertical-align:top;" | | |||
{| border="0" cellpadding="5" cellspacing="0" width="800px" | |||
|- | |||
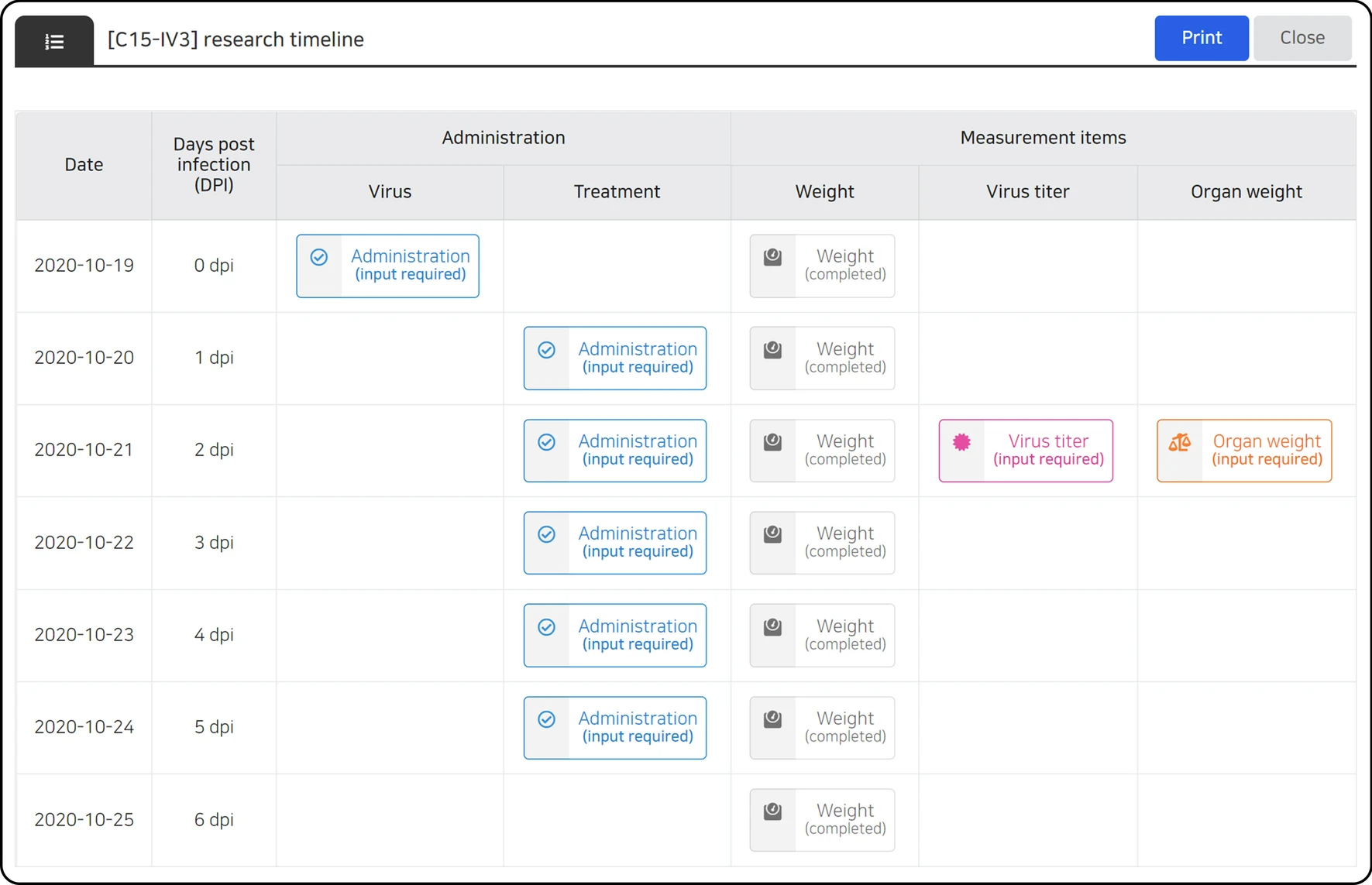
| style="background-color:white; padding-left:10px; padding-right:10px;" |<blockquote>'''Figure 3.''' LIMS calendar. The proposed LIMS provides a calendar to facilitate research under the pre-determined study plan, which allows the researchers to review the overall schedule and its corresponding data type and completion status at once.</blockquote> | |||
|- | |||
|} | |||
|} | |||
[[File:Fig4 Yoon LabAniRes22 38.png|900px]] | |||
{{clear}} | |||
{| | |||
| style="vertical-align:top;" | | |||
{| border="0" cellpadding="5" cellspacing="0" width="900px" | |||
|- | |||
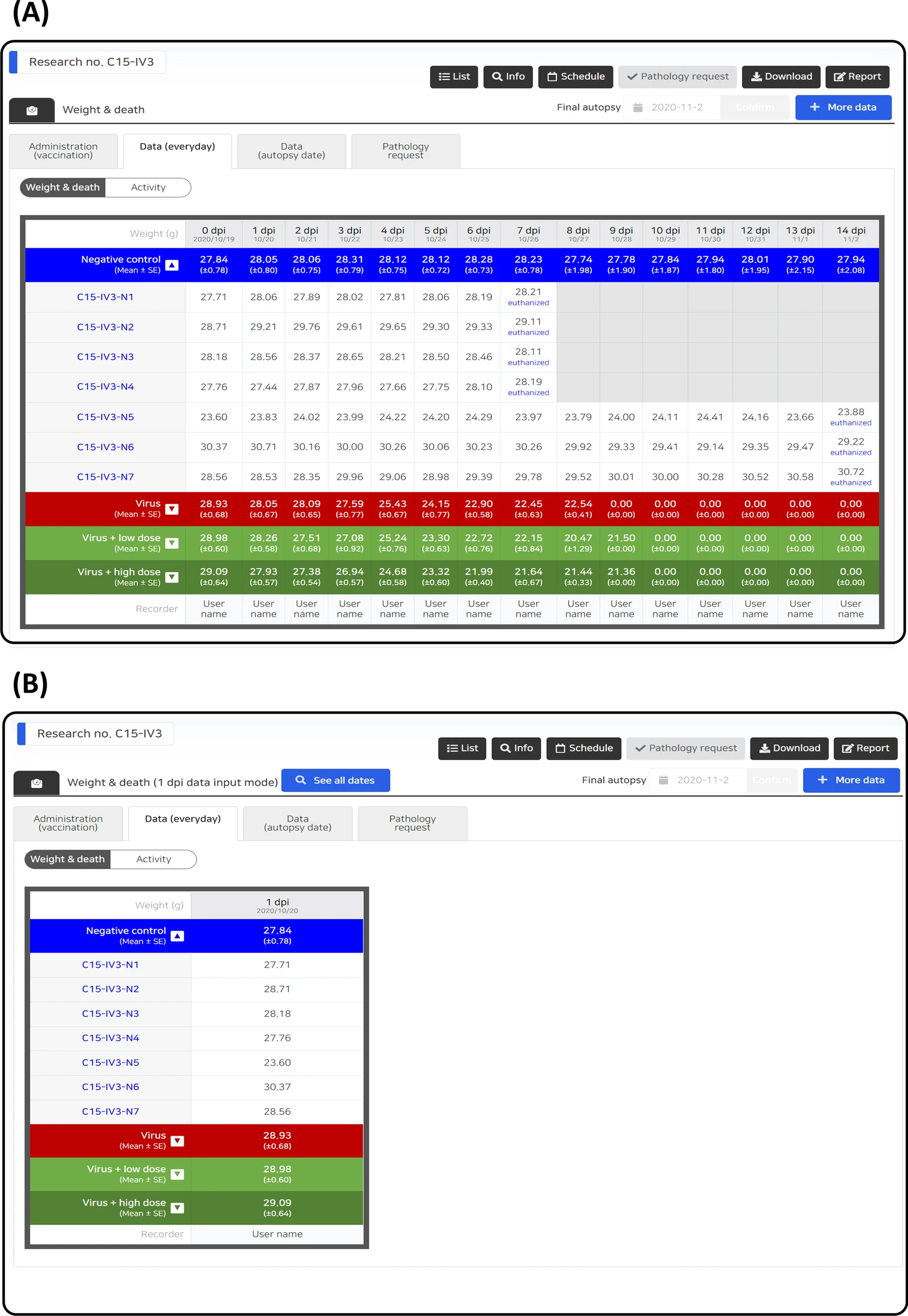
| style="background-color:white; padding-left:10px; padding-right:10px;" |<blockquote>'''Figure 4.''' LIMS sample input. '''(A)''' The data entry screen for the entire schedule. '''(B)''' The data entry screen for the selected schedule.</blockquote> | |||
|- | |||
|} | |||
|} | |||
==Discussion== | |||
Non-clinical studies are essential for identifying the potential effects of promising drug candidates prior to entering human clinical trials. [16] Numerous animal studies have been conducted in the development of therapeutic agents and vaccines during the recent COVID-19 pandemic, leading to the need for rapid and accurate non-clinical efficacy analyses in standardized animal study design protocols. [9, 10] The inoculation of hACE2 transgenic mice with SARS-CoV-2 induced weight loss, increased the viral titer in the lungs, and prompted histopathological changes with increased cytokine levels and interstitial pneumonia. [17, 18] In addition, infection with SARS-CoV-2 in a golden Syrian hamster model induced signs resembling COVID-19 disease, including decreased activity, weight loss, high lung viral load, and the presence of T-lymphocytes in the respiratory tract. [19,20,21] Therefore, systematic management of these extensive data—including mortality, clinical signs, body weight, body temperature, viral titer (viral replication and viral RNA), organ weights, and multi-organ histopathology—that can be observed in COVID-19 research using animal models such as hACE2 transgenic mice and hamsters is required for a reliable evaluation and development of a therapeutic agent or vaccine. | |||
The implemented LIMS has many distinctive features providing many user-friendly functions. First, the implemented system classifies COVID-19 non-clinical trial topics into six sub-themes (investigational product, types of investigational product, sponsor, experimental animals, research objective, and study site). It also applies access control and visualization by classifying the research accumulated in the system. Second, using a streamlined analysis, it creates a four-step process related to research progress, pathology analysis, result entry, and report generation. Access control and customized user interfaces are provided for each step, which highlights the work to be done and provides step-by-step guidance as a project planner. Third, a pathology analysis was incorporated to organize various types of pathology data and support the resulting input from pathologists. Finally, the data are organized systematically, and raw data, summary tables, and graph visualizations are documented. Many of these features were found not to be available in open-source LIMS. Table 2 provides a brief comparison of other existing open-source LIMS and the proposed COVID-19 non-clinical trial LIMS. | |||
