Journal:A model for design and implementation of a laboratory information management system specific to molecular pathology laboratory operations
| Full article title | A model for design and implementation of a laboratory information management system specific to molecular pathology laboratory operations |
|---|---|
| Journal | The Journal of Molecular Diagnostics |
| Author(s) | Tomlinson, Eban; Goodman, Jennifer; Loftus, Margaret; Bitto, Stephen; Carpenter, Erica; Oddo, Richard; Judis, LuAnn; Ali, Shabab; Robinson, Wyatt E.; Carver, Miranda; Ganea, Mariana; McDonnell, Kristen; O'Neill, Diane; Starbuck, Jennifer; Johnson, Eric; Meister, Erik; Pohl, Jonathan; Spildener, Jessica; Shurtleff, Sheila; Sovie, Sheryl; Melendez, Cathleen; Krebs, Pamela; Riley, Jacquelyn D.; Wensel, Christine; Astbury, Caroline; Azzato, Elizabeth M.; Bosler, David S.; Brock, Jay E.; Cook, James R.; Cheng, Yu-Weu; Tu, Zheng J.; Cruise, M.; Henricks, Walter H.; Farkas, Daniel H. |
| Author affiliation(s) | Semaphore Solutions, Cleveland Clinic |
| Primary contact | Email: farkasd2 at ccf dot org |
| Year published | 2022 |
| Volume and issue | S1525-1578(22) |
| Page(s) | 00012-5 |
| DOI | 10.1016/j.jmoldx.2022.01.002 |
| ISSN | 1525-1578 |
| Distribution license | Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International |
| Website | https://www.jmdjournal.org/article/S1525-1578(22)00012-5/fulltext |
| Download | https://www.jmdjournal.org/action/showPdf?pii=S1525-1578%2822%2900012-5 (PDF) |
Abstract
The Molecular Pathology Section of Cleveland Clinic (Cleveland, OH) has undergone enhancement of its testing portfolio and processes. An electronic- and paper-based data management system was replaced with a commercially available laboratory information management system (LIMS) solution, a separate bioinformatics platform, customized test-interpretation applications, a dedicated accessioning service, and a results-releasing solution. The LIMS solution manages complex workflows, large-scale data packets, and process automation. However, a customized approach was required for the LIMS since a survey of commercially available off-the-shelf (COTS) software solutions revealed none met the diverse and complex needs of Cleveland Clinic's molecular pathology service. The project utilized the expertise of clinical laboratorians, pathologists, genetics counselors, bioinformaticians, and systems analysts in partnering with software-engineering consultants to design and implement a solution. Concurrently, Agile software development best practices were formulated, which may be emulated for scalable and cost-effective laboratory-authored software.
Keywords: molecular pathology, laboratory information management system, LIMS, bioinformatics, software development, Agile-Scrum
Introduction
Data management needs in clinical molecular pathology laboratories differ in substantive ways from those in other clinical laboratories and anatomical pathology labs.[1][2][3][4] Conventional laboratory information systems (LISs) historically have not inherently supported the needs of molecular pathology laboratories to the extent that they have in other laboratory disciplines and operations.[2] Molecular pathology laboratories have often relied on a combination of manual methods, spreadsheets, and nonintegrated and/or modular software to meet data management and operational needs.
Such was the situation in the Molecular Pathology Section, Pathology & Laboratory Medicine Institute, Cleveland Clinic (Cleveland, OH) in early 2017. A revitalization and growth plan for the section—which included expansion of personnel, equipment, testing platforms, and test development—was undertaken. An improvement deemed fundamental to this re-invention process was a new laboratory information management system (LIMS) to reduce and eventually replace the outdated, largely paper- and electronic spreadsheet–based information and workflow management system.
While the overhaul of the laboratory service was substantial, the focus of this report is limited to a description of a replicable process for the modernization of information and workflow management specific to a clinical molecular pathology laboratory. The process employed a customer–vendor relationship. Out of necessity, the relationship was a partnership due to the complementary and nontechnical skill sets of each party. The overall goal was to digitize workflow based on paper and Excel (Microsoft, Redmond, WA) across a complex, multidisciplinary molecular pathology and cytogenomics clinical laboratory service. This digitization was accomplished through a custom-architected software solution.
This article describes a model for the design and implementation of a LIMS that meets the diverse information management needs of a full-service clinical molecular pathology laboratory, emphasizes the integral importance of a well-structured development process, and describes a novel application of modern software-based project management methods and third-party partnerships for building and deploying a LIMS suitable for a modern molecular pathology laboratory.