{| | |||
| style="vertical-align:top;" | | |||
{| class="wikitable" border="1" cellpadding="5" cellspacing="0" width="70%" | |||
|- | |||
| colspan="5" style="background-color:white; padding-left:10px; padding-right:10px;" |'''Table 2.''' Comparison of open-source LIMS with the proposed COVID-19 non-clinical trial LIMS | |||
|- | |||
! style="padding-left:10px; padding-right:10px;" |Attribute | |||
! style="padding-left:10px; padding-right:10px;" |[[Bika LIMS]] | |||
! style="padding-left:10px; padding-right:10px;" |[[Stanford University School of Medicine|MendeLIMS]] | |||
! style="padding-left:10px; padding-right:10px;" |[[MetaLIMS]] | |||
! style="padding-left:10px; padding-right:10px;" |COVID-19 | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Data type | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Not specific | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Clinical samples | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Environmental samples | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Animal samples | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Web-based/stand-alone | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Web-based | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Web-based | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Web-based | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Web-based | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Implementation languages | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Python | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Javascript, Ruby | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |PHP | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |PHP, nodeJS | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Database(s) | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |ZODB | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |[[MySQL]], [[PostgreSQL]], SQLLite | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |MySQL | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |MySQL | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Website | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |https://github.com/bikalabs/bika.lims | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |https://dna-discovery.stanford.edu/research/software/mendelims/ | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |https://github.com/cheinle/MetaLIMS/wiki | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |https://github.com/rexsoft-org/covid19-lims | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |COVID-19-optimized | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |No | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |No | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |No | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Yes | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Pathology report | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Limited (customizable) | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Limited (text only) | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |No | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Yes (text + image) | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Streamlined, multi-center study (customizable) | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Limited | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |No | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |No | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Yes | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Reporting | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Certificate PDF (Not COVID-19-specific; manual process is required) | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Not supported | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |Not supported | |||
| style="background-color:white; padding-left:10px; padding-right:10px;" |COVID-19-specific, fully automated, Microsoft Word format | |||
|- | |||
|} | |||
|} | |||
The COVID-19 non-clinical trial data management system developed in this study consists of a mode for the chief director, administrators, researchers of each organization, and pathologists. It aims to improve the completeness of the database by managing the data entry process for each institution. To confirm this, we applied the proposed system to an actual COVID-19 non-clinical multi-center study. For the actual application through the proposed system, 32 COVID-19 non-clinical efficacy studies were conducted by six animal [[Biosafety level|Biosafety level 3]] (BSL-3) institutions in South Korea from July 2020 to February 2021 via research funding. During the implementation period, the project progress was monitored by the chief director in real time through the management of the data entry process for each organization. In addition, the administrators of each institution were able to check and manage the data entry status of the researchers in real time. Researchers were able to review the status of the data entry by outputting descriptive statistics and graphics, such as histograms or box plots. The administrator of the institution mainly used the function for automatically generating a report based on data entry, which enabled the efficient management and monitoring of the study. | |||
As research on infectious diseases steadily increases after the COVID-19 pandemic, such studies are usually conducted as joint research involving multiple centers. [22,23,24,25,26] Therefore, a system for efficiently managing multi-center laboratory data is required, and standardization is essential for integrating the units of different studies. The proposed COVID-19 non-clinical trial LIMS, designed for organizing laboratory sample management, analysis, and reporting of non-clinical COVID-19 test results, can be beneficial for researchers in large-scale studies where multiple organizations participate in the production of data. The proposed system enables experimental data management, data sharing, and data analysis through a web-based interface. The system focuses on practical functionality such as the construction of management systems, data [[quality control]] processes, various graphics and analysis modules, result reporting