Materials and methods
Scope definition
The goal of the project was to deploy a LIMS to modernize data and workflow management of the clinical molecular pathology laboratory, including cytogenomics workflows. The project encompassed a complete overhaul of paper and electronic spreadsheet–based data handling methods into a comprehensive, integrated electronic platform. Objectives were identified to define the scope and focus priorities of the project:
- Design and implement an electronic information system to support the specialized requirements and workflow of a full-service clinical molecular pathology laboratory, and to fulfill needs unmet by conventional LISs.
- Support the range of clinical testing performed, including adult, pediatric, and somatic mutation detection in tumor tissues, blood, and bone marrow; chromosome analysis; germline genetic testing for diagnosis and/or carrier screening; and pharmacogenomics.
- Support the range of complex data management of analytical methods (and applications) in use, including polymerase chain reaction (PCR); reverse transcription polymerase chain reaction (RT-PCR); next-generation sequencing (NGS); B- and T-cell gene rearrangement (PCR + capillary electrophoresis); chromosomal microarray analysis; fluorescence in situ hybridization (FISH); cytogenetic testing; and (genotyping by) mass spectrometry.
- Integrate into the information technology (IT) environment of the institution for accessioning and resulting, particularly the electronic health record (EHR) system and the conventional clinical laboratory and anatomical pathology information systems. (More specifically, Cleveland Clinic was using Sunquest Laboratory version 7.2 [Sunquest Information Systems, Tucson, AZ] and CoPathPlus version 2014 [Cerner, N. Kansas City, MO] at the time, but these are now in the process of being replaced by Epic Beaker version November 2020 [Epic Systems, Verona, WI].)
Needs assessment and partner selection
Given the complexity and multifaceted nature of the project, two key needs were recognized: professional project management guided by domain expertise in molecular testing laboratories, and experienced and expert software development professionals who could bring to bear state-of-the-art software development tools and techniques. Between two and four software developers were dedicated to the project at various stages. One business-based quality assurance analyst was included, who served as the conduit for the translation of laboratory-specific requirements into software requirements. A Ph.D.-level molecular biologist with project management expertise was hired to apply process rigor and organization to direct LIMS development and implementation (among other projects).[5] Laboratory-based personnel participated in their various subject matter domains as their primary responsibilities to patient care allowed. Partnership with a team of software engineers was established. This team worked with molecular pathologists and laboratory personnel (subject matter experts) to design and build the full software solution. Peer-to-peer relationships were formed between the clinical laboratory project manager and software project managers as control points for the project.
Prior to the beginning of this extensive development project, due diligence was undertaken to evaluate whether, or to what extent, commercially available information systems could meet the needs at hand. This assessment indicated that some systems on the market could provide some of the functional requirements, and that development entirely from scratch was not required. NGS–centric LIMS software was selected for licensure (BaseSpace Clarity LIMS by Illumina, San Diego, CA). As licensed, this software was focused narrowly on supporting particular elements of NGS workflows. Its distinguishing characteristics were customizability and extensibility that could ultimately support the objectives of the laboratory, including a wide portfolio of molecular diagnostics tests performed and the need for systems integration. Compatibility with the third-party commercial bioinformatics software that the laboratory had previously chosen (Clinical Genomics Workspace version 6.15.1 by PierianDx, Creve Couer, MO) was another key deciding criterion.
Technology platforms and software tools
The software engineering team used Jira and Confluence web-based software (Atlassian, San Francisco, CA) to organize development tasks and to project documentation, respectively. Version control was implemented in the web-based GitHub software host (San Francisco, CA). A continuous-integration pipeline (Table 1) was built using TeamCity software version 2019.1 (JetBrains, Prague, Czech Republic). The PyCharm version 2019.1 (JetBrains), Docker Desktop Community version 2.1.0.5 (Docker, Palo Alto, CA), VirtualBox version 6.1 (Oracle, Austin, TX), and Postman version 8.1 (Postman, San Francisco, CA) software solutions were used to facilitate local testing and development.
| ||||||||||||||||||||||||||
Python (Python Software Foundation, Wilmington, DE), a popular, general-purpose, high-level programming language with well-documented, well-supported engineering standards, was used to develop external program plug-ins, automated test scripts, report templates, and other services. A report-generation and sign-out application named AVRO (analytically validated reporting object; see subsequent sections) was built on a Python server with an Angular JavaScript front-end. For LIMS workflows, the open-source Python S4-Clarity library[6] was used to support batch-analyte data (spreadsheet-formatted) parsing, laboratory instrument integrations, complex library pooling, de-multiplexing, and other computations on analytes. The Jinja web-based template engine library[7] was used for default report-content templating to support clinical interpretation and clinical report generation in AVRO.