functions, and data security and notification functions by systematically implementing access management, test execution, auditing, and comparisons. Large-scale data can therefore be managed more quickly and effectively. In particular, the proposed system is focused on multi-center laboratory data collection and the prevention of various problems owing to the non-standardization of the data coding processes at each institution. Furthermore, it was shown through a streamlined analysis that multi-center non-clinical trials can be systematically conducted. | |||
Revision as of 15:39, 17 July 2022
| Full article title | Laboratory information management system for COVID-19 non-clinical efficacy trial data |
|---|---|
| Journal | Laboratory Animal Research |
| Author(s) | Yoon, Suhyeon; Noh, Hyuna; Jin, Heejin; Lee, Sugyoung; Han, Soyul; Kim, Sung-Hee; Kim, Jiseon; Seo, Jung S.; Kim, Jeong J.; Park, In H.; Oh, Jooyeon; Bae, Joon-Yong; Lee, Gee E.; Woo, Sun-Je; Seo, Sun-Min; Kim, Na-Won; Lee, Youn W.; Jang, Hui J.; Hong, Seung-Min; An, Se-Hee; Lyoo, Kwang-Soo; Yeom, Minjoo; Lee, Hanbyeul; Jung, Bud; Yoon, Sun-Woo; Jang, Hung-Ah; Seok, Sang-Hyuk; Lee, Yu J.; Kim, Seo Y.; Kim, Young B.; Hwang, Ji-Yeon; On, Dain; Lim, Soo-Yeon; Kim, Sol P.; Jang, Ji Y.; Lee, Ho; Kim, Kyoungmi; Lee, Hyo-Jung; Kim, Hong B.; Park, Hun W.; Jeong, Dae G.; Song, Daesub; Choi, Kang-Seuk; Lee, Ho-Young; Choi, Yang-Kyu; Choi, Jung-ah; Song, Manki; Park, Man-Seong; Seo, Jun-Yeong; Nam, Ki T.; Shin, Jeon-Soo; Won, Sungho; Yun, Jun-Won; Seong, Je K. |
| Author affiliation(s) | Seoul National University, RexSoft Corp., Yonsei University College of Medicine, Korea University College of Medicine, International Vaccine Institute, Konkuk University, Seoul National University Bundang Hospital, Chonbuk National University, Korea University, Korea Research Institute of Bioscience and Biotechnology, Kangwon National University, National Cancer Center, Dongguk University, Yonsei University College of Medicine, |
| Primary contact | snumouse at snu dot ac dot kr |
| Year published | 2022 |
| Volume and issue | 38 |
| Article # | 17 |
| DOI | 10.1186/s42826-022-00127-2 |
| ISSN | 2233-7660 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://labanimres.biomedcentral.com/articles/10.1186/s42826-022-00127-2 |
| Download | https://labanimres.biomedcentral.com/track/pdf/10.1186/s42826-022-00127-2.pdf (PDF) |
|
|
This article should be considered a work in progress and incomplete. Consider this article incomplete until this notice is removed. |
Abstract
Background: As the number of large-scale research studies involving multiple organizations producing data has steadily increased, an integrated system for a common interoperable data format is needed. For example, in response to the coronavirus disease 2019 (COVID-19) pandemic, a number of global efforts are underway to develop vaccines and therapeutics. We are therefore observing an explosion in the proliferation of COVID-19 data, and interoperability is highly requested in multiple institutions participating simultaneously in COVID-19 pandemic research.
Results: In this study, a laboratory information management system (LIMS) has been adopted to systemically manage, via web interface, various COVID-19 non-clinical trial data—including mortality, clinical signs, body weight, body temperature, organ weights, viral titer (viral replication and viral RNA), and multi-organ histopathology—from multiple institutions. The main aim of the implemented system is to integrate, standardize, and organize data collected from laboratories in multiple institutes conducting COVID-19 non-clinical efficacy testing. Six animal Biosafety level 3 (BSL-3) institutions proved the feasibility of our system. Substantial benefits were shown by maximizing collaborative high-quality non-clinical research.
Conclusions: This LIMS platform can be used for current and future outbreaks, leading to accelerated medical product development through the systematic management of extensive data from non-clinical animal studies.
Keywords: SARS-CoV-2, COVID-19, non-clinical, laboratory information management system, data
Background
The novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has resulted in the serious respiratory-related coronavirus disease 2019 (COVID-19) pandemic. It was first reported in Wuhan, the capital of Hubei, China, in late December 2019 and has attracted international attention. [1] After four months, it spread to multiple countries, leading to more than one million confirmed cases worldwide. [2] SARS-CoV-2, a single-stranded RNA-enveloped virus [3], enters the host cells by binding its structural spike (S) protein to cell surface receptors, angiotensin-converting enzyme 2 (ACE2.) [4] After entering the host cells, the virus hijacks the cell to undergo viral replication. [5] During this process, an aberrant host immune condition called a cytokine storm can occur in response to SARS-CoV-2 infection, which is characterized by high concentrations of pro-inflammatory cytokines and chemokines, including tumor necrosis factor-α, interleukin-1, and interleukin-6 [6], resulting in severe COVID-19 associated with excessive inflammation. [7]
Along with understanding the mechanism of transmissibility and pathogenesis of SARS-CoV-2, significant efforts are being devoted to developing novel vaccines and therapeutic agents to prevent or treat COVID-19. In particular, various COVID-19-related research has been conducted with the participation of multiple organizations. Among such studies, animal models of monkeys, cats, ferrets, hamsters, and human angiotensin converting enzyme 2 (hACE2) transgenic mice [8], capable of conducting controlled experiments, have made significant contributions to vaccine development. [9,10,11,12,13] (Note: Though the U.S. Food and Drug Administration doesn't consider animal studies to be part of "non-clinical laboratory studies," many research organizations with animal lab capabilities still use "non-clinical" to include and describe these sorts of animal modelling studies.[1]) Because various parameters such as mortality, clinical signs, body weight, organ weights, viral titer (viral replication and viral RNA), and multi-organ histopathology should be comprehensively analyzed for a reliable interpretation of the results, there is an increasing need for an integrated system in which multiple institutions can apply a streamlined analysis and input the results simultaneously when conducting large-scale non-clinical efficacy tests of a COVID-19 vaccine.