Project management
The general project management process used was Agile-Scrum in Jira, chosen by the owner of the critical path for project completion (i.e., the software development team). In the project, Agile-Scrum was supplemented with additional roles and processes tailored to work in the hybrid clinical laboratory service/software development consultant environment. It became obvious that definitions and vocabulary differed between these two groups of subject matter experts (i.e., clinical laboratory and software engineering professionals). Selected terms are defined in Table 1. Key roles and activities of the Agile-Scrum development process, as applied in this project, are described in the next subsection.
Key roles
Four key roles corresponding to major development and validation phases were identified as gatekeepers of the development process. These gatekeepers controlled the promotion of a deliverable through the process and, thus, control of a rapidly evolving process was retained. The subject matter experts in these roles worked in tight collaboration to ensure that the highest possible quality was achieved. Note that at relevant steps, bioinformatics professionals, Ph.D.-level laboratory director-designees, and/or M.D.-level pathologists reviewed and approved changes as appropriate.
Key Role 1: Business Analyst (Extant within Software Development Team)
The business analyst assumed responsibility and accountability for the elicitation and decomposition of requirements into granular requirements that were logically defined and complete. This work output was added to the stakeholder consensus process.
Key Role 2: Software Development Team
The software development team was the consumer of approved granular requirements and the developer of software designed to meet those requirements. This team was involved in developing automated tests to validate that the software written did in fact meet the granular requirements as specified. This work output was collected into a release candidate and, upon passing the automated tests, was released into Phase 3, user acceptance testing. This team was self-organizing and chose the Scrum framework to align themselves into two-week iterations (.e., sprints).
Key Role 3: Clinical Systems Analyst (Extant within Laboratory Team)
The clinical systems analyst served as the first round of testing for the laboratory service, owing to the proximity of the role to the laboratory and to the expertise in the laboratory's process and informatics architecture. The work assigned to this role was to coordinate and, in some cases, to perform the tasks in development/validation (Phase 3), user acceptance testing. The output of this work was a pass/fail decision that either sent the release candidate back to development or promoted the release candidate to staging.
Key Role 4: Laboratory Manager
The laboratory manager signed documentation approving the release candidate for promotion to the production environment after review of successful pre-production runs that were performed using test samples in the laboratory and after verification that all test plans and checklists were completed.
Key activities
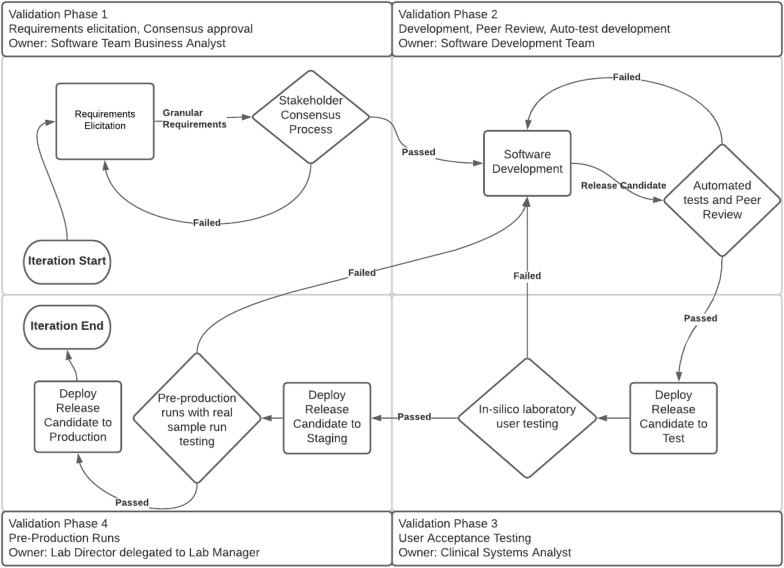
The major phases of software development and validation are shown in Figure 1. All activities were coordinated utilizing a series of modular processes that worked together to produce a performant and validated software installation consisting of licensed software, configuration of that software, and customized software to supplement the licensed software (e.g., AVRO and the Health Level 7 (HL7) version 2.3 [Health Level Seven International, Ann Arbor, MI] interface.)
|
The highest-order process consisted of four development/validation phases:
- Phase 1: Identify requirements and group them into architecturally independent release candidates logically small enough to enable rapid development (requirements elicitation).
- Phase 2: Monitor and facilitate the development of automated testing and peer-review process on release candidates (software development).
- Phase 3: Coordinate the promotion of release candidates from the development environment to the testing environment (user acceptance testing).
- Phase 4: Coordinate the promotion of the release candidate from the testing environment to the staging environment for preproduction run, and subsequently to the production in silico environments in such a way as to avoid patient-care impacts (preproduction runs, progression to user-accepted version, and deployment).
Development/Validation Phase 1: Requirements elicitation
The project consumed hundreds of person hours in elicitation meetings, the goals of which were to:
- Establish the location of relevant domain knowledge.
- Establish detailed requirements and shared terminology.
- Achieve stakeholder group consensus on each granular requirement.
- Promote granular requirements to the approved backlog of work.
Translation of end-user system functionality into workable development items was a task shared between the laboratory and software teams. The laboratory team solicited input from subject matter experts and prepared documentation. The software business analyst then used that documentation to produce software development work items that fulfilled the final system requirements specified by the laboratory team. System requirements documentation included written requirements, text-based documents, logic tables (e.g., user-role permission maps, diagrams, and flowcharts), spreadsheets, native-instrument data files, laboratory standard operating procedures, and test validation plans.
Software deliverables were detailed and tracked in Jira. Laboratory objectives were described in common language, then passed to the software team for decomposition. Software development tasks were identified during decomposition and completed by the software team using Scrum. The percentage of completion reports for each objective were tracked using Jira and reviewed weekly by the joint project management group. Impediments to progress were identified and delegated to the appropriate team members for resolution. Common impediments included:
- language ambiguity of requirements,
- technical difficulty in mapping laboratory requirements to the software, and
- requirements without a test plan or for which generating a test plan to be used in practice was challenging.
Development/Validation Phase 2: Software development
Software development proceeded in two-week iteration cycles (sprints). Members of the clinical laboratory and software engineering teams continuously evaluated and improved the software development processes throughout the project. The teams incorporated the requirements of the Health Insurance Portability and Accountability Act (HIPAA) to ensure that protected health information was handled appropriately. For example, scripting of mock samples was developed with convincing, yet mock, data. This scripting enabled the engineering team to test the system in a realistic fashion without the use of patient information.
The engineering team adhered to the following technical process while working within sprints:
- A team member chose a development task from the approved requirements to work on during the current iteration (sprint backlog).
- Once completed, each development task was peer-reviewed by other software engineers prior to inclusion in the master codebase, an industry-standard quality assurance practice.
- Automated tests were generated during development and run before and after each development task was committed to the master codebase, to identify problems in the task and in the integration of the task into the codebase, respectively. Automated tests generally consisted of scripts that generated representative mock samples and utilized the Clarity LIMS application programming interface to move those mock samples through the workflow while replicating user interaction at each step, again using the API.
- When the development of a full software deliverable was completed (release candidate), it was assembled atop all previous deliverables on a replica LIMS (test server).
- The release candidate was tested for both successful and potentially erroneous scenarios, in accordance with the requirement specification(s).
Development/Validation Phase 3: User acceptance testing
In Phase 3 of software validation, functional testing was performed by the laboratory team and led by the clinical systems analyst, who reported results and discrepancies back to the software team at each stage for collaborative triage. A go/no-go decision was made based on the severity of the discrepancy and the viability of a workaround being utilized until the next release candidate could be deployed. Discrepancies were addressed by the software team in order of priority and redeployed through reiteration of the above process.
Development/Validation Phase 4: Preproduction runs, progression to user-accepted version, and deployment
The following process was used for preproduction runs, progression to user-accepted version, and deployment:
- After all tests were passed, the software vendor deployed the validated release candidate to the staging environment of the laboratory.
- Successful execution of the user acceptance testing phase triggered full system–level preproduction runs by laboratory personnel in the staging environment.
- Successful preproduction run completion was followed by scheduled deployment of the release candidate to the production environment, which, in some cases, constituted several releases at once.
- Full deployment was facilitated jointly by software and laboratory IT personnel. Full system–level tests, as well as preproduction run checklists and end-to-end testing scripts, were utilized to validate the performant nature of the final deployment(s).