When experimental data are collected from multiple institutions, “organization-level effects” may occur owing to non-standardization in the data coding process and coding errors, which can act as a batch effect in a data analysis. In large-scale experimental multi-center research, it is necessary to prevent the loss of reliability in the data analysis caused by batch effects; however, this is a difficult goal to achieve without an integrated system. An effective approach to overcoming the limitations caused by the prior-mentioned batch effects is to use a laboratory information management system (LIMS). [14] A LIMS systematizes the research process by automating data-related tasks that were conducted manually using information technology, and simultaneously realizes various needs, such as the standardization of experimental data, sharing, problem tracking, and the creation of reports based on document templates. The accumulated data stored in a LIMS can be useful in understanding the current clinical knowledge and making decisions, which is a remarkable advantage in research on rapidly changing epidemics such as COVID-19.
Despite the advantages of a LIMS, no systems allowing multiple institutions to participate simultaneously in research related to COVID-19 have yet been proposed among developed open-source LIMSs. Given that, we aimed to clearly understand the standard procedures of non-clinical trials for COVID-19 therapeutics and vaccines and to design and implement a LIMS suitable for these research purposes.
Results
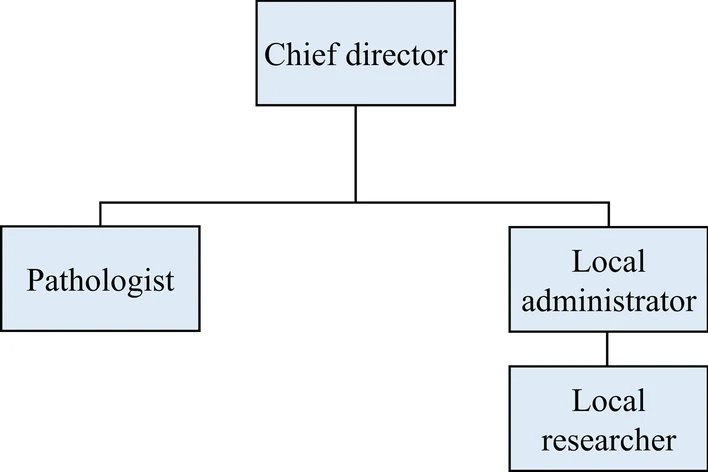
User management and functionalities
Our COVID-19 non-clinical trial LIMS has four different types of users with different authorizations to handle multiple organizations: a chief director, a local administrator and researcher from each organization, and a pathologist (Fig. 1). The role of the chief director includes new project initiation, modification, deactivation, user account management, and data management. The chief director can also control the registration of new organizations and the removal of existing organizations. The role of the local administrators is the same as that of the chief director, except that they are limited to the organization to which they belong. This enables the efficient management of each organization, as well as data security. Researchers can manage data entry and removal and check the project's progress. Pathologists can conduct pathological analyses, pathological data entry, and related functions. Furthermore, multiple user types for multiple roles are allowed. If multiple authorizations are granted, the screen manual is shown based on the user’s choice.