Results
System design: Workflow analysis and support
The LIMS organizes and handles the workflows of the clinical laboratory, from specimen accessioning through to laboratory analysis and issuing reports to ordering physicians. The system also coordinates tasks among staff. The LIMS automates several laboratory processes, such as spreadsheet parsing or container tracking, and is compatible with laboratory instrumentation, such as automated liquid handlers and quality control instruments, either through an off-the-shelf or custom-built integration. Substantial customized LIMS workflow design and engineering were required to model the testing procedures of the laboratory and to organize the constituent processes, including sample accessioning, nucleic acid (i.e., DNA, RNA, or total nucleic acid) purification, direct PCR testing, genotyping by mass spectrometry, sequencing, chromosome testing, data analysis, and all associated quality control steps.
A key project requirement was organization and refinement of workflows within the LIMS. Wherever possible, shared processes were identified and excluded from test-specific workflow building. For example, sample accessioning and DNA extraction were built out as shared protocols. Templating (sharing) at the step and protocol layers were combined with unique elements to generate end-to-end workflows specific to each test.
The LIMS was engineered to support the build of workflows in a hierarchical fashion in which workflows contain protocols which, in turn, contain steps (Table 2). In steps, the work of the end user is performed, on either a single sample or a batch of samples. Protocols are a collection of steps organized around the standard operating procedures of a laboratory, and workflows comprise the protocols required to execute a test. Custom-built workflows are shown in Table 3.
| |||||||||||||||
| ||||||||||||
System features and integration
LIS–HL7 interface for sample accessioning
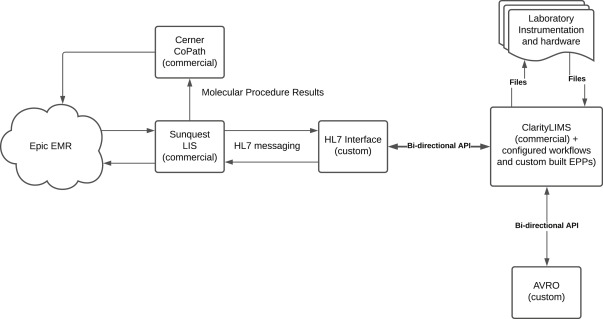
LIS applications typically connect to other patient health care applications such as health insurance policy administration systems and EHRs.[8] To function effectively, an LIS must be interoperable with these health care record systems through the adoption of a common standard.[2] The HL7 standard was utilized as the form of communication between the LIS and the LIMS through a customized interface (Figure 2).
|
The interface between the existing Sunquest Laboratory and CoPathPlus LISs of the laboratory and the new LIMS streamlined the flow of patient data received from the health care record systems into sample accessioning. A customized HL7 listener service was built and configured to interface with the LIS. This service verifies whether the HL7 message received is valid and, if so, generates a sample with all required sample information fields and accessions it directly into the LIMS. Required sample fields including, but not limited to, medical record number, last name, and collection date are extracted from the HL7 message and mapped to user-defined LIMS fields to preserve vital information for mapping test results to the correct patient downstream. The original inbound HL7 message was preserved on the sample to resolve any ambiguous data mapping in the LIMS. Error logs are generated for review when HL7 messages either cannot be processed or produce errors.
The LIS application was integrated with the HL7 interface while its essential functionality and interoperable links were preserved. After software build, the LISs still collected and organized patient health care data. However, software management of the clinical test workflows and process automation were improved with LIMS implementation.
Instrumentation
Integration with nucleic acid purification robotics and spectrophotometers (pre-analytical) and molecular diagnostics–related analytical platforms were facilitated through file transfer systems built on shared network locations. Integrations followed the same process:
- A plate map was generated and displayed to laboratory personnel at the appropriate LIMS workflow step.
- After the requisite samples were plated and the instrument completed its full cycle, an output file was directed to a shared network location available at each LIMS workstation.
- Laboratory personnel uploaded the output file to the appropriate (usually next) step in the LIMS workflow.
- An external program plug-in parsed data from the output file and saved discrete pieces to user-defined fields on the sample in the LIMS.
Clinical reporting application
To address the ultimate goal of every high-complexity clinical laboratory test (i.e., the issuance of a patient report for the physician who ordered the test), the customized AVRO was built. AVRO serves as an interface to manage the interpretation (post-analytical) component of laboratory test workflows. AVRO utilizes the authentication system of the LIMS, and therefore only LIMS users with the correct permission setting can access AVRO. AVRO runs on its own server, links directly to the LIMS, and polls specific steps to import reporting objects. It then determines which report template to assign based on the test code and displays test data to professional staff reviewers. This reporting application generates standards-compliant clinical reports for annotation and sign-out by molecular pathology professional staff. Upon sign-out, AVRO sends text reports to the automated clinical results–releasing service, which submits HL7-formatted data and issues a text-based copy of the report to the appropriate recipient.
AVRO was designed and implemented to harmonize results reporting and formatting within a dynamic multicomponent system of EMRs and LISs (Epic Beaker, Sunquest Laboratory, CoPathPlus, and Rhapsody version 6.6 [Lyniate, Boston, MA]) while maintaining a flexible LIMS configuration. (Rhapsody is an interface engine that sits between Epic and ancillary systems, e.g., Sunquest Laboratory, CoPathPlus.) Given that report fields are populated directly from LIMS user-defined fields, all LIMS data are available to the report template. Using Jinja2 Python templating enabled a programmatic approach to reporting (i.e., rules can be applied to text and data formatting). AVRO uses the LIMS application data layer and the LIMS-managed user credentials that can be integrated with Lightweight Directory Access Protocol (LDAP) systems to support data provenance and added security. This approach obviated the architectural generation of a separate database and the duplication of user credentials. AVRO thus facilitates professional actions on samples that exist entirely within the LIMS.
The concept of report-based content defaults was identified from feedback after early implementations had been tested by the reporting professional staff. Reporting templates were customized for each test and then served the correct template by mapping test codes to those templates. Templates could be mapped to multiple test codes and were maintained independently for maximum flexibility in content.
Opening a report in AVRO presents a template based on the above logic, populated with interpretation text derived from the results interpretation code applied in the analysis portion of the workflow. The molecular pathologist user is able to modify and customize all of the populated content prior to sign-out. At any point during the report-building process, AVRO allows the user to preview the report.
A customized results–releasing service was built into the LIMS as a set of external program plug-ins for report release. This step was designed such that the system monitors for newly signed-out reports in AVRO every 30 seconds via the AVRO API. The report is generated and sent back to the LIS, and then the report file(s) are attached to the completed step in the LIMS.
Audit trails were available as base functionality in the LIMS and did not require customization. Building AVRO atop the audit trail data layer facilitated auditing through the reporting process up until submission of the outbound HL7 message. HIPAA compliance as implemented in the LIMS thus persisted through the reporting process, an efficiency afforded by leveraging of the existing software validation work extant in the off-the-shelf product. The LIMS tracks all of the elements specified by audit trail-related items per the College of American Pathologists Laboratory General Accreditation Checklist.
Resulting end-to-end software solution
The test management software system of the LIMS and AVRO interfaced with the pre-existing LIS architecture via customized software services, an HL7-bridged laboratory sample accessioning service, and an automated clinical results–releasing service through the LIMS. The newly integrated solutions automated the full end-to-end process of the molecular pathology laboratory, from sample accessioning to the issuance of final patient reports. These services eliminated the use of paper to track analytics data and mitigated the risks for bottlenecks and human error that tend to result from manual control of laboratory input and output processes. For example, accessioning bottlenecks have been mitigated by streamlining accessioning through the LIS–HL7 interface, and human errors due to data transcription have been reduced through data tracking in the LIMS and subsequent automated exchange of data between the LIMS, AVRO, and LIS.
An important predictor of success was the selection of a LIMS with robust API and customization support in order to address the significant Δ between off-the-shelf functionality[2] and the functionality required by the molecular service via the only pathway remaining, custom software engineering. Software engineers worked in virtual instances of the software environment to enable rapid revisions within existing and relevant system parameters. Imagining, designing, testing, and implementing the information management system described required significant human and financial resources. A list of advantages that have been accrued by this clinical laboratory, or that are anticipated in the near future, is shown in Table 4. Specific examples of the benefits the laboratory has experienced since LIMS implementation are listed in Table 5.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||
Discussion
The Molecular Pathology Section, Pathology & Laboratory Medicine Institute, Cleveland Clinic performs high-complexity clinical laboratory testing in support of the practice of genomic medicine. A revitalization and modernization program, begun in 2016, established the need for improvements in laboratory data management. Organization of clinical laboratory testing workflows in the twenty-first century is best matched to modern software and information management tools, not tools from the previous century (i.e., paper, Excel, flash drives). Education was pivotal in accelerating the project toward success. Educating all members in this team-based approach in respective domains led to the acknowledgment that both software development and molecular pathology (with inherent regulation) were of equal importance in the LIMS development process. Education, not unlike the build process itself, was accomplished iteratively and continually throughout the project. The more that subject matter experts from all disciplines understood each other, the better the team was able to optimize and improve each build iteration and, subsequently, execute the shared process successfully. The importance of partnership between the laboratory and software engineering teams was a lesson well learned.