|
The proposed system, which applies a convenient user interface, was created to provide many functions related to data management for multi-center research. The detailed function list is presented in Table 1.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Functions can be classified into four different categories: access management, trial, audit, and comparison. The access management function forms the basis of the system and manages the user group. The trial function controls the entire project process, including initiation, progression, and finalization. The audit function records all activities that occur in the system and implements data entry, cancelation, and restoration. The comparison function provides several plots according to the investigational product (new drug/repositioning, vaccine/therapeutic), sponsor, experimental animals (hamster model/hACE2 mouse model), research objective, and study site. It performs access control and visualization by classifying all data accumulated in the system.
Data security and audit
The proposed system provides several functions for data security and audits. For security, an access management system based on the access control list (ACL) [15] was incorporated to protect data entry. Unintended data leakage was minimized by providing limited access points to users according to the user access levels. For each running instance, data security at the terminal level was obtained through secure HTTPS-based communication. For audits, the proposed system records, monitors, and manages all user activities, including data entry, by generating a separate database. Furthermore, it provides a history-tracking function to maintain data integrity. The chief director can identify the path of a data contamination and restore it to its original state, which minimizes the problem related to data entry or damage to the raw data.
Process management
Data entry and monitoring are possible for each institution, which makes it possible for the local administrator of each institution to check the data status in real time. It is also composed of a system in which the chief director supervises the overall project progress of each organization in real time. In addition, by systematically managing various types of non-clinical data and enabling collaboration between multiple organizations, researchers are expected to be able to quickly derive the desired results. Figure 2 shows the workflow of the COVID-19 non-clinical trial LIMS for vaccine and therapeutics treatments.
|
Graphics and analysis functions
The COVID-19 non-clinical trial LIMS provides various visualizations that can review the data characteristics at each stage for each institution, as well as optimized visualizations such as line diagrams, bar diagrams, and survival curves according to the data type for awareness of current situations and early detection of unusual situations. The proposed system generates suitable plots for a correspondence analysis according to the data types. In particular, a comparison function was developed to facilitate mutual comparisons according to user-selected conditions, and the system can also generate easy-to-read study-specific graphs.
Data downloading and result reporting
After obtaining approval from the chief director and administrators of each organization, the stored data can be downloaded as an Excel or Word file. In particular, it is possible to create an integrated Word file that includes a data description table, summary plot, and raw data, and all currently entered data can be downloaded as a multi-sheet Excel file with a single click for convenient data management. In addition, the proposed system can be used to monitor the data entry status of each participating organization and simultaneously create reports with a function for automatically summarizing the analysis results.
Notification
In applying important commands such as research initiation and termination, a notification system through a mail and messenger (limited to South Korea) notification service was established for relevant individuals such as chief directors and administrators. When a major change occurs in the research, the manager can quickly identify and respond to any unintended situations.
Data management and recording
Data entry control can be achieved according to the characteristics of each data type, and the user input is automatically controlled according to the condition of each stage of the study. Human error is minimized by limiting the range of unobservable values for each data parameter, and entry omission is prevented by notifying the data that need to be input according to the investigation schedule, current date, and data entry status (Fig. 3). In addition, all data entry activities were recorded, and an audit function was reviewed for each dataset. If necessary, an unsuitable data entry can be canceled to prevent data errors. On the data entry screen, users can check all data during the full research period (Fig. 4A) or partial data by filtering the data based on date (Fig. 4B).