Modeling clinical laboratory test workflows digitally and integrating a LIMS into the clinical hardware and software architectures of the laboratory required a significant amount of customized software engineering. Most commercial LIMS are touted as configurable. Principles and practices from the domains of computer science and software engineering were required to produce a maintainable and scalable LIMS implementation, alongside a greater set of laboratory software services and applications. Laboratories often do not have the resources in-house to integrate such a solution into their existing system(s). Software and process engineers were required to build effective customized test-workflow models in silico that automated processes and managed the inherent workflow data. Their role also included installing and successfully integrating the LIMS application alongside other supporting software that existed in the clinical informatics architecture. By employing a team-based collaborative approach between the clinical laboratory and software engineers, the existing software system was significantly upgraded, the use of paper and electronic spreadsheets was reduced (with the eventual goal of the complete elimination of paper), and efficiency was increased, putting the laboratory in a better position to scale up throughput and manage complex data.
The results reporting environment is complex; here we show one of the benefits of the LIMS implementation. The Pathology & Laboratory Medicine Institute (composed of two departments: anatomical pathology and laboratory medicine) uses Sunquest Laboratory to output test results. CoPathPlus is used to report anatomic pathology results and additional associated testing of tissue, cytology, and bone marrow specimens (i.e., special stains, FISH, cytogenetics, PCR, and NGS testing). Clarity LIMS streamlined reporting so that all molecular pathology tests are now reported back to Sunquest Laboratory and posted as a discrete item in the patient's EHR in Epic for the specific test. Tests that must be reported as part of an anatomic pathology report are sent back to CoPathPlus from Sunquest Laboratory so that clinicians may view a comprehensive report on that specimen.
Formulating an effective software development framework was important, considering the challenges in generating and operating customized clinical laboratory software. Such software must address testing intricacies and diverse molecular technologies within a complex clinical laboratory architecture. Rigorous building methods and techniques were needed to ensure build quality and maintenance sustainability.
Scrum best practices are valuable in the modernization of clinical laboratory information management workflows via custom software development. Scrum best practices are demonstrated to be extensible, scalable, and cost-effective. It is crucial that any given Scrum format be lightweight, with clearly defined roles and responsibilities of the participants; be comprehensively simple; and assume room for improvement.
Notably, a laboratory science framework for molecular biology research, termed LabScrum, has gained support in research-oriented molecular biology laboratories. LabScrum emulates the principles of Scrum software development. The writers of LabScrum illustrated three critical principles of the Scrum framework's utility in research laboratories.[9] These principles are valid for building customized information management software in the setting of clinical laboratory testing.
The adoption of some form of Scrum in developing customized laboratory software should focus on three essential components: transparency, inspection, and adaptation. LabScrum proponents have argued that a management framework based on empiricism is highly consistent with the scientific method.[9] Anecdotally, research scientists rigorously apply empiricism and the scientific method for results, or product, but often do not think empirically about tasks, techniques, or processes. Generating process visibility and analysis utilizing a framework for ongoing improvement can improve the quality of the science (i.e., more rigorous science), the quantity of the science (i.e., more productive science), and the quality of life of scientists (i.e., more sustainable science). In effect, to run a modern clinical laboratory necessitates having a laboratory function to interact with the processes of configuring or generating customized software, because, by definition, this software does not yet exist or, more accurately, must be customized for use in every clinical laboratory.
Limitations, lessons learned, and future work
The inability of the LIMS implementation to store unique patients in their own table with a one-to-many relationship between the patients table and the specimens table in the database is a limitation of the existing system. The system is maintained via IT department personnel, two full-time laboratorian, and one part-time laboratorian.
An important lesson learned in executing the LIMS project was the concept of a minimally viable product (MVP). Early in the project, the laboratory team was excited about the flexibility and customizability of the chosen LIMS product. In understanding what was possible and what was needed to elevate the laboratory's capabilities, overreaching and unrealistic goals were imagined. The team grew too ambitious, frustration accompanied scope creep, and project velocity decreased. Once it became clear that an MVP was, by definition, good enough, goals became simultaneously more modest and attainable. Thus the project's progress toward full execution accelerated over the last approximately 12 months of a 30-month effort from late 2017 to early 2020.
With LIMS launch, experience using the LIMS, and newly acquired local skills and knowledge, the laboratory team looks forward to independently adding more functionality via new customizations.
Acknowledgements
We thank Gordon Meyer, Justin Chant, Emily Wong, Jeremy Snell, Danny Hopkins, Mark Swinkels, Mark Luszniak, and the rest of the Semaphore team for their technical expertise; Wendy Nedlik for organizational management expertise, Hillard Meade for project management expertise; James Fenske for software implementation assistance; Michael Reese for financial analysis; Eric D. Hsi, MD, for leadership; and Jacqueline Urankar and Susan Brennan for excellent administrative support.
Funding
Supported by the Robert J. Tomsich Pathology & Laboratory Medicine Institute, Cleveland Clinic.
Conflict of interest
E.T., S.A., W.E.R., and C.M. are full-time employees of Semaphore Solutions, the vendor retained by Cleveland Clinic in the development and launch of the LIMS described in this article.
References
- ↑ Kang, Wenjun; Kadri, Sabah; Puranik, Rutika; Wurst, Michelle N.; Patil, Sushant A.; Mujacic, Ibro; Benhamed, Sonia; Niu, Nifang et al. (1 July 2018). "System for Informatics in the Molecular Pathology Laboratory: An Open-Source End-to-End Solution for Next-Generation Sequencing Clinical Data Management". The Journal of molecular diagnostics: JMD 20 (4): 522–532. doi:10.1016/j.jmoldx.2018.03.008. ISSN 1943-7811. PMC 6039793. PMID 29698836. https://pubmed.ncbi.nlm.nih.gov/29698836.
- ↑ 2.0 2.1 2.2 2.3 Myers, Charles; Swadley, Matthew; Carter, Alexis B. (1 September 2018). "Laboratory Information Systems and Instrument Software Lack Basic Functionality for Molecular Laboratories". The Journal of molecular diagnostics: JMD 20 (5): 591–599. doi:10.1016/j.jmoldx.2018.05.011. ISSN 1943-7811. PMID 30146005. https://pubmed.ncbi.nlm.nih.gov/30146005.
- ↑ Lee, Roy E.; Henricks, Walter H.; Sirintrapun, Sahussapont J. (1 March 2016). "Laboratory Information Systems in Molecular Diagnostics: Why Molecular Diagnostics Data are Different". Advances in Anatomic Pathology 23 (2): 125–133. doi:10.1097/PAP.0000000000000109. ISSN 1533-4031. PMID 26849819. https://pubmed.ncbi.nlm.nih.gov/26849819.
- ↑ Campbell, Walter S.; Carter, Alexis B.; Cushman-Vokoun, Allison M.; Greiner, Timothy C.; Dash, Rajesh C.; Routbort, Mark; de Baca, Monica E.; Campbell, James R. (1 May 2019). "A Model Information Management Plan for Molecular Pathology Sequence Data Using Standards: From Sequencer to Electronic Health Record". The Journal of molecular diagnostics: JMD 21 (3): 408–417. doi:10.1016/j.jmoldx.2018.12.002. ISSN 1943-7811. PMC 6521887. PMID 30797065. https://pubmed.ncbi.nlm.nih.gov/30797065.
- ↑ Brock, J.; Nedlik, W.; Farkas, D.H. (1 November 2018). "AMP Abstracts - TT002. A Process for New Clinical Laboratory Test Implementation" (in en). The Journal of Molecular Diagnostics 20 (6): 1016. doi:10.1016/S1525-1578(18)30401-X. PMC PMC7130024. https://linkinghub.elsevier.com/retrieve/pii/S152515781830401X.
- ↑ Semaphore Solutions (6 April 2021). "SemaphoreSolutions / s4-clarity-lib". GitHub. https://github.com/SemaphoreSolutions/s4-clarity-lib. Retrieved 04 August 2021.
- ↑ Ronacher, A. (18 May 2021). "pallets / jinja". GitHub. https://github.com/pallets/jinja. Retrieved 04 August 2021.
- ↑ Sinard, John H.; Gershkovich, Peter (2012). "Custom software development for use in a clinical laboratory". Journal of Pathology Informatics 3: 44–53. doi:10.4103/2153-3539.104906. ISSN 2153-3539. PMC 3551490. PMID 23372985. https://pubmed.ncbi.nlm.nih.gov/23372985.
- ↑ 9.0 9.1 May, L.; Runyon, T. (25 June 2019). "LabScrum: A Case Study for Agility in Academic Research Labs". Cutter. https://www.cutter.com/article/labscrum-case-study-agility-academic-research-labs-504061.
Notes
This presentation is faithful to the original, with minor changes to presentation, grammar, and punctuation. In some cases, important information was missing from the references, and that information was added. Everything else remains true to the original article, per the "NoDerivatives" portion of the distribution license.