|
|
Discussion
Non-clinical studies are essential for identifying the potential effects of promising drug candidates prior to entering human clinical trials. [16] Numerous animal studies have been conducted in the development of therapeutic agents and vaccines during the recent COVID-19 pandemic, leading to the need for rapid and accurate non-clinical efficacy analyses in standardized animal study design protocols. [9, 10] The inoculation of hACE2 transgenic mice with SARS-CoV-2 induced weight loss, increased the viral titer in the lungs, and prompted histopathological changes with increased cytokine levels and interstitial pneumonia. [17, 18] In addition, infection with SARS-CoV-2 in a golden Syrian hamster model induced signs resembling COVID-19 disease, including decreased activity, weight loss, high lung viral load, and the presence of T-lymphocytes in the respiratory tract. [19,20,21] Therefore, systematic management of these extensive data—including mortality, clinical signs, body weight, body temperature, viral titer (viral replication and viral RNA), organ weights, and multi-organ histopathology—that can be observed in COVID-19 research using animal models such as hACE2 transgenic mice and hamsters is required for a reliable evaluation and development of a therapeutic agent or vaccine.
The implemented LIMS has many distinctive features providing many user-friendly functions. First, the implemented system classifies COVID-19 non-clinical trial topics into six sub-themes (investigational product, types of investigational product, sponsor, experimental animals, research objective, and study site). It also applies access control and visualization by classifying the research accumulated in the system. Second, using a streamlined analysis, it creates a four-step process related to research progress, pathology analysis, result entry, and report generation. Access control and customized user interfaces are provided for each step, which highlights the work to be done and provides step-by-step guidance as a project planner. Third, a pathology analysis was incorporated to organize various types of pathology data and support the resulting input from pathologists. Finally, the data are organized systematically, and raw data, summary tables, and graph visualizations are documented. Many of these features were found not to be available in open-source LIMS. Table 2 provides a brief comparison of other existing open-source LIMS and the proposed COVID-19 non-clinical trial LIMS.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
The COVID-19 non-clinical trial data management system developed in this study consists of a mode for the chief director, administrators, researchers of each organization, and pathologists. It aims to improve the completeness of the database by managing the data entry process for each institution. To confirm this, we applied the proposed system to an actual COVID-19 non-clinical multi-center study. For the actual application through the proposed system, 32 COVID-19 non-clinical efficacy studies were conducted by six animal Biosafety level 3 (BSL-3) institutions in South Korea from July 2020 to February 2021 via research funding. During the implementation period, the project progress was monitored by the chief director in real time through the management of the data entry process for each organization. In addition, the administrators of each institution were able to check and manage the data entry status of the researchers in real time. Researchers were able to review the status of the data entry by outputting descriptive statistics and graphics, such as histograms or box plots. The administrator of the institution mainly used the function for automatically generating a report based on data entry, which enabled the efficient management and monitoring of the study.
As research on infectious diseases steadily increases after the COVID-19 pandemic, such studies are usually conducted as joint research involving multiple centers. [22,23,24,25,26] Therefore, a system for efficiently managing multi-center laboratory data is required, and standardization is essential for integrating the units of different studies. The proposed COVID-19 non-clinical trial LIMS, designed for organizing laboratory sample management, analysis, and reporting of non-clinical COVID-19 test results, can be beneficial for researchers in large-scale studies where multiple organizations participate in the production of data. The proposed system enables experimental data management, data sharing, and data analysis through a web-based interface. The system focuses on practical functionality such as the construction of management systems, data quality control processes, various graphics and analysis modules, result reporting functions, and data security and notification functions by systematically implementing access management, test execution, auditing, and comparisons. Large-scale data can therefore be managed more quickly and effectively. In particular, the proposed system is focused on multi-center laboratory data collection and the prevention of various problems owing to the non-standardization of the data coding processes at each institution. Furthermore, it was shown through a streamlined analysis that multi-center non-clinical trials can be systematically conducted.
References
- ↑ "Preclinical and Nonclinical Studies—What Is the Difference, and Where in Your Program Should They Fall?". Premier Consulting. 29 July 2020. https://premierconsulting.com/resources/blog/preclinical-and-nonclinical-studies-what-is-the-difference-and-where-in-your-program-should-they-fall/. Retrieved 27 May 2022.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation, grammar, and spelling. In some cases important information was missing from the references, and that information was added. The authors don't define "non-clinical trial" or "non-clinical research" in the original; for this version, a presumed definition and accompanying citation is provided to the reader for additional context.